Abstract
Invariant natural killer T (iNKT) cells recognize activating self and microbial lipids presented by CD1d. CD1d can also bind non-activating lipids, such as sphingomyelin. We hypothesized that these serve as endogenous regulators and investigated humans and mice deficient in acid sphingomyelinase (ASM), an enzyme that degrades sphingomyelin. We show that ASM absence in mice leads to diminished CD1d-restricted antigen presentation and iNKT cell selection in the thymus, resulting in decreased iNKT cell levels and resistance to iNKT cell-mediated inflammatory conditions. Defective antigen presentation and decreased iNKT cells are also observed in ASM-deficient humans with Niemann–Pick disease, and ASM activity in healthy humans correlates with iNKT cell phenotype. Pharmacological ASM administration facilitates antigen presentation and restores the levels of iNKT cells in ASM-deficient mice. Together, these results demonstrate that control of non-agonistic CD1d-associated lipids is critical for iNKT cell development and function in vivo and represents a tight link between cellular sphingolipid metabolism and immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The crystal structure is available at https://www.rcsb.org, PDB structure ID: 6CYW. The rest of the data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Brigl, M. & Brenner, M. B. CD1: antigen presentation and T cell function. Annu. Rev. Immunol. 22, 817–890 (2004).
Brennan, P. J., Brigl, M. & Brenner, M. B. Invariant natural killer T cells: an innate activation scheme linked to diverse effector functions. Nat. Rev. Immunol. 13, 101–117 (2013).
Godfrey, D. I. & Berzins, S. P. Control points in NKT cell development. Nat. Rev. Immunol. 7, 505–518 (2007).
Godfrey, D. I., Stankovic, S. & Baxter, A. G. Raising the NKT cell family. Nat. Immunol. 11, 197–206 (2010).
Gapin, L., Godfrey, D. I. & Rossjohn, J. Natural killer T cell obsession with self-antigens. Curr. Opin. Immunol. 25, 168–173 (2013).
Merrill, A. H. Sphingolipid and glycosphingolipid metabolic pathways in the era of sphingolipidomics. Chem. Rev. 111, 6387–6422 (2011).
Yuan, W., Kang, S., Evans, J. & Cresswell, P. Natural lipid ligands associated with human CD1d targeted to different subcellular compartments. J. Immunol. 182, 4784–4791 (2009).
Salio, M., Silk, J. D., Jones, Y. E. & Cerundolo, V. Biology of CD1- and MR1-restricted T cells. Annu. Rev. Immunol. 32, 323–366 (2014).
Fox, L. M. et al. Recognition of lyso-phospholipids by human natural killer T lymphocytes. PLoS Biol. 7, e1000228 (2009).
Smith, E. & Schuchman, E. The unexpected role of acid sphingomyelinase in cell death and the pathophysiology of common diseases. FASEB J. 22, 3419–3431 (2008).
Im, J. S. et al. Kinetics and cellular site of glycolipid loading control the outcome of natural killer T cell activation. Immunity 30, 888–898 (2009).
Perrotta, C. & Clementi, E. Biological roles of acid and neutral sphingomyelinases and their regulation by nitric oxide. Physiology 25, 64–71 (2010).
Zeidan, Y. H. & Hannun, Y. A. The acid sphingomyelinase/ceramide pathway: biomedical significance and mechanisms of regulation. Curr. Mol. Med. 10, 454–466 (2009).
Horinouchi, K. et al. Acid sphingomyelinase deficient mice: a model of types A and B Niemann–Pick disease. Nat. Genet. 10, 288–293 (1995).
Truman, J.-P., Gadban, M. M., Smith, K. J. & Hammad, S. M. Acid sphingomyelinase in macrophage biology. Cell Mol. Life Sci. 68, 3293–3305 (2011).
Yang, O. O. et al. CD1d on myeloid dendritic cells stimulates cytokine secretion from and cytolytic activity of Vα24JαQ T cells: a feedback mechanism for immune regulation. J. Immunol. 165, 3756–3762 (2000).
Nieuwenhuis, E. E. et al. CD1d and CD1d-restricted iNKT-cells play a pivotal role in contact hypersensitivity. Exp. Dermatol. 14, 250–258 (2005).
Takeda, K. et al. Critical contribution of liver natural killer T cells to a murine model of hepatitis. Proc. Natl Acad. Sci. USA 97, 5498–5503 (2000).
Cernadas, M. et al. Lysosomal localization of murine CD1d mediated by AP-3 is necessary for NK T cell development. J. Immunol. 171, 4149–4155 (2003).
Chiu, Y.-H. et al. Multiple defects in antigen presentation and T cell development by mice expressing cytoplasmic tail-truncated CD1d. Nat. Immunol. 3, 55–60 (2001).
Cui, J. et al. Requirement for Vα14 NKT cells in IL-12-mediated rejection of tumors. Science 278, 1623–1626 (1997).
Behar, S., Podrebarac, T. A., Roy, C., Wang, C. & Brenner, M. Diverse TCRs recognize murine CD1. J. Immunol. 162, 161–167 (1999).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA‐based α‐ and β‐chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Thedrez, A. et al. CD4 engagement by CD1d potentiates activation of CD4+ invariant NKT cells. Blood 110, 251–258 (2007).
Zeissig, S. et al. Primary deficiency of microsomal triglyceride transfer protein in human abetalipoproteinemia is associated with loss of CD1 function. J. Clin. Invest. 120, 2889–2899 (2010).
McNab, F. et al. The influence of CD1d in postselection NKT cell maturation and homeostasis. J. Immunol. 175, 3762–3768 (2005).
Balreira, A., Lacerda, L., Miranda, C. & Arosa, F. A. Evidence for a link between sphingolipid metabolism and expression of CD1d and MHC class II: monocytes from Gaucher disease patients as a model. Brit. J. Haematol. 129, 667–676 (2005).
Pereira, C. S. et al. Invariant natural killer T cells are phenotypically and functionally altered in Fabry disease. Mol. Genet. Metab. 108, 241–248 (2013).
Speak, A. O. et al. Invariant natural killer T cells are not affected by lysosomal storage in patients with Niemann–Pick disease type C. Eur. J. Immunol. 42, 1886–1892 (2012).
Salinas, F., Smith, L. & Goodman, J. Cell size distribution in the thymus as a function of age. J. Cell Physiol. 80, 339–345 (1972).
Chiu, Y.-H. et al. Distinct subsets of CD1d-restricted T cells recognize self-antigens loaded in different cellular compartments. J. Exp. Med. 189, 103–110 (1999).
Jahng, A. et al. Prevention of autoimmunity by targeting a distinct, noninvariant CD1d-reactive T cell population reactive to sulfatide. J. Exp. Med. 199, 947–957 (2004).
Girardi, E. et al. Type II natural killer T cells use features of both innate-like and conventional T cells to recognize sulfatide self antigens. Nat. Immunol. 13, 851 (2012).
Zajonc, D. M. et al. Structure and function of a potent agonist for the semi-invariant natural killer T cell receptor. Nat. Immunol. 6, 810–818 (2005).
Zajonc, D. M. et al. Structural basis for CD1d presentation of a sulfatide derived from myelin and its implications for autoimmunity. J. Exp. Med. 202, 1517–1526 (2005).
Sette, A. et al. Peptide binding to the most frequent HLA-A class I alleles measured by quantitative molecular binding assays. Mol. Immunol. 31, 813–822 (1994).
He, X. et al. Characterization of human acid sphingomyelinase purified from the media of overexpressing Chinese hamster ovary cells. Biochim. Biophys. Acta Prot. Struct. Mol. Enzymol. 1432, 251–264 (1999).
Miranda, S. et al. Infusion of recombinant human acid sphingomyelinase into Niemann–Pick disease mice leads to visceral, but not neurological, correction of the pathophysiology. FASEB J. 14, 1988–1995 (2000).
Bendelac, A., Savage, P. B. & Teyton, L. The biology of NKT cells. Annu. Rev. Immunol. 25, 297–336 (2007).
Cox, D. et al. Determination of cellular lipids bound to human CD1d molecules. PLoS ONE 4, e5325 (2009).
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
NP-C Guidelines Working Group. Recommendations on the diagnosis and management of Niemann–Pick disease type C. Mol. Genet. Metab. 98, 152–165 (2009).
Nieuwenhuis, E. et al. CD1d-dependent macrophage-mediated clearance of Pseudomonas aeruginosa from lung. Nat. Med. 8, 588 (2002).
Kinjo, Y. et al. Invariant natural killer T cells recognize glycolipids from pathogenic Gram-positive bacteria. Nat. Immunol. 12, 966–974 (2011).
McGovern, M. M. et al. Morbidity and mortality in type B Niemann–Pick disease. Genet. Med. 15, 618 (2013).
Platt, F. M. Sphingolipid lysosomal storage disorders. Nature 510, 68–75 (2014).
Fritsch, J., Tchikov, V., Hennig, L., Lucius, R. & Schütze, S. A toolbox for the immunomagnetic purification of signaling organelles. Traffic 20, 246–258 (2019).
Shaner, R. L. et al. Quantitative analysis of sphingolipids for lipidomics using triple quadrupole and quadrupole linear ion trap mass spectrometers. J. Lipid Res. 50, 1692–1707 (2009).
Zeissig, S., Olszak, T., Melum, E. & Blumberg, R. S. Analyzing antigen recognition by natural killer T cells. Methods Mol. Biol. 960, 557–572 (2013).
Yu, K. et al. Production and characterization of monoclonal antibodies against complexes of the NKT cell ligand α-galactosylceramide bound to mouse CD1d. J. Immunol. Methods 323, 11–23 (2007).
Sommer, C., Straehle, C., Kothe, U. & Hamprecht, F. A. ILASTIK: interactive learning and segmentation toolkit. Proc. 8th IEEE International Symposium on Biomedical Imaging 1, 230–233 (2011).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
Wang, J. et al. Lipid binding orientation within CD1d affects recognition of Borrelia burgorferi antigens by NKT cells. Proc. Natl Acad. Sci. USA 107, 1535–1540 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–26 (1997).
McCoy, A. J., Grosse-Kunstleve, R. W., Storoni, L. C. & Read, R. J. Likelihood-enhanced fast translation functions. Acta Crystallogr. Sect. D. Biol. Crystallogr. 61, 458–464 (2005).
Emsley, P., Lohkamp, B., Scott, W. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D. Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D. Biol. Crystallogr. 53, 240–255 (1997).
Schüttelkopf, A. W. & van Aalten, D. M. F. PRODRG: a tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr D. Biol. Crystallogr 60, 1355–1363 (2004).
D’Agostino, R. & Pearson, E. Tests for departure from normality. Empirical results for the distributions of b 2 and √b 1. Biometrika 60, 613 (1973).
Acknowledgements
This work was supported by US National Institutes of Health grants HD28607 (MERIT Award) (to E.H.S.), DK044319, DK051362, DK053056, DK088199 and the Harvard Digestive Diseases Center DK034854 (to R.S.B.), the Southeastern Norwegian Health Trust, the Unger–Vetlesen Foundation, the Caroline Musæus Aarvolds fund and the Norwegian PSC Research Center (to E.M.), the European Research Council (ERC Starting Grant agreement no. 336528), the Deutsche Forschungsgemeinschaft (DFG) (ZE 814/4-1, ZE 814/7-1) and the DFG Excellence Cluster Center for Regenerative Therapies Dresden (to S.Z.), the DFG (SCHU733/14-1) (to S.S. and J.F.) and the DFG Excellence Cluster Inflammation at Interfaces Schleswig-Holstein (EXC 306) (to S.S.). This work was partially funded by FEDER funds under the Portugal 2020 partnership agreement through the Norte Portugal Regional Operational Program (Norte 2020) (Norte-01-0145-FEDER-000012). J. Bame, J. Danielson, A. Dias, M.L. Maia, S. Torquato, J. Øgaard and J. Anmarkrud are thanked for invaluable technical help. We thank the following physicians for patient and control subject recruitment in Portugal: T. Cardoso and N. Alegrete (CHS João, Porto), E. Martins and E. Silva (CH Porto, Porto), and L. Ribeiro and A. Pereira (CHU Coimbra, Coimbra). We acknowledge the blood bank of CHS João, Porto, and the National Institutes of Health tetramer core facility for provision of PBS57-loaded CD1d tetramer and CD1d monomer.
Author information
Authors and Affiliations
Contributions
E.M. designed, performed and analyzed experiments with R.S.B. E.M., S.Z. and R.S.B. wrote the manuscript. X.J., K.D.B., C.M.D., A.P. and C.T. helped with experiments. M.F.M. and C.S.P. provided and analyzed human samples for NKT cells. S.Z. provided lentiviruses expressing CD1d and contributed to the design of experiments and the interpretation of results. J.F. and S.S. performed extraction of lysosomes. J.W. and D.M.Z. performed the IEF experiments and determined the CD1d–sphingomyelin structure. A.K., T.H.K. and M.A.E. provided scientific input. S.L.K., J.D. and A.H.M. performed mass spectometry and analyzed the lipidomics data together with E.M. E.H.S. provided Asm–/– mice and rhASM and assisted in the analysis of experiments. R.S.B. supervised the studies.
Corresponding authors
Ethics declarations
Competing interests
E.H.S. is a consultant for Sanofi Genzyme, and M.F.M. has received a research grant from Sanofi Genzyme.
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Examples of gating strategies for thymic samples.
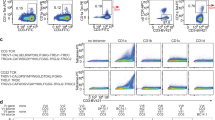
Thymocytes were prepared from the thymus and stained with monoclonal antibodies. The figure demonstrates representative dot plots and gating strategies for iNKT cells, iNKT cell stages, tetramer-negative thymocytes and CD4/CD8 distribution. Other samples analyzed in the project were gated in a similar manner. SP single positive; DN double negative; DP double positive.
Supplementary Figure 2 Acid sphingomyelinase deficient mice have reduced numbers of iNKT cells with an altered CD4/CD8 distribution.
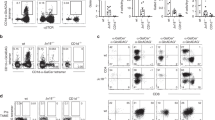
Lymphocytes were prepared from the tissues indicated in the figure followed by staining with monoclonal antibodies and a CD1d tetramer. (a) Representative flow cytometry dot plots of lymphocytes from WT and Asm–/– mice visualizing the frequency of iNKT cells in thymus and liver as defined by a PBS57-loaded CD1d tetramer and CD3. The results are representative of three independent experiments. (b) Distribution of CD4 and CD8 expression among iNKT cells from Asm–/–(n=5) and WT (n=5) mice. The results are representative of two independent experiments. In all panels the mean values are shown with the error bars representing the SEM. P-values were calculated by two-sided t-test. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001, SP single positive; DN double negative; DP double positive.
Supplementary Figure 3 T cell distribution, CD4/CD8 expression, T regulatory cells and γδ T cells in Asm–/– compared to wild-type mice.
Lymphocytes were prepared from the indicated tissues from Asm–/– (n=5) and WT (n=5) mice and stained with monoclonal antibodies. (a) The percentages indicate CD3 positive cells among PBS57-CD1d tetramer-negative lymphocytes. The results are representative of two independent experiments. (b) The percentages indicate the distribution of CD4 and CD8 expression among PBS57-CD1d tetramer-negative thymocytes (for thymus) and PBS57-CD1d tetramer-negative T cells (for spleen and liver). The results are representative of two independent experiments. (c) The percentages indicate the percentage of T regulatory cells or γδ T cells among T cells or lymphocytes excluding iNKT cells. The results are representative of two independent experiments. In all panels the mean values are shown with the error bars representing the SEM. P-values were calculated by two-sided t-test. *P<0.05, **P<0.01, SP single positive; DN double negative; DP double positive.
Supplementary Figure 4 CD1d expression in thymocytes and CD11c+ DCs from Asm–/– and wild-type mice.
Thymocytes were prepared by manual maceration through a mesh and thereafter stained with a monoclonal antibody against CD1d. CD11c+ DCs were extracted from spleens with CD11c magnetic beads and stained with a monoclonal antibody against CD1d. The two top histograms demonstrate the CD1d levels in thymocytes from young (left) and adult (right) mice. The two lower histograms demonstrate the CD1d levels in DCs from young (left) and adult (right) mice. The results are representative of three independent experiments. MFI median fluorescence intensity.
Supplementary Figure 5 Activation of NKT cells by thymocytes and DCs is blocked by a CD1d antibody and bone marrow transplantation in Asm–/– and wild-type mice.
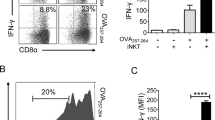
(a) Thymocytes were prepared by manual maceration through a mesh and loaded with α-GalCer for 4 hours (for 24.7 and DN32.D3 α-GC loaded) or left untreated (for 24.7 endogenous) followed by culture with the indicated iNKT hybridomas for 16–24h. Cytokine levels were measured in the culture supernatants from three independent wells. The results are representative of two independent experiments. (b) Bone marrow transplantation in Asm–/– (n=5) and WT mice (n=3). Bone marrow chimeras were made by irradiating Asm–/– and WT mice followed by injection of donor bone marrow from Asm–/– or WT mice. 3 months later the mice were sacrificed followed by flow-cytometry of tissue samples. The graphs demonstrate the percentage of CD1d-PBS57 tetramer positive cells among TCRβ positive cells (iNKT cells) from the indicated tissues. (c) CD11c+ DCs were extracted from spleens with CD11c magnetic beads and loaded with α-GalCer for 4 hours (upper panel) or left untreated (lower panel) followed by co-culture with the indicated NKT hybridomas for 16–24h. Cytokine levels were measured in the culture supernatants from three independent wells. The results are representative of two independent experiments. In all panels the mean values are shown with the error bars representing the SEM. P-values were calculated by two-sided t-test. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
Supplementary Figure 6 Transduction efficacy in transduced EBV-transformed B cells and phenotype of iNKT cells and T cells in Niemann-Pick disease patients compared to controls.
(a) EBV-transformed B cells from three healthy controls and one Niemann-Pick disease (NPD) patient were lentivirally transduced with a construct encoding human CD1d. The cells were thereafter subjected to FACS sorting. The histograms indicate the CD1d expression in the transduced and sorted cells (red lines) compared with the corresponding untransduced cells (black line). (b,c) Phenotype of iNKT cells and T cells in NPD patients compared to controls. Lymphocytes from five NPD patients (four type B and one type A) and 70 healthy controls were investigated with flow-cytometry. (b) CD4, CD8, double negative (DN) and CD161 distribution in iNKT cells. (c) CD4 and CD8 distribution in T cells. The line indicates the median. A two-sided Mann–Whitney test was used for significance testing. *P<0.05, **P<0.001 ***P<0.0001. NPD Niemann-Pick disease; DN double negative.
Supplementary Figure 7 Lipidomics results from liver samples from 2 week old Asm–/– and wild-type mice.
Sphingomyelin and ceramide levels in the liver of 2 week old Asm–/– (n=2) and wild-type (n=2) mice were quantified by mass spectrometry. (a) The graphs show the mean levels of sphingomyelins and DH-sphingomyelins with carbon chains of different lengths. (b) The graphs show the mean levels of ceramides, DH-ceramides,hexosylceramides and DH-hexosylceramides with carbon chains of different lengths.
Supplementary Figure 8 Sphingomyelin does not affect direct activation of iNKT cells and acid sphingomyelinase does not directly affect loading of lipid antigens.
(a) Cell culture plates were first coated with CD3 followed by incubation with the indicated concentrations of sphingomyelin C24:1. After thorough washing the DN32.D3 hybridoma was added and incubated for 16 hours. IL-2 levels in the supernatant were determined by ELISA. The results are representative of two independent experiments. (b) Murine CD1d was coated on cell culture plates followed by incubation with α-GalCer and Saposin-B, rhASM or BSA in the indicated concentrations overnight. Thereafter, the DN32.D3 hybridoma was added for 16 hours. IL-2 levels in the supernatant were determined by ELISA. The left panel shows the results when the experiment was performed under acidic conditions while the right panel show the results under neutral conditions. The results are representative of two independent experiments. In all panels the mean values are shown with the error bars representing the SEM.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Melum, E., Jiang, X., Baker, K.D. et al. Control of CD1d-restricted antigen presentation and inflammation by sphingomyelin. Nat Immunol 20, 1644–1655 (2019). https://doi.org/10.1038/s41590-019-0504-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0504-0
This article is cited by
-
Tryptophan metabolite norharman secreted by cultivated Lactobacillus attenuates acute pancreatitis as an antagonist of histone deacetylases
BMC Medicine (2023)
-
Imprinting of the immune system by the microbiota early in life
Mucosal Immunology (2020)
-
What one lipid giveth, another taketh away
Nature Immunology (2019)