Abstract
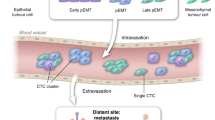
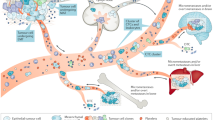
During cancer progression, many tumours shed circulating tumour cells (CTCs) and other biomarkers into the bloodstream. The analysis of CTCs offers the prospect of collecting a liquid biopsy from a patient’s blood to predict and monitor therapeutic responses and tumour recurrence. In this Review, we discuss progress towards the isolation and recovery of bulk CTCs from whole blood samples for the identification of cells with high metastatic potential. We provide an overview of the techniques that initially pointed to the clinical significance of CTCs and describe the key requirements for clinical applications of CTC analysis. We also summarize recent advances that permit the functional and biochemical phenotypes of CTCs to be characterized, and discuss the potential roles of these CTC characteristics in the formation of metastatic lesions. Moreover, we discuss the use of circulating tumour DNA and exosomes as markers for early cancer diagnosis and for the monitoring of cancer progression. Next-generation technologies and biomarkers for invasive cancers should allow for the unequivocal determination of the metastatic potential of CTCs, and for the meaningful analysis of circulating tumour DNA and exosomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Pantel, K. & Brakenhoff, R. H. Dissecting the metastatic cascade. Nat. Rev. Cancer 4, 448–456 (2004).
Alix-Panabieres, C. & Pantel, K. Circulating tumor cells: liquid biopsy of cancer. Clin. Chem. 59, 110–118 (2013).
Pantel, K., Alix-Panabières, C. & Riethdorf, S. Cancer micrometastases. Nat. Rev. Clin. Oncol. 6, 339–351 (2009).
Chaffer, C. L. & Weinberg, R. A. A perspective on cancer cell metastasis. Science 331, 1559–1564 (2011).
Joosse, S. A., Gorges, T. M. & Pantel, K. Biology, detection, and clinical implications of circulating tumor cells. EMBO Mol. Med. 7, 1–11 (2015).
Dasgupta, A., Lim, A. R. & Ghajar, C. M. Circulating and disseminated tumor cells: harbingers or initiators of metastasis? Mol. Oncol. 11, 40–61 (2017).
Riethdorf, S. et al. Detection of circulating tumor cells in peripheral blood of patients with metastatic breast cancer: a validation study of the CellSearch system. Clin. Cancer Res. 13, 920–928 (2007).
Romiti, A. et al. Circulating tumor cells count predicts survival in colorectal cancer patients. J. Gastrointestin. Liver Dis. 23, 279–284 (2014).
Bidard, F.-C. et al. Clinical validity of circulating tumour cells in patients with metastatic breast cancer: a pooled analysis of individual patient data. Lancet Oncol. 15, 406–414 (2014).
Lorente, D. et al. Decline in circulating tumor cell count and treatment outcome in advanced prostate cancer. Eur. Urol. 70, 985–992 (2016).
Franken, B. et al. Circulating tumor cells, disease recurrence and survival in newly diagnosed breast cancer. Breast Cancer Res. 14, R133 (2012).
van Dalum, G. et al. Importance of circulating tumor cells in newly diagnosed colorectal cancer. Int. J. Oncol. 46, 1361–1368 (2015).
Karl, A., Tritschler, S., Hofmann, S., Stief, C. G. & Schindlbeck, C. Perioperative search for circulating tumor cells in patients undergoing radical cystectomy for bladder cancer. Eur. J. Med. Res. 14, 487–490 (2009).
Riethdorf, S. et al. Prognostic impact of circulating tumor cells for breast cancer patients treated in the neoadjuvant ‘Geparquattro’ trial. Clin. Cancer Res. 23, 5384–5393 (2017).
Valastyan, S. & Weinberg, R. A. Tumor metastasis: molecular insights and evolving paradigms. Cell 147, 275–292 (2011).
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging biological principles of metastasis. Cell 168, 670–691 (2017).
Strilic, B. & Offermanns, S. Intravascular survival and extravasation of tumor cells. Cancer Cell 32, 282–293 (2017).
Luzzi, K. J. et al. Multistep nature of metastatic inefficiency: dormancy of solitary cells after successful extravasation and limited survival of early micrometastases. Am. J. Pathol. 153, 865–873 (1998).
Massagué, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Pantel, K. & Speicher, M. R. The biology of circulating tumor cells. Oncogene 35, 1216–1224 (2015).
Tsuji, T., Ibaragi, S. & Hu, G. Epithelial-mesenchymal transition and cell cooperativity in metastasis. Cancer Res. 69, 7135–7139 (2009).
Alix-Panabieres, C., Mader, S. & Pantel, K. Epithelial–mesenchymal plasticity in circulating tumor cells. J. Mol. Med. 95, 133–142 (2017).
Green, B. J. et al. Beyond the capture of circulating tumor cells: next-generation devices and materials. Angew. Chem. Int. Ed. 55, 1252–1265 (2016).
Kalluri, R. & Weinberg, R. A. The basics of epithelial–mesenchymal transition. J. Clin. Invest. 119, 1420–1428 (2009).
Bae, Y. K., Choi, J. E., Kang, S. H. & Lee, S. J. Epithelial–mesenchymal transition phenotype is associated with clinicopathological factors that indicate aggressive biological behavior and poor clinical outcomes in invasive breast cancer. J. Breast Cancer 18, 256–263 (2015).
Wu, S. et al. Classification of circulating tumor cells by epithelial–mesenchymal transition markers. PLoS ONE 10, e0123976 (2015).
Yao, D., Dai, C. & Peng, S. Mechanism of the mesenchymal–epithelial transition and its relationship with metastatic tumor formation. Mol. Cancer Res. 9, 1608–1620 (2011).
Ashworth, T. A case of cancer in which cells similar to those in the tumours were seen in the blood after death. Aust. Med. J. 14, 146–149 (1869).
Alexander, R. F. & Spriggs, A. I. The differential diagnosis of tumour cells in circulating blood. J. Clin. Pathol. 13, 414–424 (1960).
Salgado, I. et al. Tumour cells in the blood. Can. Med. Assoc. J. 81, 619–622 (1959).
Racila, E. et al. Detection and characterization of carcinoma cells in the blood. Proc. Natl Acad. Sci. USA 95, 4589–4594 (1998).
Mikolajczyk, S. D. et al. Detection of EpCAM-negative and cytokeratin-negative circulating tumor cells in peripheral blood. J. Oncol. 2011, 252361 (2011).
Pecot, C. V. et al. A novel platform for detection of CK+ and CK– CTCs. Cancer Discov. 1, 580–586 (2011).
Adams, A. et al. Highly efficient circulating tumor cell isolation from whole blood and label-free enumeration using polymer-based microfluidics with an integrated conductivity sensor. J. Am. Chem. Soc. 130, 8633–8641 (2008).
Yoon, H. J. et al. Sensitive capture of circulating tumour cells by functionalized graphene oxide nanosheets. Nat. Nanotech. 8, 735–741 (2013).
Stott, S. L. et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc. Natl Acad. Sci. USA 107, 18392–18397 (2010).
Besant, J. D. et al. Velocity valleys enable efficient capture, sorting and analysis of nanoparticle-bound circulating tumour cells. Nanoscale 7, 6278–6285 (2015).
Scher, H. I. et al. Association of AR-V7 on circulating tumor cells as a treatment-specific biomarker with outcomes and survival in castration-resistant prostate cancer. JAMA Oncol. 2, 1441–1449 (2016).
Antonarakis, E. S. et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N. Engl. J. Med. 371, 1028–1038 (2014).
Ferreira, M. M., Ramani, V. C. & Jeffrey, S. S. Circulating tumor cell technologies. Mol. Oncol. 10, 374–394 (2016).
Yoon, H. J., Kozminsky, M. & Nagrath, S. Emerging role of nanomaterials in circulating tumor cell isolation and analysis. ACS Nano 8, 1995–2017 (2014).
Alix-Panabieres, C. & Pantel, K. Challenges in circulating tumour cell research. Nat. Rev. Cancer 14, 623–631 (2014).
Ozkumur, E. et al. Inertial focusing for tumor antigen-dependent and -independent sorting of rare circulating tumor cells. Sci. Transl. Med. 5, 179ra47 (2013).
Mohamadi, R. M. et al. Nanoparticle-mediated binning and profiling of heterogeneous circulating tumor cell subpopulations. Angew. Chem. Int. Ed. 127, 141–145 (2015).
Poudineh, M. et al. Tracking the dynamics of circulating tumour cell phenotypes using nanoparticle-mediated magnetic ranking. Nat. Nanotech. 12, 274–281 (2017).
Mego, M. et al. Characterization of metastatic breast cancer patients with nondetectable circulating tumor cells. Int. J. Cancer 129, 417–423 (2011).
Hu, X. et al. Marker-specific sorting of rare cells using dielectrophoresis. Proc. Natl Acad. Sci. USA 102, 15757–15761 (2005).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–1239 (2007).
Talasaz, A. H. et al. Isolating highly enriched populations of circulating epithelial cells and other rare cells from blood using a magnetic sweeper device. Proc. Natl Acad. Sci. USA 106, 3970–3975 (2009).
Makker, K., Agarwal, A. & Sharma, R. K. Magnetic activated cell sorting (MACS): utility in assisted reproduction. Indian J. Exp. Biol. 46, 491–497 (2008).
Millner, L. M., Linder, M. W. & Valdes, R. Jr Circulating tumor cells: a review of present methods and the need to identify heterogeneous phenotypes. Ann. Clin. Lab. Sci. 43, 295–304 (2013).
Ibrahim, S. F. & Van Den Engh, G. Flow cytometry and cell sorting. Adv. Biochem. Eng. Biotechnol. 106, 19–39 (2007).
Wu, C.-H. et al. Versatile immunomagnetic nanocarrier platform for capturing cancer cells. ACS Nano 7, 8816–8823 (2013).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Tsai, J. H. & Yang, J. Epithelial–mesenchymal plasticity in carcinoma metastasis. Genes Dev. 27, 2192–2206 (2013).
Tellez-Gabriel, M., Brown, H. K., Young, R., Heymann, M.-F. & Heymann, D. The challenges of detecting circulating tumor cells in sarcoma. Front. Oncol. 6, 202–210 (2016).
Gabriel, M. T., Calleja, L. R., Chalopin, A., Ory, B. & Heymann, D. Circulating tumor cells: a review of non-EpCAM-based approaches for cell enrichment and isolation. Clin. Chem. 62, 571–581 (2016).
Wit, Sde et al. The detection of EpCAM+ and EpCAM– circulating tumor cells. Sci. Rep. 5, 12270 (2015).
Zhang, L. et al. The identification and characterization of breast cancer CTCs competent for brain metastasis. Sci. Transl. Med. 5, 180ra48 (2013).
Baccelli, I. et al. Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nat. Biotechnol. 31, 539–544 (2013).
Park, M. H. et al. Enhanced isolation and release of circulating tumor cells using nanoparticle binding and ligand exchange in a microfluidic chip. J. Am. Chem. Soc. 139, 2741–2749 (2017).
Che, J. et al. Classification of large circulating tumor cells isolated with ultra-high throughput microfluidic Vortex technology. Oncotarget 7, 12748–12760 (2016).
Hvichia, G. E. et al. A novel microfluidic platform for size and deformability based separation and the subsequent molecular characterization of viable circulating tumor cells. Int. J. Cancer 138, 2894–2904 (2016).
Kaldjian, E. et al. Multi-level analysis of circulating tumor cells in advanced prostate cancer using AccuCyte® – CyteFinder®. Paper presented at 22nd Annual Prostate Cancer Foundation Scientific Retreat 8–10 October (2015); http://rarecyte.com/documents/1465944263.pdf
Demierre, N., Braschler, T., Muller, R. & Renaud, P. Focusing and continuous separation of cells in a microfluidic device using lateral dielectrophoresis. Sens. Actuators B Chem. 132, 388–396 (2008).
Ng, S. Y. et al. Label-free impedance detection of low levels of circulating endothelial progenitor cells for point-of-care diagnosis. Biosens. Bioelectron. 25, 1095–1101 (2010).
Jin, C. et al. Technologies for label-free separation of circulating tumor cells: from historical foundations to recent developments. Lab Chip 14, 32–44 (2014).
Abonnenc, M. et al. Programmable interactions of functionalized single bioparticles in a dielectrophoresis-based microarray chip. Anal. Chem. 85, 8219–8224 (2013).
Manaresi, N. et al. A CMOS chip for individual cell manipulation and detection. IEEE J. Solid-State Circuits 38, 2297–2305 (2003).
Werner, S. L. et al. Analytical validation and capabilities of the Epic CTC platform: enrichment-free circulating tumour cell detection and characterization. J. Circ. Biomark. 4, 1 (2015).
Kowalik, A., Kowalewska, M. & Góźdź, S. Current approaches for avoiding the limitations of circulating tumor cells detection methods — implications for diagnosis and treatment of patients with solid tumors. Transl. Res. 185, 58–84.e15 (2017).
Halo, T. L. et al. Nanoflares for the detection, isolation, and culture of live tumor cells from human blood. Proc. Natl Acad. Sci. USA 111, 17104–17109 (2014).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2012).
Balkwill, F. Cancer and the chemokine network. Nat. Rev. Cancer 4, 540–550 (2004).
Kakinuma, T. & Hwang, S. T. Chemokines, chemokine receptors, and cancer metastasis. J. Leukoc. Biol. 79, 639–651 (2006).
Chow, M. T. & Luster, A. D. Chemokines in cancer. Cancer Immunol. Res. 2, 1125–1131 (2014).
Wong, I. Y. et al. Collective and individual migration following the epithelial-mesenchymal transition. Nat. Mater. 13, 1063–1071 (2014).
Zhang, Y., Zhang, W. & Qin, L. Mesenchymal-mode migration assay and antimetastatic drug screening with high-throughput microfluidic channel networks. Angew. Chem. Int. Ed. 126, 2376–2380 (2014).
Roussos, E. T., Condeelis, J. S. & Antonia, P. Chemotaxis in cancer. Nat. Rev. Cancer 11, 573–587 (2011).
Chen, Y.-C. et al. Single-cell migration chip for chemotaxis-based microfluidic selection of heterogeneous cell populations. Sci. Rep. 5, 9980 (2015).
Poudineh, M. et al. Profiling functional and biochemical phenotypes of circulating tumor cells using a two-dimensional sorting device. Angew. Chem. Int. Ed. 56, 163–168 (2017).
Zijlstra, A., Lewis, J., DeGryse, B., Stuhlmann, H. & Quigley, J. P. The inhibition of tumor cell invasation and subsequent metastasis through the regulation of in vivo tumor cell motility by the tetrspanin CD151. Cancer Cell 13, 221–234 (2011).
Reymond, N., d’Água, B. B. & Ridley, A. J. Crossing the endothelial barrier during metastasis. Nat. Rev. Cancer 13, 858–870 (2013).
William, J., Zervantonakis, I. K., Roger, D. & Link, C. Tumor cell migration in complex microenvironments. Cell. Mol. Life Sci. 70, 1335–1356 (2013).
Zervantonakis, I. K. et al. Three-dimensional microfluidic model for tumor cell intravasation and endothelial barrier function. Proc. Natl Acad. Sci. USA 109, 13515–13520 (2012).
Jeon, J. S., Zervantonakis, I. K., Chung, S., Kamm, R. D. & Charest, J. L. In vitro model of tumor cell extravasation. PLoS ONE 8, e56910 (2013).
Yee, S. S. et al. A novel approach for next-generation sequencing of circulating tumor cells. Mol. Genet. Genom. Med. 4, 395–406 (2016).
De Luca, F. et al. Mutational analysis of single circulating tumor cells by next generation sequencing in metastatic breast cancer. Oncotarget 7, 26107–26119 (2016).
Krebs, M. G. et al. Molecular analysis of circulating tumour cells — biology and biomarkers. Nat. Rev. Clin. Oncol. 11, 129–144 (2014).
Carter, L. et al. Molecular analysis of circulating tumor cells identifies distinct copy-number profiles in patients with chemosensitive and chemorefractory small-cell lung cancer. Nat. Med. 23, 114–119 (2016).
Ramirez, J.-M. et al. Prognostic relevance of viable circulating tumor cells detected by EPISPOT in metastatic breast cancer patients. Clin. Chem. 60, 214–221 (2014).
Alix-Panabières, C. & Pantel, K. Clinical prospects of liquid biopsies. Nat. Biomed. Eng. 1, 0065 (2017).
Denève, E. et al. Capture of viable circulating tumor cells in the liver of colorectal cancer patients. Clin. Chem. 59, 1384–1392 (2013).
Alix-Panabie’res, C. et al. Detection and characterization of putative metastatic precursor cells in cancer patients. Clin. Chem. 53, 536–537 (2007).
Sinkala, E. et al. Profiling protein expression in circulating tumour cells using microfluidic western blotting. Nat. Commun. 8, 14622 (2017).
Han, T. et al. How does cancer cell metabolism affect tumor migration and invasion? Cell Adh. Migr. 1, 395–403 (2013).
Zhang, Y. et al. Single-cell codetection of metabolic activity, intracellular functional proteins, and genetic mutations from rare circulating tumor cells. Anal. Chem. 87, 9761–9768 (2015).
Phan, L. M., Yeung, S.-C. J. & Lee, M.-H. Cancer metabolic reprogramming: importance, main features, and potentials for precise targeted anti-cancer therapies. Cancer Biol. Med. 11, 1–19 (2014).
Gialeli, C., Theocharis, A. D. & Karamanos, N. K. Roles of matrix metalloproteinases in cancer progression and their pharmacological targeting. FEBS J. 278, 16–27 (2011).
Ren, F. et al. Overexpression of MMP family members functions as prognostic biomarker for breast cancer patients: a systematic review and meta-analysis. PLoS ONE 10, e0135544 (2015).
Son, K. J., Shin, D. S., Kwa, T., Gao, Y. & Revzin, A. Micropatterned sensing hydrogels integrated with reconfigurable microfluidics for detecting protease release from cells. Anal. Chem. 85, 11893–11901 (2013).
Yang, G., Li, L., Rana, R. K. & Zhu, J. J. Assembled gold nanoparticles on nitrogen-doped graphene for ultrasensitive electrochemical detection of matrix metalloproteinase. Carbon NY 61, 357–366 (2013).
Timm, K. N., Kennedy, B. W. C. & Brindle, K. M. Imaging tumor metabolism to assess disease progression and treatment response. Clin. Cancer Res. 22, 5196–5203 (2016).
Andree, K. C., van Dalum, G. & Terstappen, L. W. M. M. Challenges in circulating tumor cell detection by the CellSearch system. Mol. Oncol. 10, 395–407 (2016).
Stoecklein, N. H., Fischer, J. C., Niederacher, D. & Terstappen, L. W. M. M. Challenges for CTC-based liquid biopsies: low CTC frequency and diagnostic leukapheresis as a potential solution. Expert Rev. Mol. Diagn. 16, 147–164 (2016).
Aceto, N. et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158, 1110–1122 (2014).
Fidler, I. J. Immune stimualtion–inhibition of experimental cancer metastasis. Cancer Res. 34, 491–498 (1974).
Hong, Y., Fang, F. & Zhang, Q. Circulating tumor cell clusters: what we know and what we expect (Review). Int. J. Oncol. 49, 2206–2216 (2016).
Sharma, D., Brummel-Ziedins, K. E., Bouchard, B. A. & Holmes, C. E. Platelets in tumor progression: a host factor that offers multiple potential targets in the treatment of cancer. J. Cell. Physiol. 229, 1005–1015 (2014).
Sarioglu, A. F. et al. A microfluidic device for label-free, physical capture of circulating tumor cell clusters. Nat. Methods 12, 685–691 (2015).
Au, S. H. et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proc. Natl Acad. Sci. USA 113, 4947–4952 (2016).
Gkountela, S. & Aceto, N. Stem-like features of cancer cells on their way to metastasis. Biol. Direct 11, 33 (2016).
Yu, M. et al. RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis. Nature 487, 510–513 (2012).
Perakis, S. & Speicher, M. R. Emerging concepts in liquid biopsies. BMC Med. 15, 75 (2017).
Ma, M. et al. ‘Liquid biopsy’ — ctDNA detection with great potential and challenges. Ann. Transl. Med 16, 235 (2015).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Keller, S., Ridinger, J., Rupp, A.-K., Janssen, J. W. & Altevogt, P. Body fluid derived exosomes as a novel template for clinical diagnostics. J. Transl. Med. 9, 86 (2011).
Shao, H., Chung, J. & Issadore, D. Diagnostic technologies for circulating tumour cells and exosomes. Biosci. Rep. 36, e00292 (2016).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Whiteside, T. L. The potential of tumor-derived exosomes for noninvasive cancer monitoring. Expert Rev. Mol. Diagn. 15, 1293–1310 (2015).
Hoshino, A. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015).
Im, H. et al. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat. Biotechnol. 32, 490–495 (2014).
Zhou, Y. G. et al. Interrogating circulating microsomes and exosomes using metal nanoparticles. Small 12, 727–732 (2016).
Sharma, S., Gillespie, B. M., Palanisamy, V. & Gimzewski, J. K. Quantitative nanostructural and single-molecule force spectroscopy biomolecular analysis of human-saliva-derived exosomes. Langmuir 27, 14394–14400 (2011).
Malloy, A. & Carr, B. Nanoparticle tracking analysis — the halo system. Part. Part. Syst. Charact. 23, 197–204 (2006).
Wunsch, B. H. et al. Nanoscale lateral displacement arrays for the separation of exosomes and colloids down to 20 nm. Nat. Nanotech. 11, 936–940 (2016).
Alix-Panabières, C. & Pantel, K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 6, 479–491 (2016).
Alix-Panabières, C. & Pantel, K. Real-time liquid biopsy: circulating tumor cells versus circulating tumor DNA. Ann. Transl. Med. 1, 18 (2013).
Madic, J. et al. Circulating tumor DNA and circulating tumor cells in metastatic triple negative breast cancer patients. Int. J. Cancer 136, 2158–2165 (2015).
Bardelli, A. & Pantel, K. Liquid biopsies, what we do not know (yet). Cancer Cell 31, 172–179 (2017).
Pantel, K. Blood-based analysis of circulating cell-free DNA and tumor cells for early cancer detection. PLoS Med. 13, e1002205 (2016).
Fernandez-Cuesta, L. et al. Identification of circulating tumor DNA for the early detection of mmall-cell lung cancer. EBioMedicine 10, 117–123 (2016).
Dawson, S.-J. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 368, 1199–1209 (2013).
Francis, G. & Stein, S. Circulating cell-free tumour DNA in the management of cancer. Int. J. Mol. Sci. 16, 14122–14142 (2015).
Olsson, E. et al. Serial monitoring of circulating tumor DNA in patients with primary breast cancer for detection of occult metastatic disease. EMBO Mol. Med. 7, 1034–1047 (2015).
Siravegna, G. & Bardelli, A. Genotyping cell-free tumor DNA in the blood to detect residual disease and drug resistance. Genome Biol. 15, 449 (2014).
Gold, B., Cankovic, M., Furtado, L. V., Meier, F. & Gocke, C. D. Do circulating tumor cells, exosomes, and circulating tumor nucleic acids have clinical utility?: A report of the association for molecular pathology. J. Mol. Diagn. 17, 209–224 (2015).
Han, X., Wang, J. & Sun, Y. Circulating tumor DNA as biomarkers for cancer detection. Genom. Proteom. Bioinforma. 15, 59–72 (2017).
Bettegowda., C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra24 (2014).
Newman, A. M. et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat. Biotechnol. 34, 547–555 (2016).
Gorges, T. M. et al. Accession of tumor heterogeneity by multiplex transcriptome profiling of single circulating tumor cells. Clin. Chem. 62, 1504–1515 (2016).
Acknowledgements
S.O.K. acknowledges the Canadian Institutes of Health Research, the National Science and Engineering Research Council, the National Institutes of Health, and the Ontario Research Fund for their support of work in this area. K.P. acknowledges European Research Council Advanced Investigator Grant DISSECT, European Research Council Proof of Concept grant CTCapture_2.0 and European Union–Innovative Medicines Initiative grant CANCER-ID for support.
Author information
Authors and Affiliations
Contributions
All authors contributed to compiling content for this work, and to writing and editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.O.K., M.P. and E.H.S. are inventors on patents related to technologies for liquid biopsy and rare-cell profiling. The other authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Poudineh, M., Sargent, E.H., Pantel, K. et al. Profiling circulating tumour cells and other biomarkers of invasive cancers. Nat Biomed Eng 2, 72–84 (2018). https://doi.org/10.1038/s41551-018-0190-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0190-5
This article is cited by
-
Capturing nascent extracellular vesicles by metabolic glycan labeling-assisted microfluidics
Nature Communications (2023)
-
CdSe@CdS quantum dot–sensitized Au/α-Fe2O3 structure for photoelectrochemical detection of circulating tumor cells
Microchimica Acta (2023)
-
Peptide nano-blanket impedes fibroblasts activation and subsequent formation of pre-metastatic niche
Nature Communications (2022)
-
Cancer cells spread aggressively during sleep
Nature (2022)
-
Integrated microfluidic single-cell immunoblotting chip enables high-throughput isolation, enrichment and direct protein analysis of circulating tumor cells
Microsystems & Nanoengineering (2022)