Abstract
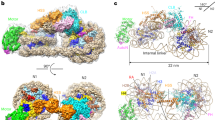
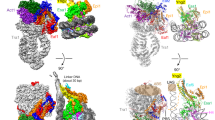
Access to DNA within nucleosomes is required for a variety of processes in cells including transcription, replication and repair. Consequently, cells encode multiple systems that remodel nucleosomes. These complexes can be simple, involving one or a few protein subunits, or more complicated multi-subunit machines1. Biochemical studies2,3,4 have placed the motor domains of several chromatin remodellers in the superhelical location 2 region of the nucleosome. Structural studies of yeast Chd1 and Snf2—a subunit in the complex with the capacity to remodel the structure of chromatin (RSC)—in complex with nucleosomes5,6,7 have provided insights into the basic mechanism of nucleosome sliding performed by these complexes. However, how larger, multi-subunit remodelling complexes such as INO80 interact with nucleosomes and how remodellers carry out functions such as nucleosome sliding8, histone exchange9 and nucleosome spacing10,11,12 remain poorly understood. Although some remodellers work as monomers13, others work as highly cooperative dimers11, 14, 15. Here we present the structure of the human INO80 chromatin remodeller with a bound nucleosome, which reveals that INO80 interacts with nucleosomes in a previously undescribed manner: the motor domains are located on the DNA at the entry point to the nucleosome, rather than at superhelical location 2. The ARP5–IES6 module of INO80 makes additional contacts on the opposite side of the nucleosome. This arrangement enables the histone H3 tails of the nucleosome to have a role in the regulation of the activities of the INO80 motor domain—unlike in other characterized remodellers, for which H4 tails have been shown to regulate the motor domains.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Dechassa, M. L. et al. Architecture of the SWI/SNF-nucleosome complex. Mol. Cell. Biol. 28, 6010–6021 (2008).
Brahma, S. et al. INO80 exchanges H2A.Z for H2A by translocating on DNA proximal to histone dimers. Nat. Commun. 8, 15616 (2017).
Nodelman, I. M. et al. Interdomain communication of the Chd1 chromatin remodeler across the DNA gyres of the nucleosome. Mol. Cell 65, 447–459 (2017).
Farnung, L., Vos, S. M., Wigge, C. & Cramer, P. Nucleosome–Chd1 structure and implications for chromatin remodelling. Nature 550, 539–542 (2017).
Liu, X., Li, M., Xia, X., Li, X. & Chen, Z. Mechanism of chromatin remodelling revealed by the Snf2–nucleosome structure. Nature 544, 440–445 (2017).
Sundaramoorthy, R. et al. Structural reorganization of the chromatin remodeling enzyme Chd1 upon engagement with nucleosomes. eLife 6, e22510 (2017).
Shen, X., Mizuguchi, G., Hamiche, A. & Wu, C. A chromatin remodelling complex involved in transcription and DNA processing. Nature 406, 541–544 (2000).
Papamichos-Chronakis, M., Watanabe, S., Rando, O. J. & Peterson, C. L. Global regulation of H2A.Z localization by the INO80 chromatin-remodeling enzyme is essential for genome integrity. Cell 144, 200–213 (2011).
Willhoft, O., Bythell-Douglas, R., McCormack, E. A. & Wigley, D. B. Synergy and antagonism in regulation of recombinant human INO80 chromatin remodeling complex. Nucleic Acids Res. 44, 8179–8188 (2016).
Leonard, J. D. & Narlikar, G. J. A nucleotide-driven switch regulates flanking DNA length sensing by a dimeric chromatin remodeler. Mol. Cell 57, 850–859 (2015).
Udugama, M., Sabri, A. & Bartholomew, B. The INO80 ATP-dependent chromatin remodeling complex is a nucleosome spacing factor. Mol. Cell. Biol. 31, 662–673 (2011).
Qiu, Y. et al. The Chd1 chromatin remodeler shifts nucleosomal DNA bidirectionally as a monomer. Mol. Cell 68, 76–88e76 (2017).
Willhoft, O. et al. Crosstalk within a functional INO80 complex dimer regulates nucleosome sliding. eLife 6, e25782 (2017).
Racki, L. R. et al. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature 462, 1016–1021 (2009).
Aramayo, R. J. et al. Cryo-EM structures of the human INO80 chromatin-remodeling complex. Nat. Struct. Mol. Biol. 25, 37–44 (2018).
Tosi, A. et al. Structure and subunit topology of the INO80 chromatin remodeler and its nucleosome complex. Cell 154, 1207–1219 (2013).
Chen, L., Conaway, R. C. & Conaway, J. W. Multiple modes of regulation of the human Ino80 SNF2 ATPase by subunits of the INO80 chromatin-remodeling complex. Proc. Natl Acad. Sci. USA 110, 20497–20502 (2013).
Watanabe, S. et al. Structural analyses of the chromatin remodelling enzymes INO80-C and SWR-C. Nat. Commun. 6, 7108 (2015).
Yao, W. et al. Assembly of the Arp5 (actin-related protein) subunit involved in distinct INO80 chromatin remodeling activities. J. Biol. Chem. 290, 25700–25709 (2015).
Gu, M. & Rice, C. M. Three conformational snapshots of the hepatitis C virus NS3 helicase reveal a ratchet translocation mechanism. Proc. Natl Acad. Sci. USA 107, 521–528 (2010).
Saravanan, M. et al. Interactions between the nucleosome histone core and Arp8 in the INO80 chromatin remodeling complex. Proc. Natl Acad. Sci. USA 109, 20883–20888 (2012).
Gerhold, C. B. et al. Structure of actin-related protein 8 and its contribution to nucleosome binding. Nucleic Acids Res. 40, 11036–11046 (2012).
Stockdale, C., Flaus, A., Ferreira, H. & Owen-Hughes, T. Analysis of nucleosome repositioning by yeast ISWI and Chd1 chromatin remodeling complexes. J. Biol. Chem. 281, 16279–16288 (2006).
Saha, A., Wittmeyer, J. & Cairns, B. R. Chromatin remodeling through directional DNA translocation from an internal nucleosomal site. Nat. Struct. Mol. Biol. 12, 747–755 (2005).
Ranjan, A. et al. H2A histone-fold and DNA elements in nucleosome activate SWR1-mediated H2A.Z replacement in budding yeast. eLife 4, e06845 (2015).
Mizuguchi, G. et al. ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 303, 343–348 (2004).
Ranjan, A. et al. Nucleosome-free region dominates histone acetylation in targeting SWR1 to promoters for H2A.Z replacement. Cell 154, 1232–1245 (2013).
Singleton, M. R., Dillingham, M. S. & Wigley, D. B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007).
Blosser, T. R., Yang, J. G., Stone, M. D., Narlikar, G. J. & Zhuang, X. Dynamics of nucleosome remodelling by individual ACF complexes. Nature 462, 1022–1027 (2009).
Deindl, S. et al. ISWI remodelers slide nucleosomes with coordinated multi-base-pair entry steps and single-base-pair exit steps. Cell 152, 442–452 (2013).
Krietenstein, N. et al. Genomic nucleosome organization reconstituted with pure proteins. Cell 167, 709–721.e712 (2016).
Clapier, C. R., Längst, G., Corona, D. F., Becker, P. B. & Nightingale, K. P. Critical role for the histone H4 N terminus in nucleosome remodeling by ISWI. Mol. Cell. Biol. 21, 875–883 (2001).
Clapier, C. R. & Cairns, B. R. Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes. Nature 492, 280–284 (2012).
Dang, W., Kagalwala, M. N. & Bartholomew, B. Regulation of ISW2 by concerted action of histone H4 tail and extranucleosomal DNA. Mol. Cell. Biol. 26, 7388–7396 (2006).
Yan, L., Wang, L., Tian, Y., Xia, X. & Chen, Z. Structure and regulation of the chromatin remodeller ISWI. Nature 540, 466–469 (2016).
Davey, C. A., Sargent, D. F., Luger, K., Maeder, A. W. & Richmond, T. J. Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 Å resolution. J. Mol. Biol. 319, 1097–1113 (2002).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Yang, J. et al. The I-TASSER suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Kiefer, F., Arnold, K., Künzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Acknowledgements
We thank A. Siebert and Y. Chaban at eBIC for assistance with data collection, and C. Aylett for help in preparing the nucleosome model. Electron microscopy access and support were provided by the UK national electron Bio-Imaging Centre (eBIC) (proposal EM14769), funded by the Wellcome Trust, MRC and BBSRC. The work was funded by the Wellcome Trust (D.B.W. and X.Z.), Cancer Research UK (D.B.W.) and an Imperial College President’s PhD Scholarship (R.A.).
Reviewer information
Nature thanks B. Bartholomew, O. Llorca and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
D.B.W. and X.Z. designed the studies. R.A., R.J.A., M.W. and O.W. performed the cryo-electron microscopy analysis, and built and refined the structural models. O.W., E.A.M. and L.O. prepared the samples. O.W. conducted the biochemical experiments. D.B.W. and X.Z. wrote the manuscript with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Analysis of INO80–nucleosome complex sample.
a, MST experiment of INO80 with nucleosome flanked by 60 and 12 base pair overhangs (60N12) (±3 mM ADP•BeF3). Raw data (top) were processed to analyse binding and cooperativity (bottom). Data points represent mean values with s.d.; n = 3 experimentally independent replicates. b, Gel of electron microscopy sample (INO80 + nucleosome). Two loadings are shown to enable assessment of INO80 stoichiometry (left) or histones (right). n = 3 independent experimental measurements. c, DNA sequence of the 50N25 nucleosome used for the structure determination. The Widom sequence (yellow) is flanked by 50 base pairs on one side and 25 base pairs on the other. A three-base single strand overhang that remained from the restriction cleavage site is depicted in lowercase. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 2 Cryo-electron microscopy data processing of INO80–nucleosome complex.
a, A typical micrograph out of the 5,479 micrographs generated. b, Representative 2D classes (from 100 generated) obtained with RELION from 775,804 particles. c, Image processing scheme. Data were processed by two parallel pathways to obtain maps for model building.
Extended Data Fig. 3 Quality of the structures.
a, Local resolution map of the INO80–nucleosome complex (4.8 Å) (left) and cut away (right). b, Angular distribution of these particles. c, Local resolution map of the INO80 complex (3.8 Å) (left) and cut away (right). d, Angular distribution of these particles. e, Corrected FSC curves of the reconstructions.
Extended Data Fig. 4 Assessment of various structural features in the INO80–nucleosome complex.
a, Overall fold of the INO80-I and motor domains. b, Locations of the INO80-I, motor domains and IES2 regions relative to the RUVBL1–RUVBL2 hexamer. c, Sequence alignment of the C-terminal regions of human and yeast IES2. The built part of the human IES2 structure is indicated by a yellow bar. Asterisks indicate lysine residues in yeast Ies2 that crosslink to Ino80-HN (red) or Ino80-HC (blue). d, Representative density from two regions of the INO80. Insert: top, Density in the deposited 4.8 Å INO80-nucleosome map; bottom, improvement in density in the 3.8 Å map, which facilitated model building. e, Coordinates of IES2 showing formation of β-sheet secondary structure with RUVBL1 (chain E) and RUVBL2 (chain D) within the 3.8 Å map. f, Left, fit of ARP5 into 4.8 Å map (left). Centre, DNA and motor domains fit into the 4.8 Å map (centre). Right, perpendicular view of centre panel that shows the DNA crossing the motor domains.
Extended Data Fig. 5 Comparisons of INO80–nucleosome interactions with those of Chd1 and Snf2.
Images are viewed from the top of nucleosome, showing that all the motor domains are located on one side and that ARP5–IES6 (green) contacts the other side of the DNA wrap. Chd1 induces an unwrapping of the DNA at the SHL −7 position owing in a large part to interactions with the accessory SANT and SLIDE domains. Despite this unwrapping, the histone core remains largely unaltered. Although the Snf2–nucleosome structure does not induce unwrapping of DNA, it is only a fragment of the motor subunit and also lacks other accessory subunits of the SWI–SNF complex and so probably presents an incomplete picture of interactions or DNA distortions within the nucleosome in the complex.
Extended Data Fig. 6 Interaction of human actin, ARP5 and ARP8 with human H2A–H2B dimers assessed by in vitro pulldown.
a, Actin and actin-related proteins were all expressed with a C-terminal double-Strep tag and used as bait to capture untagged H2A–H2B dimers. The result supports the position of ARP5 in the reported structure. Assay products were visualized by SDS–PAGE and Coomassie staining. n = 1. b, A comparison of ARP5–IES6 and ARP5 nucleosome-binding activity assayed by electrophoretic mobility shift assay, which demonstrates a lack of nucleosome-binding activity by ARP5 at in vivo relevant concentrations in the absence of IES6. Nucleosomes were labelled with Alexa Fluor 488. Reaction species were visualized by fluorescent scan. n = 1. c, ARP5–IES6 and 0N100 nucleosome interaction measured by MST. d, ARP5–IES6 and H2A–H2B interaction measured by MST. For gel source data, see Supplementary Fig. 1. n = 2 biologically independent experiments in all the graphs. Error bars represent s.d. from the mean values.
Extended Data Fig. 7 INO80 SC1 is flexible in the INO80–nucleosome complex.
a, Individual particles (selected out of 775,804 particles in total) with RUVBL1–RUVBL2 oriented similarly, to show different orientations of SC1 (dashed lines). b, Two-dimensional class averages (~30 particles each) showing different orientations of SC1 relative to RUVBL1–RUVBL2. c, Projections of the 3D reconstruction along the same angles of those in b, confirming the extra density as SC1. Scale bar, 100 Å.
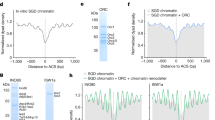
Extended Data Fig. 8 INO80 is regulated by H3 tails.
a, Schematic of histone tail truncations used in this study. b, Initial nucleosome sliding rates of human nucleosomes that lacked different histone tails. Plots of raw data for each histone tail deletion, with Vmax obtained after fitting the data shown as a dotted line. Data are summarized in Fig. 4a. n = 2 biologically independent experiments in all the graphs. Error bars represent s.d. from the mean values.
Extended Data Fig. 9 INO80 is regulated by H3 tails.
a, ATPase data and Hill coefficients for data shown in Fig. 4c. b, ATPase rates for mutations of the H3 tails. c, Nucleosomes carrying wild-type or mutated H3 tails show similar salt stability, which indicates that the mutations have not altered the stability of nucleosomes. n = 2 biologically independent experiments in all the panels. Error bars represent s.d. from the mean values.
Supplementary information
Supplementary Figure
This file contains original gel images for Extended Data Fig. 1b, Extended Data Fig. 6a, 6b
Video 1: Rotating movie of INO80-nucleosome
The 3D reconstruction is shown with structural models fitted in as in Fig. 1. The video was created using Chimera
Rights and permissions
About this article
Cite this article
Ayala, R., Willhoft, O., Aramayo, R.J. et al. Structure and regulation of the human INO80–nucleosome complex. Nature 556, 391–395 (2018). https://doi.org/10.1038/s41586-018-0021-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0021-6
This article is cited by
-
Energy-driven genome regulation by ATP-dependent chromatin remodellers
Nature Reviews Molecular Cell Biology (2024)
-
Integration of multi-omics summary data reveals the role of N6-methyladenosine in neuropsychiatric disorders
Molecular Psychiatry (2024)
-
Context-specific functions of chromatin remodellers in development and disease
Nature Reviews Genetics (2024)
-
Integration of genetic, transcriptomic, and clinical data provides insight into 16p11.2 and 22q11.2 CNV genes
Genome Medicine (2021)
-
Genome information processing by the INO80 chromatin remodeler positions nucleosomes
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.