Abstract
Selfing is a frequent evolutionary trend in angiosperms, and is a suitable model for studying the recurrent patterns underlying adaptive evolution. Many plants avoid self-fertilization by physiological processes referred to as self-incompatibility (SI). In the Brassicaceae, direct and specific interactions between the male ligand SP11/SCR and the female receptor kinase SRK are required for the SI response. Although Arabidopsis thaliana acquired autogamy through loss of these genes, molecular evolution contributed to the spread of self-compatibility alleles requires further investigation. We show here that in this species, dominant SRK silencing genes have evolved at least twice. Different inverted repeat sequences were found in the relic SRK region of the Col-0 and C24 strains. Both types of inverted repeats suppress the functional SRK sequence in a dominant fashion with different target specificities. It is possible that these dominant suppressors of SI contributed to the rapid fixation of self-compatibility in A. thaliana.
Similar content being viewed by others
Introduction
Self-incompatibility (SI) refers to mechanisms that prevent self-fertilization and promote outcrossing between different individuals adopted by over 40% of flowering plant species1. In the Brassicaceae family, SI is controlled by two multi-allelic genes: the S-locus protein 11 (SP11; also known as the S-locus cysteine-rich protein, SCR) and the S-locus receptor kinase (SRK)2,3. SP11/SCR is a peptide ligand localized to the pollen coat surface4,5, and SRK is a stigma surface receptor6. Proteins translated from a tightly linked pair (haplotype) of SP11/SCR and SRK directly and specifically bind to one another, triggering a self-pollen rejection signal in the stigma7.
On the other hand, the evolution of self-compatibility (SC) is one of the most frequently occurred convergent evolution known in plants8. In the model plant species Arabidopsis thaliana, SC is considered to have evolved at least three times independently because three major haplogroups (A, B, and C) derived from a much larger set of diverged S haplotypes in the closely related outcrossers, such as A. halleri, are found9. A previous study further classified the alleles into 12 haplotypes10. Strains such as Col-0, Uk-3 and Wei-1 belong to haplogroup A in which Col-0 carries a pseudo SRK and a non-functional SP11/SCR, whereas Uk-3 and Wei-1 carry functional SRK genes11. The inversion of SP11/SCR is considered to be the primary mutation disrupting the SI of this haplogroup11. The Cvi-0 strain belongs to haplogroup B in which a premature stop codon is found in SRK9. Strains such as C24 belong to haplotype R2 formed by recombination between the A and C haplogroups, followed by large deletions of both SRKA and SRKC10. Notwithstanding these facts, molecular forces contributed to the spread of SC in this species still requires understanding.
Recently, some laboratory observations that provide support for this question have been reported. Although all known A. thaliana strains are self-compatible, SI can be reconstituted in this species when functional forms of SP11/SCR and/or SRK are introduced2,11,12. These findings indicate that at least in some strains, non-S-locus components required to express SI are still retained; however, there has been some controversy over the requirements for genetic factors to express the female SI-phenotype13,14. When the Col-0 strain was transformed with the Sb-haplotype, SRKb and SCRb genes from A. lyrata (an allele corresponding to SRK20 (ref. 15), the transgenic plants showed only weak or transient SI phenotypes12. This transient SI phenotype was evident in stigmas at flower stage 13 (at anthesis) but was attenuated in later stages. On the other hand, robust SI was expressed regardless of the flower stage in other A. thaliana strains such as C24, when transformed with the same genes. Liu et al. reported that this phenotypic variation is due to natural variation in the S-locus-linked PUB8 gene, which encodes an ARM-repeat U-box-containing protein16. In contrast, Indoriolo et al. reported that another ARM-repeat U-box-containing protein, ARC1, is required to express strong SI in the Col-0 and Sha strains13. ARC1 is involved in the expression of SI in Brassica species17, and the ARC1 gene has been independently deleted in many self-compatible species of this genus18. The discrepancies among these reports have not yet been resolved. Identifying the prerequisites for reconstituting SI in A. thaliana directly relates to solving how SC evolved in this species.
Through the process of identifying the differences between Col-0 and C24 in this study, we found that an inverted repeat sequence (SRKIRCol-0) in the relic SRKA region of Col-0 suppresses SRK mRNA accumulation by different haplotypes. Interference by the small RNAs possibly produced from SRKIRCol-0 can explain the instability of the reconstituted SI phenotype in this strain. Furthermore, we found a distinct inverted repeat in C24 (SRKIRC24) that may also suppress the expression of allelic SRK genes with a target specificity different from SRKIRCol-0. These two sequence structures evolved independently in A. thaliana. We discuss the possible contributions of these dominant suppressors in the fixation of SC in A. thaliana. Our results may also support the idea that inverted repeats are a recurrent evolutionary form of allelic dominance.
Results
The Col-0 strain may carry a dominant suppressor of SRK mRNA
To confirm the instability of the SI phenotype in Col-0, we generated a transgenic line carrying SRKb and SCRb that were cloned from the A. lyrata Sb (S20) haplotype (Col-0 + SRKb-SCRb). In a previous study, we produced a C24 transgenic line carrying single copies of SRKb and SCRb (called SI-C24 in this study for brevity)19. SI-C24 exhibited SI at both flower stages 13 and 14 (Supplementary Fig. 1). Col-0 + SRKb-SCRb pistils at stage 13 were only partially able to reject pollen from SI-C24 flowers, and at stage 14 they failed to reject (Supplementary Fig. 1). These observations were in agreement with the previous results of Nasrallah et al.12.
When Col-0 + SRKb-SCRb was used as the pollen donor with SI-C24 pistils from flower stage 14, virtually no pollen tube growth was observed (Fig. 1a, b). Therefore, Col-0 was able to express the SCRb gene and show the male side of the SI phenotype but was unable to express the female side of the SI phenotype. The F1 generation of SI-C24 and Col-0 failed to express a strong female SI phenotype at stage 14 (Fig. 1b) and the SRKb mRNA levels were significantly reduced (Fig. 1c). We, therefore, hypothesized that Col-0 carries a dominant endogenous factor, not present in C24, that suppresses the expression of SRKb. This was not in agreement with the previous study by Liu et al. reported that the weak SI expression phenotype is recessive16.

a Representative fluorescent images of aniline blue-stained pistils at stage 14 three hours after pollination. Scale bars = 100 µm. b Summary of pollination tests. S × C F1 is the hybrid obtained from the cross: ♀SI-C24 × ♂Col-0. Compatibility phenotypes were scored as described in the methods. Statistically significant differences based on Dunnett’s test are indicated by ***(p < 0.005), with the exact p-value being 7.16e−06. Error bars indicate standard deviations. The number of pistils pollinated (from left to right): n = 8, n = 8, n = 8, n = 4, n = 3, n = 7. c Relative mRNA accumulation levels of SRKb compared to ACTIN8 (ACT8) in stigmatic tissues of the indicated lines. Two independent S × C F1 individuals (#1 and #2) were analyzed. Error bars indicate standard deviations from independent RNA extraction trials (n = 3). d Illustration comparing the S loci of A. thaliana Col-0 and C24, and the A. halleri S4 haplotype. Col-0 carries an inverted repeat with homology to the sequence encoding the kinase domain of SRK. The C24 strain carries an inverted repeat with homology to the sequence encoding the ecto-domain of SRK. e A histogram showing the numbers of small RNAs from stigmas that map to the pseudogenized SRKA gene (ψSRKA) of Col-0; most small RNAs map to the inverted repeat region (see Supplementary Fig. 2a for the mapping results for the flanking regions).
Small RNAs are possibly produced from SRKIR Col-0
Col-0 carries a relic S locus sequence belonging to the A haplogroup (At4g21370: ψSRKA). From the available genomic information archived in The Arabidopsis Information Resource (TAIR), we found an inverted repeat structure within the Col-0 allele of ψSRKA and named this structure SRKIRCol-0. SRKIRCol-0 consists of a 502-bp inverted repeat, which corresponds to the fragment encoding the SRK kinase domain in the orthologous SRK sequence from the A. halleri S4 haplotype (Fig. 1d). Such a hairpin RNA sequence can form a duplex-RNA and could be processed into small RNAs by Dicers20. We suspected that SRKIRCol-0 serves as the source of small RNAs that act to suppress expression of the introduced SRKb. Consistent with this hypothesis, a transcriptome analysis revealed that 25,671 small RNA reads from Col-0 stigmas mapped to the SRKIRCol-0 region (Fig. 1e, Supplementary Fig. 2a). Over 99.5% of the reads (25,563) were uniquely mapped to the SRKIRCol-0 region; thus, we considered that these small RNAs were likely produced from this hairpin. The majority of these small RNAs had lengths of 20 or 21 nucleotides (Supplementary Fig. 2b). SRKIRCol-0 showed high sequence homology with the DNA sequence encoding the SRK kinase domain in SRKb. In particular, the fragment between base-pairs 102 to 253 of the SRKIRCol-0 sequence was 89.0% identical to the corresponding region of the SRKb gene (Supplementary Fig. 3). We used the psRNAtarget program21 to predict if these small RNAs that mapped to SRKIRCol-0 can target SRKb. As a result, 58 non-redundant reads perfectly matched (expectation = 0 in the psRNAtarget program) within the kinase domain of SRKb (Supplementary Fig. 4), suggesting that these small RNAs found within the SRKIRCol-0 sequence may potentially target SRKb.
SRKIR Col-0 suppresses SRKb expression in a dominant fashion
To show whether SRKIRCol-0 could suppress SRKb, we introduced a 5,353-bp genomic fragment containing the SRKIRCol-0 sequence (full SRKIRCol-0) into the SI-C24 line (Fig. 2a). In a previous study, we showed that the promoter of ψSRKA is actively transcribed in stigmas19. As a result, the SI phenotype observed in the SI-C24 line at stage 14 was compromised in the transformants (Fig. 2b–d). This observation further indicated that the SRKIRCol-0 sequence can suppress SRKb in a dominant fashion. When we introduced a partial fragment that contains only the first repeat of the SRKIRCol-0 sequence (partial SRKIRCol-0), this fragment was unable to suppress the function of SRKb (Fig. 2b–d).

a Schematic drawing of the ψSRKA locus of Col-0. b Representative fluorescent images of aniline blue-stained pistils at stage 14 three hours after pollination. #1 and #2 indicate pistils from two independent first generation transformants. Scale bars = 100 µm. c Summary of pollination tests. Statistically significant differences based on Dunnett’s test are indicated by ***(p < 0.005). The exact p-values found by Dunnett’s test compared to the ♀SI-C24 × ♂SI-C24 cross were 4.66e−07 and 1.84e−05, from left to right. Error bars indicate standard deviations. The number of pistils pollinated (from left to right): n = 8, n = 8, n = 4, n = 3, n = 6. d Relative mRNA accumulation levels of SRKb compared to ACT8 in stigmatic tissues of the indicated lines. Error bars indicate standard deviations from independent RNA extraction trials (n = 3).
We also investigated the effect of a T-DNA insertion in SRKIRCol-0. Compared to the individual that did not carry the T-DNA insertion (−/−), SRKb mRNA accumulation was recovered in the line carrying the T-DNA insertion homozygously (+/+) (Supplementary Fig 5b). These results suggested that the continuous inverted repeat is probably required for SRKIRCol-0 to function in compromising SI. Taken together, we showed that full SRKIRCol-0 is necessary and is likely to be sufficient to cause breakdown of SI in the C24 background.
SRKIR Col-0 suppresses the expression of endogenous SRKs
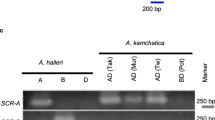
We next tested whether the SRKIRCol-0 sequence could suppress expression of the endogenous SRKb gene. To achieve this objective, we generated inter-specific hybrids of A. thaliana and A. lyrata. We crossed A. thaliana Col-0 and C24 with an individual of A. lyrata carrying the Sa (intermediate dominance class22) and Sb (most dominant class22) haplotypes. Pistils carrying the Sb allele from the A. lyrata parent and the C24 genotype from the A. thaliana parent were incompatible with pollen from the SaSb A. lyrata individual (Fig. 3a,b). It should be noted that pollen from SaSb A. lyrata shows only Sb specificity because Sb is dominant over Sa for SCR expression23. In contrast, pistils with the Sb allele from A. lyrata and the Col-0 genotype from A. thaliana were compatible with SaSb pollen (Fig. 3a, b). We also found that the levels of SRKb mRNA were reduced by 6.5-fold in the Col-0/Sb stigmas when compared with the A. lyrata SaSb stigmas, and this was not the case with the C24/Sb stigmas (Fig. 3c). Therefore, SRKb expression was suppressed in the stigmas with the Col-0 genetic background.

a Representative fluorescent images of aniline blue-stained pistils at stage 14 of A. thaliana–lyrata inter-species hybrids with different genotypes 3 h after pollination. C24/Sa and C24/Sb are individuals obtained from the cross A. lyrata SaSb × C24 that inherited the Sa and Sb alleles, respectively. Similarly, Col-0/Sa and Col-0/Sb are A. lyrata SaSb x Col-0 hybrids that inherited the Sa and Sb alleles, respectively. Pollen from an A. lyrata SaSb individual was used for the pollination test. Scale bars = 100 µm. b Summary of pollination tests. A statistically significant difference between C24/Sb and Col-0/Sb (***p < 0.005) was detected using Dunnett’s test, with the exact p-value being 0.00e+00. Error bars indicate standard deviations. Number of pistils pollinated (from left to right): n = 8, n = 6, n = 7, n = 8. c Relative mRNA accumulation levels of SRKb compared to the A. lyrata ACTIN8 ortholog in stigmatic tissues of the indicated lines. Error bars indicate standard deviations from independent RNA extraction trials (n = 4). d, e Relative mRNA accumulation levels of the SRK genes compared to ACT8 in stigmatic tissues of the indicated lines. For each hybrid cross, two individuals (#1 and #2) were tested. In c–e error bars indicate standard deviations from independent RNA extraction trials (n = 3). The broken lines indicate the expected mRNA accumulation level of the SRK genes in the hybrids compared to that of the parents (theoretically one-half). Dunnett’s tests were used to find significant differences (***p < 0.005) compared to the expected mRNA level. d The exact p-values from left to right: 5.02e−13, 2.22e−16, 0.00e+00, 0.00e+00, 1.17e−06, 1.05e−04, 7.71e−12, and 8.88e−16. e The exact p values from left to right; 1.63e−05, 9.15e−05, 2.00e−15, and 7.77e−16.
In some A. thaliana strains, including Uk-3 and Wei-1, SRK function is retained11; both strains carry an intact SRK gene that belongs to haplogroup A (SRKA), and Wei-1 can reject pollen from A.halleri carrying SCRA11. The SRK sequence in the Cvi-0 strain belongs to the B haplotype and is a pseudogene (ψSRKBCvi-0) but is transcribed in flowers9. We expected that expression of the SRK genes from these strains would be suppressed in the presence of SRKIRCol-0 because of their high sequence identities with SRKIRCol-0 (Supplementary Fig. 6). The psRNAtarget program also predicted that 1,023 and 91 non-redundant small RNA reads found within the SRKIRCol-0 sequence matched perfectly within the kinase domains of SRKAWei-1 and ψSRKBCvi-0, respectively (Supplementary Fig. 4). To verify this finding, we crossed these strains with Col-0 and analyzed the SRK mRNA levels in the F1 generations. Similar to the SI-C24 x Col-0 F1 hybrids (Fig. 1c), SRK expression in the Uk-3 x Col-0, Wei-1 x Col-0, and Cvi-0 x Col-0 F1 hybrids was significantly reduced when compared with the expected levels of transcript accumulation in the hybrids (Fig. 3d, e). This result indicated a broad function of SRKIRCol-0 in SRK suppression.
A putative inverted repeat structure in S locus of C24
Since C24 can successfully express externally introduced SRKb (Fig. 1), the reduction in SRK mRNA levels in the Uk-3 × C24, Wei-1 × C24, and Cvi-0 × C24 F1 hybrids when compared with the parental Uk-3, Wei-1, and Cvi-0 strains was unexpected (Fig. 3d, e). C24 carries a relic S locus belonging to haplotype R2 that carries small fragments of SRKA and SRKC10. We rechecked the bacterial artificial chromosome sequence carrying the relic S-locus in C24, reported in a previous study24, and found a 434-bp putative inverted repeat sequence within the pseudo SRKA region (Supplementary Fig. 7a). We named the sequence SRKIRC24. Using the publicly available small RNA transcriptome dataset for the C24 strain25, we detected 3235 reads that mapped within the SRKIRC24 sequence in flower buds (Supplementary Fig. 7b). SRKIRC24 shared 99.7% sequence identity with SRKAWei-1 and 84.4% sequence identity with and ψSRKBCvi-0 (Supplementary Fig. 8). On the other hand, the DNA sequence identity between SRKIRC24 and SRKb was limited to 73.1%. The psRNAtarget prediction found that 329 and 20 non-redundant small RNA reads detected in SRKIRC24 matched perfectly to the kinase domains of SRKAWei-1 and ψSRKBCvi-0, respectively (Supplementary Fig. 4). In contrast, none of these reads perfectly matched SRKb (Supplementary Fig. 4a), suggesting that these small RNAs may not target SRKb. This finding is in line with our experimental results showing that SRKb can be stably expressed in the C24 background12,19 (Fig. 1).
A codon substituted SRKb is not suppressed by SRKIR Col-0
The above studies suggested involvement of the sequence complementarity-based small RNA silencing pathway in the suppression of SRKb in Col-0. Therefore, we generated a synthetic SRKb sequence with synonymous substitutions in the sequence encoding the kinase domain (synSRKb) without altering its protein sequence and introduced this synthetic SRKb sequence into Col-0 (Fig. 4a). In contrast with the native SRKb, which shared 89.0% sequence identity with SRKIRCol-0, the sequence identity between synSRKb and SRKIRCol-0 was 66.6% (Supplementary Fig. 3). The synonymous substitutions almost completely removed the predicted targetability of the small RNAs found in SRKIRCol-0 when examined in synSRKb (Supplementary Fig. 4). We hypothesized that this reduced targetability would decrease the likelihood of silencing. Lines carrying the synSRKb sequence exhibited full strength SI phenotypes similar to SI-C24 (Fig. 4b, c). The levels of SRKb mRNA were not suppressed in these lines, unlike those transformed with the native SRKb gene (Fig. 4d). These data suggest that the suppression of SRKb by SRKIRCol-0 requires shared sequence homology between the target and the small RNAs produced.

a The artificially constructed SynSRKb carries synonymous codon substitutions and shares 66.0% identity with SRKIRCol-0, whereas the native SRKb shares 89.0% identity with SRKIRCol-0. b Representative fluorescent images of aniline blue-stained pistils three hours after pollination. #1 and #2 indicate samples from two independent first generation transformants. Scale bars = 100 µm. c Summary of pollination tests. Statistically significant differences based on Dunnett’s test are indicated by ***(p < 0.005). The exact p-values were 8.55e−11 and 1.91e−11, from left to right. Error bars indicate standard deviations. Number of pistils pollinated (from left to right): n = 8 n = 11, n = 9, n = 8. d Relative mRNA accumulation levels of the SRK genes compared to ACT8 in stigmatic tissues of the indicated lines. Error bars indicate standard deviations from independent RNA extraction trials (n = 3).
Discussion
In this study, we explored the involvement of inverted repeats in relic S-loci of A. thaliana in the suppression of SRK mRNA accumulation. We showed that SRKIRCol-0 can directly suppress mRNA accumulation from the introduced SRKb transgene, and that this reduced transcript abundance may explain the weak reconstitution of the SI phenotype in the Col-0 strain12. From the finding that the promoter of ψSRKA is actively transcribed in stigma19, it is likely that this inverted repeat is transcribed by an RNA polymerase II as is characteristic of naturally occurring hairpin RNAs20. It is possible that the RNA structure of SRKIRCol-0 is processed into small RNAs by a pathway previously described in Drosophila20 for naturally occurring hairpin RNAs. Further studies are necessary to understand how small RNAs are produced from SRKIRCol-0.
Furthermore, we noticed a putative inverted repeat structure, SRKIRC24, in the relic S-locus of the C24 strain. As is the case for SRKIRCol-0, SRKIRC24 may act directly to suppress mRNA accumulation from homologous SRK sequences in a dominant fashion. The finding that SRKIRCol-0 was able to suppress all SRK alleles investigated in this study, whereas SRKIRC24 was not able to suppress SRKb may be attributable to the location of the inverted structure. SRKIRCol-0 corresponds to a fragment encoding the kinase domain, which is relatively well conserved within the SRK sequence. SRKIRC24 was located within the variable ecto-domain, suggesting that this structure may not have the ability to suppress a broad range of diverse SRK. Based on these results, we propose that in silico prediction of SRK alleles that can express its function when introduced into Col-0 or C24 is possible.
According to a previous study, sequence structures of the SRKA regions among the A1, R0, and R1 haplotypes and those for the A2 and R2 haplotypes are largely shared10. Therefore, from a rough estimation, we propose that about 80.3% of A. thaliana strains (the total of A1, A2, R0, R1, and R2 haplotypes) carry either one of these two inverted repeat types (SRKIRCol-0 or SRKIRC24) based on a survey of the S-locus sequences in 1083 strains10. Several studies suggest that the initial step in the evolution of self-compatibility in the A haplotype of A. thaliana was a loss-of-function of the SP11/SCR gene because functional forms of this gene are absent from all strains investigated8,10,11.
Whether the dominant SRK suppressors played a role in the evolution of SC in A. thaliana is an intriguing question. In the genus Brassica, S-locus-linked small non-coding RNA genes in the dominant alleles silence mRNA expression of the recessive SP11/SCR26,27. Similarly, several SP11/SCR dominance classes are predicted in species such as A. lyrata22,28 and A. halleri29. Further study of the A. halleri system found that there are at least 17 small RNA-producing loci in the region that contribute to the establishment of dominance relationships30. It has been argued that gene-disrupting mutations of SP11/SCR or SRK in dominant S-haplogroups are predicted to be more likely spread than recessive mutations when SC is favored8,31. It is possible that in certain climatic situations such as the glacial periods when pollen availability is limited11,32, such a dominant SRK suppressor mutation may spread throughout the population faster than a recessive mutation. More theoretical studies on the evolution of SC by incorporating the allelic dominance effect would be worthwhile using existing frameworks32,33.
The fact that SRKIRCol-0 and SRKIRC24 independently evolved twice in the evolution of A. thaliana is reminiscent of the classical example of the spread of a dominant phenotype: the melanization of the peppered moth (Biston betularia) during the Industrial Revolution34. The melanic phenotype is dominant over other color types and also independently evolved twice in this species35. Our results support the model that inverted repeats are an easy way to establish an allelic dominance relationship. It is possible that inverted repeat structures may explain other dominant-recessive genetic systems.
Methods
Plant materials
All plant materials were grown in mixed soil in a growth chamber under controlled conditions (light intensity, 120–150 µmol m−2 s−1; 14-h light/10-h dark cycle; 22 ± 2 °C). The T-DNA insertion line of SRKIRCol-0 (SALK_137645C) was obtained from the Arabidopsis Biological Resource Center. A. lyrata SaSb seeds were gifts from Dr. Yoshinobu Takada, Tohoku University.
Transgenic experiments
The SRKb-SCRb/pBI121 construct was produced by the below procedure. The Sb-SP11/SCR and Sb-SRK genes were derived from the Sb-haplotype of self-incompatible A. lyrata19. Sb-SP11/SCR was fused to the promoter region of B. rapa S9-SP11/SCR36. Construct for expressing Sb-SP11/SCR (SCRb/pBI121) was generated in the previous study19. For Sb-SRK expression in papilla, the ΨSRKA upstream region was cloned from Col-0 and used as the promoter sequence to enable papilla–cell specific expression. ΨSRKA upstream region was amplified using the primers listed in Supplementary Table 1. The PCR fragment was cloned into pGEM T-Easy vector (Promega) using the manufacturer’s protocol (ΨSRKA/pGEM). The NOS terminator was amplified from pBI121 using the primers listed Supplementary Table 1, and then was cloned into pGEM T-Easy vector to obtain NOS/pGEM. Both ΨSRKA/pGEM and NOS/pGEM plasmids were digested with KpnI (Takara) and SacII (Takara), and the NOS terminator fragment was inserted into ΨSRKA/pGEM to obtain the ΨSRKA-NOS/pGEM plasmid. The SRKb coding fragment was amplified with primers listed in Supplementary Table 1, and then cloned into the KpnI site of the ΨSRKA-NOS/pGEM plasmid to obtain ΨSRKA-SRKb-NOS/pGEM. The ΨSRKA-SRKb-NOS cassette was digested out with EcoRI (Takara), and then cloned into the EcoRI site of the SCRb/pBI121 plasmid to obtain SRKb-SCRb/pBI121.
The synSRKb sequence (Supplementary Fig. 9) was chemically synthesized by Medical & Biological Laboratories. The SRKb-SCRb/ pBI121construct was digested with KpnI to cut out the SRKb moiety and was treated with calf intestine alkaline phosphatase (Takara). The synSRKb fragment PCR amplified by primers listed in Supplementary Table 1 to add the KpnI restriction site. Amplified fragment was digested by KpnI and then introduced into the vector.
A 5,353-bp genomic fragment containing the full SRKIRCol-0 sequence or the fragment containing the partial SRKIRCol-0 sequence was amplified by PCR using Col-0 DNA as the template with the primers listed in Supplementary Table 1. Fragments were introduced into pCambia1300 digested with HindIII (Takara) and SacI (Takara) using the In-Fusion HD Cloning Kit (Takara-Bio, Kusatsu, Japan). These constructs were introduced into the SI-C24 line. Five independent T1 plants introduced with partial SRKIRCol-0 were obtained. Since they all expressed strong SI comparable to SI-C24, we analyzed one out of them for SRKb transcript accumulation.
All transgenic plants were generated using the Agrobacterium infiltration procedure, as previously reported37. Plants transfected with pBI121 binary vectors were selected for kanamycin resistance and pCambia1300 were selected for hygromycin resistance.
Genetic analysis of the T-DNA insertion line
The Col-0 + SRKb-SCRb line was crossed with the homozygous T-DNA insertion line SALK_137645C to obtain an F1 population. Individuals from the segregating F2 generation were genotyped for the presence of SRKb, and then genotyped for the presence of T-DNA insertions. Primers used for genotyping are listed in Supplementary Table 1.
Pollination experiments
Flowers were emasculated before anthesis. Pistils of flowers at stage 14 were harvested, transferred to 1% (w/v) agar plates, pollinated with pollen from stage 13 flowers, and then incubated for three hours under controlled conditions (22 ± 2 °C, humidity 50 ± 5%). For Supplementary Fig. 1, pistils from stage 13 flowers were also used in pollination tests. The pollinated pistils were fixed overnight at room temperature in a freshly prepared ethanol/acetate 3:1 (v/v) solution. After replacing the solution with 1 M sodium hydroxide, the pistils were incubated at 60 °C for 30 min (A. thaliana) or 60 min (A. lyrata). The solution was then replaced with the aniline blue staining solution (2% (w/v) tripotassium phosphate, 0.01% (w/v) aniline blue) and incubated at room temperature for 3 h. To facilitate phenotyping, we defined arbitrary compatibility scores based on the number of pollen tubes in the styles: 1, no tubes observed; 2, 1–10 tubes; 3, 11–30 tubes; 4, 31–60 tubes; 5, ≥60 tubes.
Gene expression studies
Total RNA was extracted from stigmas at stage 13 with the RNeasy Plant Mini Kit (Qiagen, Hilden, Germany). cDNA was synthesized from the RNA using the SuperScript IV Reverse Transcriptase Kit (Thermo Fischer Scientific, Massachusetts, USA). The real-time PCR reactions were performed with the QuantiTect SYBRGreen PCR Kit (Qiagen) using the LightCycler 96 system (Roche, Basel, Switzerland). Relative quantification was calculated relative to the endogenous ACTIN8 (ACT8: AT1G49240) gene, using the Relative quantification application mode of the LightCycler 96 system. The same set of primers was used to quantify the ACT8 ortholog from A. lyrata (genesh2_kg.1__4019__AT1G49240.1). Primers used for the quantitative reverse transcription-PCR analyses are listed in Supplementary Table 1.
Small RNA transcriptome analysis
Small RNAs were extracted from Col-0 stigmas at stage 13 using the mirVana™ miRNA Isolation Kit (Ambion). Library preparation and Illumina Hiseq single-end sequencing were conducted by Hokkaido System Science Co., Ltd. Small RNA transcriptome data for the C24 flower buds (under run IDs SRR4026067 and SRR4026068) were downloaded from the Sequence Read Archive at the National Center for Biotechnology Information [NCBI]. After adaptor trimming with the cutadapt version 1.1 software38 using parameters -O 5/-e 10/-g GTTCAGAGTTCTACAGTCCGACGATC/-a TGGAATTCTCGGGTGCCAAGG and rest set as defaults. The reads were mapped to the Arabidopsis genome (version TAIR10) or within the bacterial artificial chromosome sequence of the S locus in C24 (NCBI ID: EF182720) with Bowtie version 1.0.039 using parameters -v 2/-a/--best/--strata/-m 20 and rest set as defaults. Mapped reads were extracted using the functions implemented in the SAMtools suite40 with default parameters. The results were visualized with the Integrated Genome Viewer 2.3.9341.
Small RNA target prediction
Small RNA targetability predictions within the SRK sequences (SRKb; AB052756, SRKAWei-1; GU723787, ψSRKBCvi-0; AY772644) were evaluated using the psRNAtarget program21 with default parameters and an expectation score of 3.0 as a cut-off. An expectation score of 0 in this program indicates a perfect match; prediction scores ranging from 0.5 to 3.0 indicate potential targets with mismatches. From the small RNAs mapped to SRKIRCol-0 or SRKIRC24, those sized from 19 to 24 nucleotides (typical sizes for those functioning in gene silencing) were used for the analysis. Non-redundant reads were selected using the Fastx_Collapser program42.
Statistical analyses and exact p values
All of the statistical analyses and bar plot visualizations were carried out using R43. Dunnett’s test was performed using the glht function from the multcomp package44. For all bar plots, the values indicate means and the error bars indicate standard deviations. The number of replicates and the exact p-values are defined in the figure captions.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw sequence data files were deposited at the National Center for Biotechnology Information Sequence Read Archive (NCBI-SRA) under BioProject ID PRJNA561260. The data underlying Figs. 1a, 1b, 1c, 2b, 2c, 2d, 3a, 3b, 3c, 3d, 3e, 4b, 4c and 4d as well as Supplementary Figs. 1, 5b, 5c and 5d are provided as a Source Data file. All other data are available in the manuscript or in the supplementary materials.
References
Igic, B., Lande, R. & Kohn, J. R. Loss of self-incompatibility and its evolutionary consequences. Int. J. Plant Sci. 169, 93–104 (2008).
Fujii, S., Kubo, K. & Takayama, S. Non-self- and self-recognition models in plant self-incompatibility. Nat. Plants 2, 16130 (2016).
Takayama, S. & Isogai, A. Self-incompatibility in plants. Annu. Rev. Plant Biol. 56, 467–489 (2005).
Takayama, S. et al. The pollen determinant of self-incompatibility in Brassica campestris. Proc. Natl Acad. Sci. USA. 97, 1920–5 (2000).
Schopfer, C. R., Nasrallah, M. E. & Nasrallah, J. B. The male determinant of self-incompatibility in Brassica. Science 286, 1697–1700 (1999).
Takasaki, T. et al. The S receptor kinase determines self-incompatibility in Brassica stigma. Nature 403, 913–916 (2000).
Takayama, S. et al. Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature 413, 534–538 (2001).
Shimizu, K. K. & Tsuchimatsu, T. Evolution of selfing: recurrent patterns in molecular adaptation. Annu. Rev. Ecol. Evol. Syst. 46, 593–622 (2015).
Shimizu, K. K., Shimizu-Inatsugi, R., Tsuchimatsu, T. & Purugganan, M. D. Independent origins of self-compatibility in Arabidopsis thaliana. Mol. Ecol. 17, 704–14 (2008).
Tsuchimatsu, T. et al. Patterns of polymorphism at the self-incompatibility Locus in 1,083 Arabidopsis thaliana genomes. Mol. Biol. Evol. 34, 1878–1889 (2017).
Tsuchimatsu, T. et al. Evolution of self-compatibility in Arabidopsis by a mutation in the male specificity gene. Nature 464, 1342–6 (2010).
Nasrallah, M. E., Liu, P., Sherman-Broyles, S., Boggs, Na & Nasrallah, J. B. Natural variation in expression of self-incompatibility in Arabidopsis thaliana: implications for the evolution of selfing. Proc. Natl Acad. Sci. U. S. A. 101, 16070–4 (2004).
Indriolo, E., Safavian, D. & Goring, D. R. The ARC1 E3 ligase promotes two different self-pollen avoidance traits in Arabidopsis. Plant Cell 26, 1525–1543 (2014).
Nasrallah, J. B. & Nasrallah, M. E. Robust self-Incompatibility in the absence of a functional ARC1 gene in Arabidopsis thaliana. Plant Cell 26, 3838–3841 (2014).
Schierup, M. H., Mable, B. K., Awadalla, P. & Charlesworth, D. Identification and characterization of a polymorphic receptor kinase gene linked to the self-incompatibility locus of Arabidopsis lyrata. Genetics 158, 387–99 (2001).
Liu, P., Sherman-Broyles, S., Nasrallah, M. E. & Nasrallah, J. B. A cryptic modifier causing transient self-incompatibility in Arabidopsis thaliana. Curr. Biol. 17, 734–740 (2007).
Stone, S. L., Arnoldo, M. & Goring, D. R. A breakdown of Brassica self-incompatibility in ARC1 antisense transgenic plants. Science 286, 1729–31 (1999).
Indriolo, E., Tharmapalan, P., Wright, S. I. & Goring, D. R. The ARC1 E3 ligase gene is frequently deleted in self-compatible Brassicaceae species and has a conserved role in Arabidopsis lyrata self-pollen rejection. Plant Cell 24, 4607–20 (2012).
Iwano, M. et al. Calcium signalling mediates self-incompatibility response in the Brassicaceae. Nat. Plants 1, 15128 (2015).
Okamura, K., Chung, W.-J. & Lai, E. C. The long and short of inverted repeat genes in animals: microRNAs, mirtrons and hairpin RNAs. Cell Cycle 7, 2840–5 (2008).
Dai, X., Zhuang, Z. & Zhao, P.X. psRNATarget: a plant small RNA target analysis server (2017 release). Nucleic Acids Res. 46, W49–W54 (2018).
Prigoda, N. L., Nassuth, A. & Mable, B. K. Phenotypic and genotypic expression of self-incompatibility haplotypes in Arabidopsis lyrata suggests unique origin of alleles in different dominance classes. Mol. Biol. Evol. 22, 1609–1620 (2005).
Kusaba, M., Tung, C.-W., Nasrallah, M. E. & Nasrallah, J. B. Monoallelic expression and dominance interactions in anthers of self-incompatible Arabidopsis lyrata. Plant Physiol. 128, 17–20 (2002).
Sherman-Broyles, S. et al. S locus genes and the evolution of self-fertility in Arabidopsis thaliana. Plant Cell 19, 94–106 (2007).
Greaves, I. K. et al. Twenty-four-nucleotide siRNAs produce heritable trans-chromosomal methylation in F1 Arabidopsis hybrids. Proc. Natl Acad. Sci. USA 113, E6895–E6902 (2016).
Tarutani, Y. et al. Trans -acting small RNA determines dominance relationships in Brassica self-incompatibility. Nature 466, 983–986 (2010).
Yasuda, S. et al. A complex dominance hierarchy is controlled by polymorphism of small RNAs and their targets. Nat. Plants 3, 16206 (2016).
Mable, B. K., Schierup, M. H. & Charlesworth, D. Estimating the number, frequency, and dominance of S-alleles in a natural population of Arabidopsis lyrata (Brassicaceae) with sporophytic control of self-incompatibility. Heredity 90, 422–431 (2003).
Llaurens, V. et al. Does frequency-dependent selection with complex dominance interactions accurately predict allelic frequencies at the self-incompatibility locus in Arabidopsis halleri? Evolution 62, 2545–57 (2008).
Durand, E. et al. Dominance hierarchy arising from the evolution of a complex small RNA regulatory network. Science 346, 1200–5 (2014).
Shimizu, K. K., Kudoh, H. & Kobayashi, M. J. Plant sexual reproduction during climate change: Gene function in natura studied by ecological and evolutionary systems biology. Ann. Bot. 108, 777–787 (2011).
Tsuchimatsu, T. & Shimizu, K. K. Effects of pollen availability and the mutation bias on the fixation of mutations disabling the male specificity of self-incompatibility. J. Evol. Biol. 26, 2221–2232 (2013).
Billiard, S., Castric, V. & Vekemans, X. A general model to explore complex dominance patterns in plant sporophytic self-incompatibility systems. Genetics 175, 1351–1369 (2007).
Cook, L. M. & Saccheri, I. J. The peppered moth and industrial melanism: evolution of a natural selection case study. Heredity 110, 207–12 (2013).
Grant, B. S. Allelic melanism in American and British peppered moths. J. Hered. 95, 97–102 (2004).
Shiba, H. et al. A pollen coat protein, SP11/SCR, determines the pollen S-specificity in the self-incompatibility of Brassica species. Plant Physiol. 125, 2095–103 (2001).
Iwano, M. et al. Fine-tuning of the cytoplasmic Ca2+ concentration is essential for pollen tube growth. Plant Physiol. 150, 1322–34 (2009).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–9 (2009).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24 (2011).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2016. (2017).
Hothorn, T., Bretz, F. & Westfall, P. Simultaneous inference in general parametric models. Biom. J. 50, 346–63 (2008).
Acknowledgements
We thank M. Okamura, M. Nara, T. Manabe, and Y. Yamamoto for their technical assistance. This work was supported in part by Grants-in-Aid for Scientific Research on Innovative Areas (23113002, 16H06467, 16H06464 to S.T.; 16H01467, 18H04776 to S.F.), Grants-in-Aid for Scientific Research (21248014, 25252021, 16H06380 to S.T.; 18H02456 to S.F.), a Grant-in-Aid for Challenging Exploratory Research (15K14626 to S.F.) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT) and the Japan Science and Technology Agency (JST) PRESTO program (JPMJPR16Q8) to S.F.
Author information
Authors and Affiliations
Contributions
S.T. conceived the study. S.F and S.T. wrote the paper. S.F. and H.S.-A. conducted the majority of the experiments and analyzed the data. M.K., T.K., M.I. contributed to transgenic experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Vincent Castric, Barbara Mable and Marcus Samuel for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fujii, S., Shimosato-Asano, H., Kakita, M. et al. Parallel evolution of dominant pistil-side self-incompatibility suppressors in Arabidopsis. Nat Commun 11, 1404 (2020). https://doi.org/10.1038/s41467-020-15212-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-15212-0
This article is cited by
-
MLPK function is not required for self-incompatibility in the S29 haplotype of Brassica rapa L.
Plant Reproduction (2023)
-
Genetics Behind Sexual Incompatibility in Plants: How Much We Know and What More to Uncover?
Journal of Plant Growth Regulation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.