Abstract
Gastrointestinal fungal dysbiosis is a hallmark of several diseases marked by systemic immune activation. Whether persistent pathobiont colonization during immune alterations and impaired gut barrier function has a durable impact on host immunity is unknown. We found that elevated levels of Candida albicans immunoglobulin G (IgG) antibodies marked patients with severe COVID-19 (sCOVID-19) who had intestinal Candida overgrowth, mycobiota dysbiosis and systemic neutrophilia. Analysis of hematopoietic stem cell progenitors in sCOVID-19 revealed transcriptional changes in antifungal immunity pathways and reprogramming of granulocyte myeloid progenitors (GMPs) for up to a year. Mice colonized with C. albicans patient isolates experienced increased lung neutrophilia and pulmonary NETosis during severe acute respiratory syndrome coronavirus-2 infection, which were partially resolved with antifungal treatment or by interleukin-6 receptor blockade. sCOVID-19 patients treated with tocilizumab experienced sustained reductions in C. albicans IgG antibodies titers and GMP transcriptional changes. These findings suggest that gut fungal pathobionts may contribute to immune activation during inflammatory diseases, offering potential mycobiota-immune therapeutic strategies for sCOVID-19 with prolonged symptoms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
16S and ITS sequencing data are available in the National Center for Biotechnology Information (NCBI) Sequence Read Archive with the accession code PRJNA732432. The single-cell transcriptome and ATAC sequencing data have been deposited in NCBI GEO under the accession number GSE196990. Source data are provided with this paper.
References
COVID-19 Dashboard. The Center for Systems Science and Engineering (CSSE) (Johns Hopkins University (JHU)) https://coronavirus.jhu.edu/map.html (2023)
Guan, W.-J. et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med 382, 1708–1720 (2020).
Al-Aly, Z., Bowe, B. & Xie, Y. Long COVID after breakthrough SARS-CoV-2 infection. Nat. Med. 28, 1461–1467 (2022).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Lau, R. I. et al. Gut microbiota in COVID-19: key microbial changes, potential mechanisms and clinical applications. Nat. Rev. Gastroenterol. Hepatol. 20, 323–337 (2023).
Miyauchi, E., Shimokawa, C., Steimle, A., Desai, M. S. & Ohno, H. The impact of the gut microbiome on extra-intestinal autoimmune diseases. Nat. Rev. Immunol. 23, 9–23 (2023).
Hou, K. et al. Microbiota in health and diseases. Signal Transduct. Target. Ther. 7, 135 (2022).
Liu, Q. et al. Multi-kingdom gut microbiota analyses define COVID-19 severity and post-acute COVID-19 syndrome. Nat. Commun. 13, 6806 (2022).
Arunachalam, P. S. et al. Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science 369, 1210–1220 (2020).
Li, X. V., Leonardi, I. & Iliev, I. D. Gut mycobiota in immunity and inflammatory disease. Immunity 50, 1365–1379 (2019).
Bacher, P. et al. Human anti-fungal Th17 immunity and pathology rely on cross-reactivity against Candida albicans. Cell 176, 1340–1355 (2019).
Leonardi, I. et al. CX3CR1+ mononuclear phagocytes control immunity to intestinal fungi. Science 359, 232–236 (2018).
Salmanton-García, J. et al. COVID-19-associated pulmonary aspergillosis, March–August 2020. Emerg. Infect. Dis. J. 27, 1077 (2021).
Zuo, T. et al. Alterations in fecal fungal microbiome of patients with COVID-19 during time of hospitalization until discharge. Gastroenterology 159, 1302–1310.e1305 (2020).
Lv, L. et al. Gut mycobiota alterations in patients with COVID-19 and H1N1 infections and their associations with clinical features. Commun. Biol. 4, 480 (2021).
Standaert–Vitse, A. et al. Candida albicans is an immunogen for anti Saccharomyces cerevisiae antibody markers of Crohn’s disease. Gastroenterology 130, 1764–1775 (2006).
Doron, I. et al. Human gut mycobiota tune immunity via CARD9-dependent induction of anti-fungal IgG antibodies. Cell 184, 1017–1031 (2021).
Wang, Z. Z., Shi, K. & Peng, J. Serologic testing of a panel of five antibodies in inflammatory bowel diseases: diagnostic value and correlation with disease phenotype. Biomed. Rep. 6, 401–410 (2017).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv113 (2012).
Koutsakos, M. et al. Integrated immune dynamics define correlates of COVID-19 severity and antibody responses. Cell Rep. Med. 2, 100208 (2021).
Hoenigl, M. et al. COVID-19-associated fungal infections. Nat. Microbiol. 7, 1127–1140 (2022).
Proctor, D. M. et al. Integrated genomic, epidemiologic investigation of Candida auris skin colonization in a skilled nursing facility. Nat. Med. 27, 1401–1409 (2021).
Leonardi, I. et al. Fungal trans-kingdom dynamics linked to responsiveness to fecal microbiota transplantation (FMT) therapy in ulcerative colitis. Cell Host Microbe 27, 823–829 (2020).
Zhai, B. et al. High-resolution mycobiota analysis reveals dynamic intestinal translocation preceding invasive candidiasis. Nat. Med. 26, 59–64 (2020).
Li, X. et al. Response to fungal dysbiosis by gut-resident CX3CR1+ mononuclear phagocytes aggravates allergic airway disease. Cell Host Microbe 24, 847–856 (2018).
Li, X. V. et al. Immune regulation by fungal strain diversity in inflammatory bowel disease. Nature 603, 672–678 (2022).
Silvin, A. et al. Elevated calprotectin and abnormal myeloid cell subsets discriminate severe from mild COVID-19. Cell 182, 1401–1418 (2020).
Rendeiro, A. F. et al. Profiling of immune dysfunction in COVID-19 patients allows early prediction of disease progression. Life Sci. Alliance 4, e202000955 (2021).
Mann, E. R. et al. Longitudinal immune profiling reveals key myeloid signatures associated with COVID-19. Sci. Immunol. 5, eabd6197 (2020).
Shao, T. Y. et al. Commensal Candida albicans positively calibrates systemic Th17 immunological responses. Cell Host Microbe 25, 404–417 (2019).
Al-Aly, Z., Xie, Y. & Bowe, B. High-dimensional characterization of post-acute sequelae of COVID-19. Nature 594, 259–264 (2021).
George, P. M. et al. A persistent neutrophil-associated immune signature characterizes post-COVID-19 pulmonary sequelae. Sci. Transl. Med. 14, eabo5795 (2022).
Cheong, J.-G. et al. Epigenetic memory of coronavirus infection in innate immune cells and their progenitors. Cell 186, 3882–3902 (2023).
Leonardi, I. et al. Mucosal fungi promote gut barrier function and social behavior via type 17 immunity. Cell 185, 831–846 (2022).
Rosas, I. O. et al. Tocilizumab in hospitalized patients with severe COVID-19 pneumonia. N. Engl. J. Med. 384, 1503–1516 (2021).
Zuo, Y. et al. Neutrophil extracellular traps in COVID-19. JCI Insight 5, e138999 (2020).
Dinnon, K. H. et al. A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature 586, 560–566 (2020).
Çavuş, M. A. & Sav, H. Opportunistic infections in critical COVID-19 patients. Pol. J. Microbiol. 71, 411–419 (2022).
Hoenigl, M. et al. The emergence of COVID-19 associated mucormycosis: a review of cases from 18 countries. Lancet Microbe 3, e543–e552 (2022).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Yeoh, Y. K. et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 70, 698–706 (2021).
Kayaaslan, B. et al. Incidence and risk factors for COVID-19 associated candidemia (CAC) in ICU patients. Mycoses 65, 508–516 (2022).
Giron, L. B. et al. Markers of fungal translocation are elevated during post-acute sequelae of SARS-CoV-2 and induce NF-κB signaling. JCI Insight 7, e160989 (2022).
Sun, Z. et al. Gut microbiome alterations and gut barrier dysfunction are associated with host immune homeostasis in COVID-19 patients. BMC Med. 20, 24 (2022).
Chen, Y.-H. et al. Rewilding of laboratory mice enhances granulopoiesis and immunity through intestinal fungal colonization. Sci. Immunol. 8, eadd6910 (2023).
McGonagle, D., Sharif, K., O’Regan, A. & Bridgewood, C. The role of cytokines including interleukin-6 in COVID-19 induced pneumonia and macrophage activation syndrome-like disease. Autoimmun. Rev. 19, 102537 (2020).
Veras, F. P. et al. SARS-CoV-2–triggered neutrophil extracellular traps mediate COVID-19 pathology. J. Exp. Med. 217, e20201129 (2020).
Wigerblad, G. & Kaplan, M. J. Neutrophil extracellular traps in systemic autoimmune and autoinflammatory diseases. Nat. Rev. Immunol. 23, 274–288 (2023).
Tanaka, T., Narazaki, M. & Kishimoto, T. IL-6 in inflammation, immunity, and disease. Cold. Spring Harb. Perspect. Biol. 6, a016295 (2014).
Beigel, J. H. et al. Remdesivir for the treatment of COVID-19—final report. N. Engl. J. Med. 383, 1813–1826 (2020).
Doron, I. et al. Mycobiota-induced IgA antibodies regulate fungal commensalism in the gut and are dysregulated in Crohn’s disease. Nat. Microbiol. 6, 1493–1504 (2021).
Yeung, S. T., Ovando, L. J., Russo, A. J., Rathinam, V. A. & Khanna, K. M. CD169+ macrophage intrinsic IL-10 production regulates immune homeostasis during sepsis. Cell Rep. 42, 112171 (2023).
Ural, B. B. et al. Identification of a nerve-associated, lung-resident interstitial macrophage subset with distinct localization and immunoregulatory properties. Sci. Immunol. 5, eaax8756 (2020).
Tang, J., Iliev, I. D., Brown, J., Underhill, D. M. & Funari, V. A. Mycobiome: approaches to analysis of intestinal fungi. J. Immunol. Methods 421, 112–121 (2015).
Acknowledgements
We thank members of the Iliev Laboratory for their critical reviews of the manuscript. We thank all contributing members of the Department of Pathology and Laboratory Medicine, of the JRI IBD Live Cell Bank Consortium and the Microbiome Core Laboratory of Weill Cornell Medicine, the NYU Langone Health Microscopy Laboratory (NCI P30CA016087), Khanna Laboratory (supported by R01AI143861 and R01AI143861-02S1) and R. Albrecht, R. Cadogan, D. Flores for support with the BSL3 facility and procedures. The authors were supported by WCM WCG COVID-19; WCGS Merit Fellowship (to W.-Y.L); Asan foundation, ROK (to J.-G.C.); K08MH130773 (to C.N.P.); R01AI148416, R01AI148416-S1, R01AI148416-S2, Burrough Welcome Trust PATH Award, and the Hirschl Weill-Caulier Award (to S.Z.J); R01AI160706 and R01DK130425 (to M. Schotsaert); CRIPT, CEIRR (contract # 75N93021C00014) U19AI135972, U19AI168631 and U19AI142733 (to A.G.-S.). Research in the Iliev Laboratory is supported by the US National Institutes of Health (R01DK113136, R01DK121977 and R01AI137157), the Leona M. and Harry B. Helmsley Charitable Trust, the Irma T. Hirschl Career Scientist Award, the Research Corporation for Science Advancement Award, the Cancer Research Institute Lloyd J. Old STAR Award, and the Burrough Welcome Trust PATH Award. I.D.I. is a fellow of the CIFAR program Fungal Kingdom—Threats and Opportunities.
Author information
Authors and Affiliations
Contributions
T.K. and I.D.I. conceived and conceptualized the study. T.K., G.S., J.C., A.R., S.T.Y. and M. Schotsaert designed experiments and analyzed data. M. Salvatore was involved in the investigation, clinical samples and clinical data acquisition. T.K., W.-Y.L., J.C., G.S., G.C, S.T.Y., C.J.G., M.M., M.B.D., I.D. and G.G.P. performed experiments and acquired and analyzed data. S.R., M.C., L.W., C.N.P., Z.Z. and G.I. provided clinical samples and participated in clinical data acquisition. S.W., L.N. contributed to experimental interpretation. I.D.I., M. Salvatore, A.G.-S. and S.Z.J. supervised the study. T.K., J.C., W.-Y.L. and I.D.I. generated figures and legends from analyzed data. I.D.I. administered the project and acquired funding. T.K. and I.D.I. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The AGS Laboratory has received research support from GSK, Pfizer, Senhwa Biosciences, Kenall Manufacturing, Blade Therapeutics, Avimex, Johnson & Johnson, Dynavax, 7 Hills Pharma, Pharmamar, ImmunityBio, Accurius, Nanocomposix, Hexamer, N-fold LLC, Model Medicines, Atea Pharma, Applied Biological Laboratories and Merck, outside of the reported work. A.G.-S. has consulting agreements for the following companies involving cash and/or stock: Castlevax, Amovir, Vivaldi Biosciences, Contrafect, 7 Hills Pharma, Avimex, Pagoda, Accurius, Esperovax, Farmak, Applied Biological Laboratories, Pharmamar, CureLab Oncology, CureLab Veterinary, Synairgen, Paratus, Pfizer and Prosetta, outside of the reported work. A.G.-S. has been an invited speaker in meeting events organized by Seqirus, Janssen, Abbott and AstraZeneca; and is inventor on patents and patent applications on the use of antivirals and vaccines for the treatment and prevention of virus infections and cancer, owned by the Icahn School of Medicine at Mount Sinai, New York, outside of the reported work. M. Schotsaert has received unrelated research funding in sponsored research agreements from ArgenX BV, Moderna, 7 Hills Pharma and Phio Pharmaceuticals, which has no competing interest with this work. The other authors declare no competing interests related to this study.
Peer review
Peer review information
Nature Immunology thanks Gordon Brown and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ioana Visan, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Serological testing in Cohorts 1 and 2.
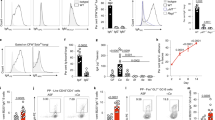
a-b, SARS-CoV-2 RBD IgG titration curves in HD (n = 36, a) and sCOVID-19 (n = 66, b). c, Plasma IgG antibody titers to SARS-CoV-2 RBD in HD (n = 36), mCOVID-19 (n = 25) and sCOVID-19 (n = 66) (Extended Data Table 1). The data are shown as endpoint titers normalized to ELISA reciprocal dilution as in a and b. The dotted line indicates the limit of detection. In the boxplots, the center is drawn through the median of the measurement, and the lower and upper bounds of the box correspond to the first and third quartile. The whiskers go down to the smallest value and up to the largest. Statistical significance was determined by the one-way ANOVA followed by Tukey’s multiple-comparison. Related to Fig. 1.
Extended Data Fig. 2 Compositional analysis of gut mycobiota.
a-b, Ratio between Ascomycota and Basidiomycota (a) and relative abundance of fungal species (b) in ITS1 sequencing of fungal rDNA from stool samples of HD (n = 10) and COVID-19 patients (n = 10). Lower and upper hinges correspond to the first and third quartile; dots represent individual patients’ samples. P values were calculated using a two-tailed Mann-Whitney testing between all groups. Related to Fig. 2.
Extended Data Fig. 3 Immune responses to C.albicans isolates from COVID-19 patients.
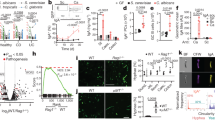
a, Representative graphs depicting the flow cytometry gating strategy for defining murine neutrophil populations in tissues. b-d, Anti-C. albicans specific IgG titers (b), the amount of neutrophils in peripheral blood (c) and lung (d) in antibiotic-treated mice orally gavaged or not (PBS, n = 7) with either C. glabrata (CgCOV3, n = 7) or C. albicans (CaCOV1, CaCOV5, CaCOV2, n = 7) isolated from COVID-19 patient’s stool. Immune responses were assessed at 2 weeks after colonization. In boxplots in b, the center is drawn through the median of the measurement, and the lower and upper bounds of the box correspond to the first and third quartile. The whiskers go down to the smallest value and up to the largest. The bar graphs in c and d presented as mean ± SEM. The results were pooled from two experiments. P values were calculated using the one-way ANOVA followed by Tukey’s multiple comparison. ns = not significant. Related to Figs. 2 and 3.
Extended Data Fig. 4 Linear regression analysis of the Immune cell frequencies and levels of ACAL IgG in peripheral blood of mCOVID-19 and sCOVID-19.
a-e, Comparison of linear regression analysis of CD3+ T cell (a), CD19+CD20+ B cell (b), NK cell (c), CD4+ T cell (d) and CD8+ T cell (e) frequencies and levels of ACAL IgG in peripheral blood of sCOVID-19 (n = 13) and mCOVID-19 (n = 17), see Extended Data Table 1. Crosses indicate deceased patients. Red and black lines show linear fits within severe and low to moderate groups, respectively, with 95% confidence intervals shown in gray. Spearman correlation estimates (rho) and associated p values are shown in red and black for sCOVID-19 and mCOVID19, respectively. Related to Fig. 3.
Extended Data Fig. 5 Strong correlation between ACAL IgG and GMPs among multiple progenitor cell types.
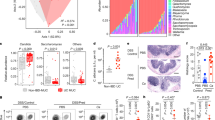
Correlation and linear regression of frequency of hematopoietic stem cells/multipotent progenitor cells (HSC/MPP), lymphoid-primed multipotent progenitor cells (LMPP), granulocyte-macrophage progenitor cells (GMP), erythroid progenitor cells (Ery), megakaryocyte-erythroid progenitor cells (MEP) with ACAL IgG (log10 titer) in enrichment of CD34+ HSPC from PBMC of HD(n = 5), sCOVID-19 (n = 12) and nonCOV-19 (n = 5), see Extended Data Table 2. Blue line indicates the regression line for all patients. The associated linear regression equation, Pearson’s correlation. Coefficient and significance are shown. Related to Fig. 4.
Extended Data Fig. 6 HLA-related transcriptional signatures of GMPs form HD, sCOVID19 and sCOVID19 treated with tocilizumab.
a, Heatmap showing antigen presentation marker that are differentially expressed in GMP of HD (n = 7) and sCOVID-19 who received (n = 6) or not (n = 7) IL-6R blockade treatment. Data are average of normalized expression for each gene in each group. b, Antibiotic treated mice were colonized with C. albicans CaCOV5 by oral gavage twice for two weeks prior to SARS-COV-2 challenge and were harvested at Day 20. Water with or without antifungal fluconazole was provided to mice two days after second oral gavage. Related to Fig. 5.
Supplementary information
Source data
Source Data Fig. 1
Raw data.
Source Data Fig. 2
Raw data.
Source Data Fig. 3
Raw data.
Source Data Fig. 4
Raw data.
Source Data Fig. 5
Raw data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kusakabe, T., Lin, WY., Cheong, JG. et al. Fungal microbiota sustains lasting immune activation of neutrophils and their progenitors in severe COVID-19. Nat Immunol 24, 1879–1889 (2023). https://doi.org/10.1038/s41590-023-01637-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01637-4
This article is cited by
-
Fungi in cancer
Nature Reviews Cancer (2024)
-
Gut fungi have lasting effect on immune response to severe COVID-19
Nature Reviews Immunology (2023)
-
Candida makes a lasting impression in COVID-19
Nature Immunology (2023)
-
Inflammation in severe COVID linked to bad fungal microbiome
Nature (2023)