Abstract
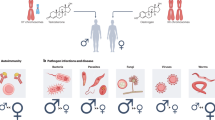
Rheumatic diseases affect a wide range of individuals of all ages, but the most common diseases occur more frequently in women than in men, at ratios of up to ten women to one man. Despite a growing number of studies on sex bias in rheumatic diseases, sex-specific health care is limited and sex specificity is not systematically integrated into treatment regimens. Women and men differ in three major biological points: the number of X chromosomes per cell, the type and quantities of sex hormones present and the ability to be pregnant, all of which have immunological consequences. Could a greater understanding of these differences lead to a new era of personalized sex-specific medicine? This Review focuses on the main genetic and epigenetic mechanisms that have been put forward to explain sex bias in rheumatic diseases, including X chromosome inactivation, sex chromosome aneuploidy and microchimerism. The influence of sex hormones is not discussed in detail in this Review, as it has been well described elsewhere. Understanding the sex-specific factors that contribute to the initiation and progression of rheumatic diseases will enable progress to be made in the diagnosis, treatment and management of all patients with these conditions.
Key points
-
Overall, women are more frequently affected than men by rheumatic diseases and, to date, little sex-specific health care exists.
-
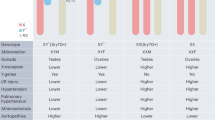
Men often have a stronger genetic predisposition for rheumatic diseases than women, who are predisposed by other factors (for example, pregnancy or carrying two X chromosomes).
-
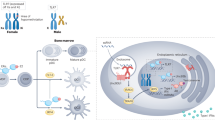
The X chromosome is enriched for immunity-related genes, thus immune functions and immune dysregulation can result from skewed X chromosome inactivation or escape from X chromosome inactivation.
-
Individuals with sex chromosome aneuploidy have an increased risk of autoimmune disorders.
-
Feto–maternal traffic of cells during pregnancy and their long-term persistence in their respective hosts might contribute to the high prevalence of rheumatic diseases in women.
-
The collection and analysis of genetic and epigenetic data in a sex-stratified manner for the development of sex-specific medicine remain challenging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hootman, J. M. & Helmick, C. G. Projections of US prevalence of arthritis and associated activity limitations. Arthritis Rheum. 54, 226–229 (2006).
Jafarzadeh, S. R. & Felson, D. T. Corrected estimates for the prevalence of self-reported doctor-diagnosed arthritis among US adults: comment on the article by Hootman et al. Arthritis Rheumatol. 69, 1701–1702 (2017).
Al Maini, M. et al. The global challenges and opportunities in the practice of rheumatology: white paper by the World Forum on Rheumatic and Musculoskeletal Diseases. Clin. Rheumatol. 34, 819–829 (2015).
Cooper, G. S. & Stroehla, B. C. The epidemiology of autoimmune diseases. Autoimmun. Rev. 2, 119–125 (2003).
van der Slik, B. et al. Although female patients with ankylosing spondylitis score worse on disease activity than male patients and improvement in disease activity is comparable, male patients show more radiographic progression during treatment with TNF-α inhibitors. Semin. Arthritis Rheum. 48, 828–833 (2019).
Cattalini, M., Soliani, M., Caparello, M. C. & Cimaz, R. Sex differences in pediatric rheumatology. Clin. Rev. Allergy Immunol. 56, 293–307 (2019).
Cathcart, E. S. & O’Sullivan, J. B. Rheumatoid arthritis in a New England town. A prevalence study in Sudbury, Massachusetts. N. Engl. J. Med. 282, 421–424 (1970).
Harvey, J., Lotze, M., Stevens, M. B., Lambert, G. & Jacobson, D. Rheumatoid arthritis in a Chippewa Band. I. Pilot screening study of disease prevalence. Arthritis Rheum. 24, 717–721 (1981).
Molokhia, M. & McKeigue, P. Risk for rheumatic disease in relation to ethnicity and admixture. Arthritis Res. 2, 115–125 (2000).
Regitz-Zagrosek, V. Sex and gender differences in health. Science & Society Series on Sex and Science. EMBO Rep. 13, 596–603 (2012).
Dospinescu, P., Jones, G. T. & Basu, N. Environmental risk factors in systemic sclerosis. Curr. Opin. Rheumatol. 25, 179–183 (2013).
Krasselt, M. & Baerwald, C. Sex, symptom severity, and quality of life in rheumatology. Clin. Rev. Allergy Immunol. 56, 346–361 (2019).
Billi, A. C., Kahlenberg, J. M. & Gudjonsson, J. E. Sex bias in autoimmunity. Curr. Opin. Rheumatol. 31, 53–61 (2019).
Kirino, Y. & Remmers, E. F. Genetic architectures of seropositive and seronegative rheumatic diseases. Nat. Rev. Rheumatol. 11, 401–414 (2015).
Deng, Y. & Tsao, B. P. Genetic susceptibility to systemic lupus erythematosus in the genomic era. Nat. Rev. Rheumatol. 6, 683–692 (2010).
Radstake, T. R. et al. Genome-wide association study of systemic sclerosis identifies CD247 as a new susceptibility locus. Nat. Genet. 42, 426–429 (2010).
Dieudé, P. et al. Independent replication establishes the CD247 gene as a genetic systemic sclerosis susceptibility factor. Ann. Rheum. Dis. 70, 1695–1696 (2011).
Allanore, Y. et al. Genome-wide scan identifies TNIP1, PSORS1C1, and RHOB as novel risk loci for systemic sclerosis. PLOS Genet. 7, e1002091 (2011).
Eyre, S., Orozco, G. & Worthington, J. The genetics revolution in rheumatology: large scale genomic arrays and genetic mapping. Nat. Rev. Rheumatol. 13, 421–432 (2017).
MacGregor, A., Ollier, W., Thomson, W., Jawaheer, D. & Silman, A. HLA-DRB1*0401/0404 genotype and rheumatoid arthritis: increased association in men, young age at onset, and disease severity. J. Rheumatol. 22, 1032–1036 (1995).
Hughes, T. et al. Analysis of autosomal genes reveals gene-sex interactions and higher total genetic risk in men with systemic lupus erythematosus. Ann. Rheum. Dis. 71, 694–699 (2012).
Gregersen, P. K., Silver, J. & Winchester, R. J. The shared epitope hypothesis. An approach to understanding the molecular genetics of susceptibility to rheumatoid arthritis. Arthritis Rheum. 30, 1205–1213 (1987).
Lambert, N. C. et al. HLA-DQA1*0501 is associated with diffuse systemic sclerosis in Caucasian men. Arthritis Rheum. 43, 2005–2010 (2000).
Ciurea, A. et al. Age at symptom onset in ankylosing spondylitis: is there a gender difference? Ann. Rheum. Dis. 73, 1908–1910 (2014).
Plenge, R. M. et al. Replication of putative candidate-gene associations with rheumatoid arthritis in >4,000 samples from North America and Sweden: association of susceptibility with PTPN22, CTLA4, and PADI4. Am. J. Hum. Genet. 77, 1044–1060 (2005).
Hinks, A. et al. Brief report: the genetic profile of rheumatoid factor-positive polyarticular juvenile idiopathic arthritis resembles that of adult rheumatoid arthritis. Arthritis Rheumatol. 70, 957–962 (2018).
Webb, R. et al. Early disease onset is predicted by a higher genetic risk for lupus and is associated with a more severe phenotype in lupus patients. Ann. Rheum. Dis. 70, 151–156 (2011).
Ross, M. T. et al. The DNA sequence of the human X chromosome. Nature 434, 325–337 (2005).
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lahn, B. T. & Page, D. C. Functional coherence of the human Y chromosome. Science 278, 675–680 (1997).
Wise, A. L., Gyi, L. & Manolio, T. A. eXclusion: toward integrating the X chromosome in genome-wide association analyses. Am. J. Hum. Genet. 92, 643–647 (2013).
Eyre, S. et al. High-density genetic mapping identifies new susceptibility loci for rheumatoid arthritis. Nat. Genet. 44, 1336–1340 (2012).
Jacob, C. O. et al. Identification of IRAK1 as a risk gene with critical role in the pathogenesis of systemic lupus erythematosus. Proc. Natl Acad. Sci. USA 106, 6256–6261 (2009).
Kaufman, K. M. et al. Fine mapping of Xq28: both MECP2 and IRAK1 contribute to risk for systemic lupus erythematosus in multiple ancestral groups. Ann. Rheum. Dis. 72, 437–444 (2013).
Liu, G., Tsuruta, Y., Gao, Z., Park, Y. J. & Abraham, E. Variant IL-1 receptor-associated kinase-1 mediates increased NF-κB activity. J. Immunol. 179, 4125–4134 (2007).
He, F. et al. Detection of parent-of-origin effects for quantitative traits in complete and incomplete nuclear families with multiple children. Am. J. Epidemiol. 174, 226–233 (2011).
He, H. Q. et al. Detection of parent-of-origin effects for quantitative traits using general pedigree data. J. Genet. 93, 339–347 (2014).
Morison, I. M., Ramsay, J. P. & Spencer, H. G. A census of mammalian imprinting. Trends Genet. 21, 457–465 (2005).
Zou, Q. L., You, X. P., Li, J. L., Fung, W. K. & Zhou, J. Y. A powerful parent-of-origin effects test for qualitative traits on X chromosome in general pedigrees. BMC Bioinformatics 19, 8 (2018).
Bianchi, I. et al. The X chromosome and immune associated genes. J. Autoimmun. 38, J187–J192 (2012).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Souyris, M., Mejia, J. E., Chaumeil, J. & Guery, J. C. Female predisposition to TLR7-driven autoimmunity: gene dosage and the escape from X chromosome inactivation. Semin. Immunopathol. 41, 153–164 (2019).
Chamberlain, N. D. et al. Ligation of TLR7 by rheumatoid arthritis synovial fluid single strand RNA induces transcription of TNFα in monocytes. Ann. Rheum. Dis. 72, 418–426 (2013).
Young, N. A. et al. Estrogen modulation of endosome-associated Toll-like receptor 8: an IFNα-independent mechanism of sex-bias in systemic lupus erythematosus. Clin. Immunol. 151, 66–77 (2014).
Komatsuda, A. et al. Up-regulated expression of Toll-like receptors mRNAs in peripheral blood mononuclear cells from patients with systemic lupus erythematosus. Clin. Exp. Immunol. 152, 482–487 (2008).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Radstake, T. R. et al. Increased frequency and compromised function of T regulatory cells in systemic sclerosis (SSc) is related to a diminished CD69 and TGFβ expression. PLOS ONE 4, e5981 (2009).
Ehrenstein, M. R. et al. Compromised function of regulatory T cells in rheumatoid arthritis and reversal by anti-TNFα therapy. J. Exp. Med. 2, 277–285 (2004).
van Amelsfort, J. M. et al. Proinflammatory mediator-induced reversal of CD4+,CD25+ regulatory T cell-mediated suppression in rheumatoid arthritis. Arthritis Rheum. 56, 732–742 (2007).
Yan, B. et al. Dysfunctional CD4+,CD25+ regulatory T cells in untreated active systemic lupus erythematosus secondary to interferon-alpha-producing antigen-presenting cells. Arthritis Rheum. 58, 801–812 (2008).
Bonelli, M., von Dalwigk, K., Savitskaya, A., Smolen, J. S. & Scheinecker, C. Foxp3 expression in CD4+ T cells of patients with systemic lupus erythematosus: a comparative phenotypic analysis. Ann. Rheum. Dis. 67, 664–671 (2008).
Peters, A. L. CD40 and autoimmunity: the dark side of a great activator. Semin. Immunol. 21, 293–300 (2009).
Croft, M. & Siegel, R. M. Beyond TNF: TNF superfamily cytokines as targets for the treatment of rheumatic diseases. Nat. Rev. Rheumatol. 13, 217–233 (2017).
Durie, F. H. et al. Prevention of collagen-induced arthritis with an antibody to gp39, the ligand for CD40. Science 261, 1328–1330 (1993).
Early, G. S., Zhao, W. & Burns, C. M. Anti-CD40 ligand antibody treatment prevents the development of lupus-like nephritis in a subset of New Zealand black x New Zealand white mice. Response correlates with the absence of an anti-antibody response. J. Immunol. 157, 3159–3164 (1996).
Mohan, C., Shi, Y., Laman, J. D. & Datta, S. K. Interaction between CD40 and its ligand gp39 in the development of murine lupus nephritis. J. Immunol. 154, 1470–1480 (1995).
Seillet, C. et al. The TLR-mediated response of plasmacytoid dendritic cells is positively regulated by estradiol in vivo through cell-intrinsic estrogen receptor alpha signaling. Blood 119, 454–464 (2012).
Laffont, S. et al. X-chromosome complement and estrogen receptor signaling independently contribute to the enhanced TLR7-mediated IFN-alpha production of plasmacytoid dendritic cells from women. J. Immunol. 193, 5444–5452 (2014).
Meier, A. et al. Sex differences in the Toll-like receptor-mediated response of plasmacytoid dendritic cells to HIV-1. Nat. Med. 15, 955–959 (2009).
Jefferies, C. A. Regulating IRFs in IFN driven disease. Front. Immunol. 10, 325 (2019).
Schoenemeyer, A. et al. The interferon regulatory factor, IRF5, is a central mediator of Toll-like receptor 7 signaling. J. Biol. Chem. 280, 17005–17012 (2005).
Takaoka, A. et al. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature 434, 243–249 (2005).
Laffont, S., Seillet, C. & Guery, J. C. Estrogen receptor-dependent regulation of dendritic cell development and function. Front. Immunol 8, 108 (2017).
Ban, T., Sato, G. R. & Tamura, T. Regulation and role of the transcription factor IRF5 in innate immune responses and systemic lupus erythematosus. Int. Immunol. 30, 529–536 (2018).
Cham, C. M., Ko, K. & Niewold, T. B. Interferon regulatory factor 5 in the pathogenesis of systemic lupus erythematosus. Clin. Dev. Immunol. 2012, 780436 (2012).
Niewold, T. B. et al. Association of the IRF5 risk haplotype with high serum interferon-alpha activity in systemic lupus erythematosus patients. Arthritis Rheum. 58, 2481–2487 (2008).
Griesbeck, M. et al. Sex differences in plasmacytoid dendritic cell levels of IRF5 drive higher IFN-alpha production in women. J. Immunol. 195, 5327–5336 (2015).
Souyris, M. et al. TLR7 escapes X chromosome inactivation in immune cells. Sci. Immunol. 3, eaap8855 (2018).
Smith-Bouvier, D. L. et al. A role for sex chromosome complement in the female bias in autoimmune disease. J. Exp. Med. 205, 1099–1108 (2008).
Smith, D. L. et al. A female preponderance for chemically induced lupus in SJL/J mice. Clin. Immunol. 122, 101–107 (2007).
Pisitkun, P. et al. Autoreactive B cell responses to RNA-related antigens due to TLR7 gene duplication. Science 312, 1669–1672 (2006).
Deane, J. A. et al. Control of Toll-like receptor 7 expression is essential to restrict autoimmunity and dendritic cell proliferation. Immunity 27, 801–810 (2007).
Koelsch, K. A. et al. Functional characterization of the MECP2/IRAK1 lupus risk haplotype in human T cells and a human MECP2 transgenic mouse. J. Autoimmun. 41, 168–174 (2013).
Case, L. K. et al. The Y chromosome as a regulatory element shaping immune cell transcriptomes and susceptibility to autoimmune disease. Genome Res. 23, 1474–1485 (2013).
Kanaan, S. B. et al. Evaluation of X chromosome inactivation with respect to HLA genetic susceptibility in rheumatoid arthritis and systemic sclerosis. PLOS ONE 11, e0158550 (2016).
Ozbalkan, Z. et al. Skewed X chromosome inactivation in blood cells of women with scleroderma. Arthritis Rheum. 52, 1564–1570 (2005).
Chabchoub, G. et al. Analysis of skewed X-chromosome inactivation in females with rheumatoid arthritis and autoimmune thyroid diseases. Arthritis Res. Ther. 11, R106 (2009).
Lambert, N. C. The price of silence. Arthritis Rheum. 60, 3164–3167 (2009).
Brown, C. J. & Robinson, W. P. The causes and consequences of random and non-random X chromosome inactivation in humans. Clin. Genet. 58, 353–363 (2000).
Asplund, A., Guo, Z., Hu, X., Wassberg, C. & Ponten, F. Mosaic pattern of maternal and paternal keratinocyte clones in normal human epidermis revealed by analysis of X-chromosome inactivation. J. Invest. Dermatol. 117, 128–131 (2001).
Busque, L. et al. Skewing of X-inactivation ratios in blood cells of aging women is confirmed by independent methodologies. Blood 113, 3472–3474 (2009).
Broen, J. C. et al. Skewed X chromosomal inactivation impacts T regulatory cell function in systemic sclerosis. Ann. Rheum. Dis. 69, 2213–2216 (2010).
Uz, E. et al. Increased frequency of extremely skewed X chromosome inactivation in juvenile idiopathic arthritis. Arthritis Rheum. 60, 3410–3412 (2009).
Ek, W. et al. Mapping QTL affecting a systemic sclerosis-like disorder in a cross between UCD-200 and red jungle fowl chickens. Dev. Comp. Immunol. 38, 352–359 (2012).
Itoh, Y. et al. Dosage compensation is less effective in birds than in mammals. J. Biol. 6, 2 (2007).
Tukiainen, T. et al. Landscape of X chromosome inactivation across human tissues. Nature 550, 244–248 (2017).
Carrel, L. & Willard, H. F. X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434, 400–404 (2005).
Garieri, M. et al. Extensive cellular heterogeneity of X inactivation revealed by single-cell allele-specific expression in human fibroblasts. Proc. Natl Acad. Sci. USA 115, 13015–13020 (2018).
Wang, J. et al. Unusual maintenance of X chromosome inactivation predisposes female lymphocytes for increased expression from the inactive X. Proc. Natl Acad. Sci. USA 113, E2029–E2038 (2016).
Lu, Q. et al. Demethylation of CD40LG on the inactive X in T cells from women with lupus. J. Immunol. 179, 6352–6358 (2007).
Hewagama, A. et al. Overexpression of X-linked genes in T cells from women with lupus. J. Autoimmun. 41, 60–71 (2013).
Syrett, C. M. et al. Altered X-chromosome inactivation in T cells may promote sex-biased autoimmune diseases. JCI Insight 4, e126751 (2019).
Prothero, K. E., Stahl, J. M. & Carrel, L. Dosage compensation and gene expression on the mammalian X chromosome: one plus one does not always equal two. Chromosome Res. 17, 637–648 (2009).
Fuks, F. et al. The methyl-CpG-binding protein MeCP2 links DNA methylation to histone methylation. J. Biol. Chem. 278, 4035–4040 (2003).
Hammond, S. M. An overview of microRNAs. Adv. Drug Deliv. Rev. 87, 3–14 (2015).
Pinheiro, I., Dejager, L. & Libert, C. X-chromosome-located microRNAs in immunity: might they explain male/female differences? The X chromosome-genomic context may affect X-located miRNAs and downstream signaling, thereby contributing to the enhanced immune response of females. Bioessays 33, 791–802 (2011).
Kozomara, A., Birgaoanu, M. & Griffiths-Jones, S. miRBase: from microRNA sequences to function. Nucleic Acids Res. 47, D155–D162 (2019).
Hashemi, M. et al. Association of pre-miRNA-146a rs2910164 and pre-miRNA-499 rs3746444 polymorphisms and susceptibility to rheumatoid arthritis. Mol. Med. Rep. 7, 287–291 (2013).
Chatzikyriakidou, A. et al. A polymorphism in the 3’-UTR of interleukin-1 receptor-associated kinase (IRAK1), a target gene of miR-146a, is associated with rheumatoid arthritis susceptibility. Joint Bone Spine 77, 411–413 (2010).
Yang, X. K. et al. Association between IRAK1 rs3027898 and miRNA-499 rs3746444 polymorphisms and rheumatoid arthritis: a case control study and meta-analysis. Z. Rheumatol. 76, 622–629 (2017).
Khalifa, O. et al. X-linked miRNAs associated with gender differences in rheumatoid arthritis. Int. J. Mol. Sci. 17, 1852 (2016).
Scofield, R. H. et al. Klinefelter’s syndrome (47,XXY) in male systemic lupus erythematosus patients: support for the notion of a gene-dose effect from the X chromosome. Arthritis Rheum. 58, 2511–2517 (2008).
Harris, V. M. et al. Klinefelter’s syndrome (47,XXY) is in excess among men with Sjögren’s syndrome. Clin. Immunol. 168, 25–29 (2016).
Rovensky, J., Imrich, R., Lazurova, I. & Payer, J. Rheumatic diseases and Klinefelter’s syndrome. Ann. NY Acad. Sci. 1193, 1–9 (2010).
Liu, K. et al. X chromosome dose and sex bias in autoimmune diseases: increased prevalence of 47,XXX in systemic lupus erythematosus and Sjögren’s syndrome. Arthritis Rheumatol. 68, 1290–1300 (2016).
Bojesen, A., Juul, S. & Gravholt, C. H. Prenatal and postnatal prevalence of Klinefelter syndrome: a national registry study. J. Clin. Endocrinol. Metab. 88, 622–626 (2003).
Nielsen, J. & Wohlert, M. Chromosome abnormalities found among 34,910 newborn children: results from a 13-year incidence study in Århus, Denmark. Hum. Genet. 87, 81–83 (1991).
Otter, M., Schrander-Stumpel, C. T. & Curfs, L. M. Triple X syndrome: a review of the literature. Eur. J. Hum. Genet. 18, 265–271 (2010).
Sharma, R. et al. Rare X chromosome abnormalities in systemic lupus erythematosus and Sjögren’s syndrome. Arthritis Rheumatol. 69, 2187–2192 (2017).
Abdelmoula, N. B. et al. Cytogenetics and fluorescence in situ hybridization assessment of sex-chromosome mosaicism in Klinefelter’s syndrome. Ann. Genet. 47, 163–175 (2004).
Samplaski, M. K. et al. Phenotypic differences in mosaic Klinefelter patients as compared with non-mosaic Klinefelter patients. Fertil. Steril. 101, 950–955 (2014).
Martin, G. V. et al. Mosaicism with XX and XXY cells accounts for high copy number of Toll like receptor 7 and 8 genes in peripheral blood of men with rheumatoid arthritis. Sci. Rep. 9, 12880 (2019).
Sharma, A. et al. DNA methylation signature in peripheral blood reveals distinct characteristics of human X chromosome numerical aberrations. Clin. Epigenetics 7, 76 (2015).
Rocca, M. S. et al. The Klinefelter syndrome is associated with high recurrence of copy number variations on the X chromosome with a potential role in the clinical phenotype. Andrology 4, 328–334 (2016).
Jorgensen, K. T. et al. Autoimmune diseases in women with Turner’s syndrome. Arthritis Rheum. 62, 658–666 (2010).
Armagan, O., Ekim, A., Dinc, A. & Oner, C. Ankylosing spondylitis in a patient with Turner syndrome: a case report. Rheumatol. Int. 27, 1177–1180 (2007).
Wihlborg, C. E., Babyn, P. S. & Schneider, R. The association between Turner’s syndrome and juvenile rheumatoid arthritis. Pediatr. Radiol. 29, 676–681 (1999).
Invernizzi, P. et al. X chromosome monosomy: a common mechanism for autoimmune diseases. J. Immunol. 175, 575–578 (2005).
Castellanos, M. V. et al. Chromosomal abnormalities are related to location and grade of osteoarthritis. Osteoarthr. Cartil. 12, 982–985 (2004).
Schaschl, H., Aitman, T. J. & Vyse, T. J. Copy number variation in the human genome and its implication in autoimmunity. Clin. Exp. Immunol. 156, 12–16 (2009).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006).
Lee, Y. H. et al. Association between FCGR3B copy number variations and susceptibility to autoimmune diseases: a meta-analysis. Inflamm. Res. 64, 983–991 (2015).
McKinney, C. et al. Evidence for an influence of chemokine ligand 3-like 1 (CCL3L1) gene copy number on susceptibility to rheumatoid arthritis. Ann. Rheum. Dis. 67, 409–413 (2008).
Bailey, J. A., Carrel, L., Chakravarti, A. & Eichler, E. E. Molecular evidence for a relationship between LINE-1 elements and X chromosome inactivation: the Lyon repeat hypothesis. Proc. Natl Acad. Sci. USA 97, 6634–6639 (2000).
Mavragani, C. P. et al. Expression of long interspersed nuclear element 1 retroelements and induction of type I interferon in patients with systemic autoimmune disease. Arthritis Rheumatol. 68, 2686–2696 (2016).
Ali, M. et al. Overexpression of transcripts containing LINE-1 in the synovia of patients with rheumatoid arthritis. Ann. Rheum. Dis. 62, 663–666 (2003).
Matsuno, Y., Yamashita, T., Wagatsuma, M. & Yamakage, H. Convergence in LINE-1 nucleotide variations can benefit redundantly forming triplexes with lncRNA in mammalian X chromosome inactivation. Mob. DNA 10, 33 (2019).
Carrel, L. et al. Genomic environment predicts expression patterns on the human inactive X chromosome. PLOS Genet. 2, e151 (2006).
Robinson, H. P. & Caines, J. S. Sonar evidence of early pregnancy failure in patients with twin conceptions. Br. J. Obstet. Gynaecol. 84, 22–25 (1977).
Nelson, J. L. Microchimerism: incidental byproduct of pregnancy or active participant in human health? Trends Mol. Med. 8, 109–113 (2002).
Lambert, N. C. in Chimerism: A Clinical Guide (ed. Draper, N. L.) 153–179 (Springer, 2018).
Mold, J. E. et al. Maternal alloantigens promote the development of tolerogenic fetal regulatory T cells in utero. Science 322, 1562–1565 (2008).
Nelson, J. L. Maternal-fetal immunology and autoimmune disease: is some autoimmune disease auto-alloimmune or allo-autoimmune? Arthritis Rheum. 39, 191–194 (1996).
Lambert, N. C. et al. Cutting edge: persistent fetal microchimerism in T lymphocytes is associated with HLA-DQA1*0501: implications in autoimmunity. J. Immunol. 164, 5545–5548 (2000).
Nelson, J. L. et al. Microchimerism and HLA-compatible relationships of pregnancy in scleroderma. Lancet 351, 559–562 (1998).
Di Cristofaro, J. et al. Soluble HLA-G expression inversely correlates with fetal microchimerism levels in peripheral blood from women with scleroderma. Front. Immunol. 9, 1685 (2018).
Rak, J. M. et al. Transfer of the shared epitope through microchimerism in women with rheumatoid arthritis. Arthritis Rheum. 60, 73–80 (2009).
Yan, Z., Aydelotte, T., Gadi, V. K., Guthrie, K. A. & Nelson, J. L. Acquisition of the rheumatoid arthritis HLA shared epitope through microchimerism. Arthritis Rheum. 63, 640–644 (2011).
Cruz, G. I. et al. Mother-child histocompatibility and risk of rheumatoid arthritis and systemic lupus erythematosus among mothers. Genes Immun. https://doi.org/10.1038/s41435-018-0055-7 (2019).
Lo, Y. M. et al. Quantitative abnormalities of fetal DNA in maternal serum in preeclampsia. Clin. Chem. 45, 184–188 (1999).
Khosrotehrani, K. et al. The influence of fetal loss on the presence of fetal cell microchimerism: a systematic review. Arthritis Rheum. 48, 3237–3241 (2003).
van Wyk, L. et al. Increased incidence of pregnancy complications in women who later develop scleroderma: a case control study. Arthritis Res. Ther. 13, R183 (2011).
Silman, A. J. & Black, C. Increased incidence of spontaneous abortion and infertility in women with scleroderma before disease onset: a controlled study. Ann. Rheum. Dis. 47, 441–444 (1988).
Silman, A. J., Roman, E., Beral, V. & Brown, A. Adverse reproductive outcomes in women who subsequently develop rheumatoid arthritis. Ann. Rheum. Dis. 47, 979–981 (1988).
Cruz, G. I. et al. Increased risk of rheumatoid arthritis among mothers with children who carry DRB1 risk-associated alleles. Ann. Rheum. Dis. 76, 1405–1410 (2017).
Nelson, J. L. & Lambert, N. C. Rheumatoid arthritis: forward and reverse inheritance—the yin and the yang. Nat. Rev. Rheumatol. 13, 396–397 (2017).
Artlett, C. M. et al. Chimeric cells of maternal origin in juvenile idiopathic inflammatory myopathies. Childhood Myositis Heterogeneity Collaborative Group. Lancet 356, 2155–2156 (2000).
Reed, A. M., Picornell, Y. J., Harwood, A. & Kredich, D. W. Chimerism in children with juvenile dermatomyositis. Lancet 356, 2156–2157 (2000).
Stevens, A. M. Maternal microchimerism in health and disease. Best Pract. Res. Clin. Obstet. Gynaecol. 31, 121–130 (2016).
Stevens, A. M., Hermes, H. M., Rutledge, J. C., Buyon, J. P. & Nelson, J. L. Myocardial-tissue-specific phenotype of maternal microchimerism in neonatal lupus congenital heart block. Lancet 362, 1617–1623 (2003).
de Bellefon, L. M. et al. Cells from a vanished twin as a source of microchimerism 40 years later. Chimerism 1, 56–60 (2010).
Lambert, N. C. et al. Quantification of maternal microchimerism by HLA-specific real-time polymerase chain reaction: studies of healthy women and women with scleroderma. Arthritis Rheum. 50, 906–914 (2004).
Stevens, A. M., Hermes, H. M., Kiefer, M. M., Rutledge, J. C. & Nelson, J. L. Chimeric maternal cells with tissue-specific antigen expression and morphology are common in infant tissues. Pediatr. Dev. Pathol. 12, 337–346 (2009).
Gleichmann, E., Pals, S. T., Rolink, A. G., Radaszkiewicz, T. & Gleichmann, H. Graft-versus-host reactions: clues to the etiopathology of a spectrum of immunological diseases. Immunol. Today 5, 324–332 (1984).
Via, C. S. Advances in lupus stemming from the parent-into-F1 model. Trends Immunol. 31, 236–245 (2010).
Kanold, A. M. et al. A research study of the association between maternal microchimerism and systemic lupus erythematosus in adults: a comparison between patients and healthy controls based on single-nucleotide polymorphism using quantitative real-time PCR. PLOS ONE 8, e74534 (2013).
Abbud Filho, M. et al. Systemic lupus erythematosus and microchimerism in autoimmunity. Transplant. Proc. 34, 2951–2952 (2002).
Stevens, A. M. et al. Maternal HLA class II compatibility in men with systemic lupus erythematosus. Arthritis Rheum. 52, 2768–2773 (2005).
Kaneda, T., Shiraki, K., Hirano, K. & Nagata, I. Detection of maternofetal transfusion by placental alkaline phosphatase levels. J. Pediatr. 130, 730–735 (1997).
Zhou, L. et al. Two independent pathways of maternal cell transmission to offspring: through placenta during pregnancy and by breast-feeding after birth. Immunology 101, 570–580 (2000).
Laursen, M. F. et al. Having older siblings is associated with gut microbiota development during early childhood. BMC Microbiol. 15, 154 (2015).
Peterson, S. E. et al. Prospective assessment of fetal-maternal cell transfer in miscarriage and pregnancy termination. Hum. Reprod. 27, 2607–2612 (2012).
Huurre, A. et al. Mode of delivery—effects on gut microbiota and humoral immunity. Neonatology 93, 236–240 (2008).
Shree, R. et al. Fetal microchimerism by mode of delivery: a prospective cohort study. BJOG 126, 24–31 (2019).
Chaudhari, M. et al. Impaired reproductive fitness in mothers of children with juvenile autoimmune arthropathies. Rheumatology 45, 1282–1287 (2006).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 12, 861–874 (2011).
Djouad, F., Bouffi, C., Ghannam, S., Noel, D. & Jorgensen, C. Mesenchymal stem cells: innovative therapeutic tools for rheumatic diseases. Nat. Rev. Rheumatol. 5, 392–399 (2009).
Maumus, M., Jorgensen, C. & Noel, D. Mesenchymal stem cells in regenerative medicine applied to rheumatic diseases: role of secretome and exosomes. Biochimie 95, 2229–2234 (2013).
Hügle, T. & van Laar, J. M. Allogeneic stem cell transplantation for rheumatic autoimmune diseases. F1000 Med. Rep. 2, 22 (2010).
Silva, M. F. J. et al. Allogeneic hematopoietic stem cell transplantation for severe, refractory juvenile idiopathic arthritis. Blood Adv. 2, 777–786 (2018).
Kinder, J. M., Stelzer, I. A., Arck, P. C. & Way, S. S. Immunological implications of pregnancy-induced microchimerism. Nat. Rev. Immunol. 17, 483–494 (2017).
Eikmans, M. et al. Naturally acquired microchimerism: implications for transplantation outcome and novel methodologies for detection. Chimerism 5, 24–39 (2014).
Dutta, P. et al. Microchimerism is strongly correlated with tolerance to noninherited maternal antigens in mice. Blood 114, 3578–3587 (2009).
Molitor-Dart, M. L. et al. Developmental exposure to noninherited maternal antigens induces CD4+ T regulatory cells: relevance to mechanism of heart allograft tolerance. J. Immunol. 179, 6749–6761 (2007).
Burlingham, W. J. A lesson in tolerance–maternal instruction to fetal cells. N. Engl. J. Med. 360, 1355–1357 (2009).
Maria, A. T. et al. Human adipose mesenchymal stem cells as potent anti-fibrosis therapy for systemic sclerosis. J. Autoimmun. 70, 31–39 (2016).
Liang, Y. et al. A gene network regulated by the transcription factor VGLL3 as a promoter of sex-biased autoimmune diseases. Nat. Immunol. 18, 152–160 (2017).
Amur, S., Parekh, A. & Mummaneni, P. Sex differences and genomics in autoimmune diseases. J. Autoimmun. 38, J254–J265 (2012).
Rehman, W., Arfons, L. M. & Lazarus, H. M. The rise, fall and subsequent triumph of thalidomide: lessons learned in drug development. Ther. Adv. Hematol. 2, 291–308 (2011).
Poon, R. et al. Participation of women and sex analyses in late-phase clinical trials of new molecular entity drugs and biologics approved by the FDA in 2007-2009. J. Womens Health 22, 604–616 (2013).
Atzeni, F. et al. Predicting response to anti-TNF treatment in rheumatoid arthritis patients. Autoimmun. Rev. 8, 431–437 (2009).
van der Horst-Bruinsma, I. E., Zack, D. J., Szumski, A. & Koenig, A. S. Female patients with ankylosing spondylitis: analysis of the impact of gender across treatment studies. Ann. Rheum. Dis. 72, 1221–1224 (2013).
Mueller, S. et al. Differences in fecal microbiota in different European study populations in relation to age, gender, and country: a cross-sectional study. Appl. Environ. Microbiol. 72, 1027–1033 (2006).
Lyon, M. F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190, 372–373 (1961).
Pinheiro, I. & Heard, E. X chromosome inactivation: new players in the initiation of gene silencing. F1000Res 6, 344 (2017).
Chang, S. C., Tucker, T., Thorogood, N. P. & Brown, C. J. Mechanisms of X-chromosome inactivation. Front. Biosci. 11, 852–866 (2006).
Avner, P. & Heard, E. X-chromosome inactivation: counting, choice and initiation. Nat. Rev. Genet. 2, 59–67 (2001).
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002).
Yavuz, S. et al. Dual effects of testosterone in Behcet’s disease: implications for a role in disease pathogenesis. Genes Immun. 17, 335–341 (2016).
Kochi, Y. et al. PADI4 polymorphism predisposes male smokers to rheumatoid arthritis. Ann. Rheum. Dis. 70, 512–515 (2011).
Kwon, Y. C. et al. Male-specific association of the FCGR2A His167Arg polymorphism with Kawasaki disease. PLOS ONE 12, e0184248 (2017).
Robinson, J. I. et al. Dissection of the FCGR3A association with RA: increased association in men and with autoantibody positive disease. Ann. Rheum. Dis. 69, 1054–1057 (2010).
Torcia, M. G. et al. Sex differences in the response to viral infections: TLR8 and TLR9 ligand stimulation induce higher IL10 production in males. PLOS ONE 7, e39853 (2012).
Hu, K. et al. STAT4 polymorphism in a Chinese Han population with Vogt-Koyanagi-Harada syndrome and Behçet’s disease. Hum. Immunol. 71, 723–726 (2010).
Gonzalez-Escribano, M. F., Aguilar, F., Sanchez-Roman, J. & Nunez-Roldan, A. FcγRIIA, FcγRIIIA and FcγRIIIB polymorphisms in Spanish patients with systemic lupus erythematosus. Eur. J. Immunogenet. 29, 301–306 (2002).
Prokunina, L. et al. Association of the PD-1.3A allele of the PDCD1 gene in patients with rheumatoid arthritis negative for rheumatoid factor and the shared epitope. Arthritis Rheum. 50, 1770–1773 (2004).
Ferreiros-Vidal, I. et al. Association of PDCD1 with susceptibility to systemic lupus erythematosus: evidence of population-specific effects. Arthritis Rheum. 50, 2590–2597 (2004).
Lee, S. H. et al. Association of the programmed cell death 1 (PDCD1) gene polymorphism with ankylosing spondylitis in the Korean population. Arthritis Res. Ther. 8, R163 (2006).
Kadota, K. et al. Analysis of gender differences in genetic risk: association of TNFAIP3 polymorphism with male childhood-onset systemic lupus erythematosus in the Japanese population. PLOS ONE 8, e72551 (2013).
Pers, Y. M. et al. Association of TRAF1-C5 with risk of uveitis in juvenile idiopathic arthritis. Joint Bone Spine 84, 305–308 (2017).
Albers, H. M. et al. The TRAF1/C5 region is a risk factor for polyarthritis in juvenile idiopathic arthritis. Ann. Rheum. Dis. 67, 1578–1580 (2008).
Hinks, A. et al. Association of the IL2RA/CD25 gene with juvenile idiopathic arthritis. Arthritis Rheum. 60, 251–257 (2009).
Liu, R. et al. Influence of MIF, CD40, and CD226 polymorphisms on risk of rheumatoid arthritis. Mol. Biol. Rep. 39, 6915–6922 (2012).
Acknowledgements
N.C.L. thanks J. Roudier, I. Auger, N. Balandraud, D.F. Azzouz, S.B. Kanaan and G.V. Martin for constructive discussions and J. Buand for editorial assistance. The work of N.L.C. was supported financially by INSERM, Région PACA, Arthritis-Fondation Courtin and Groupe Francophone de Recherche sur la Sclérodermie (GFRS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Peer review information
Nature Reviews Rheumatology thanks R. H. Scofield, M. Anguerra, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
miRBase: http://mirbase.org
Supplementary information
Glossary
- Microchimerism
-
The presence, in small quantities, of foreign DNA or cells in an individual.
- Shared epitope
-
A characteristic five amino acid sequence in the HLA-DRβ1 chain, encoded by allelic variants associated with risk of rheumatoid arthritis.
- Submetacentric
-
When the centromere is located on the chromosome so that chromosomal arm lengths are unequal, the chromosome is said to be submetacentric.
- Acrocentric
-
When the centromere is located on the chromosome so that one chromosomal arm is much shorter than the other, the chromosome is said to be acrocentric.
- Mosaicism
-
A mixture of two or more populations of genetically different cells within an individual.
- Mouse constructions
-
The creation of genetically engineered mice as tools for studying human diseases.
Rights and permissions
About this article
Cite this article
Lambert, N.C. Nonendocrine mechanisms of sex bias in rheumatic diseases. Nat Rev Rheumatol 15, 673–686 (2019). https://doi.org/10.1038/s41584-019-0307-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41584-019-0307-6
This article is cited by
-
DNA methylation profiling of labial salivary gland tissues revealed hypomethylation of B-cell-related genes in primary Sjögren’s syndrome
Immunologic Research (2024)
-
Sex steroids and autoimmune rheumatic diseases: state of the art
Nature Reviews Rheumatology (2020)