Abstract
Motile cilia serve vital functions in development, homeostasis, and regeneration. We recently demonstrated that TAp73 is an essential transcriptional regulator of respiratory multiciliogenesis. Here, we show that TAp73 is expressed in multiciliated cells (MCCs) of diverse tissues. Analysis of TAp73 mutant animals revealed that TAp73 regulates Foxj1, Rfx2, Rfx3, axonemal dyneins Dnali1 and Dnai1, plays a pivotal role in the generation of MCCs in male and female reproductive ducts, and contributes to fertility. However, the function of MCCs in the brain appears to be preserved despite the loss of TAp73, and robust activity of cilia-related networks is maintained in the absence of TAp73. Notably, TAp73 loss leads to distinct changes in ciliogenic microRNAs: miR34bc expression is reduced, whereas the miR449 cluster is induced in diverse multiciliated epithelia. Among different MCCs, choroid plexus (CP) epithelial cells in the brain display prominent miR449 expression, whereas brain ventricles exhibit significant increase in miR449 levels along with an increase in the activity of ciliogenic E2F4/MCIDAS circuit in TAp73 mutant animals. Conversely, E2F4 induces robust transcriptional response from miR449 genomic regions. To address whether increased miR449 levels in the brain maintain the multiciliogenesis program in the absence of TAp73, we deleted both TAp73 and miR449 in mice. Although loss of miR449 alone led to a mild ciliary defect in the CP, more pronounced ciliary defects and hydrocephalus were observed in the brain lacking both TAp73 and miR449. In contrast, miR449 loss in other MCCs failed to enhance ciliary defects associated with TAp73 loss. Together, our study shows that, in addition to the airways, TAp73 is essential for generation of MCCs in male and female reproductive ducts, whereas miR449 and TAp73 complement each other to support multiciliogenesis and CP development in the brain.
Similar content being viewed by others
Introduction
Cilia are hair-like appendages protruding from the cell membrane into the surrounding environment. Solitary immotile primary cilia are a common organelle in most mammalian cells, whereas motile cilia are restricted to a subset of cell types. This subset includes multiciliated cells (MCCs) lining brain ventricles, tracheal, and bronchial epithelium as well as the epithelium of male efferent ducts (EDs) and fallopian tubes (FTs) in females [1].
Multiciliogenesis requires precise regulation of the production, transport and assembly of a large number of different structural components, a process critically dependent on a hierarchical network of transcriptional and post-transcriptional regulators [2]. Geminin coiled-coil domain containing 1 (GEMC1) [3,4,5] and multiciliate differentiation and DNA synthesis associated cell cycle protein (MCIDAS or Multicilin) [6,7,8], members of the Geminin family, are early regulators of the MCC fate, downstream of the Notch pathway. MCC differentiation is also regulated by post-transcriptional mechanisms including microRNAs (miRNAs). miR-34/449 constitutes a conserved family that encodes six homologous miRNAs (miR34a, 34b, 34c, 449a, 449b, and 449c) from three genomic loci in vertebrates. Inhibition of the Notch pathway e.g., by miR449 is required for multiciliogenesis through derepression of the transcriptional network of GEMC1, MCIDAS, E2F transcription factors (E2F4, E2F5), forkhead box J1 (FOXJ1), and v-myb avian myeloblastosis viral oncogene homolog (MYB) [9,10,11]. Disturbance of the molecular circuit leads to defective multiciliogenesis and ciliopathies in the airways, reproductive tracts, and the brain [1].
Transformation-related protein 73 (Trp73) is a member of the p53 family with distinct isoforms generated from two alternative promoters: isoforms containing the N-terminal transactivation domain (TAp73) and N-terminally truncated dominant-negative isoforms (ΔNp73). Recently, we and others showed that TAp73 is essential for airway multiciliogenesis [12, 13]. Gene expression analysis and chromatin immunoprecipitation (ChIP) identified TAp73 as a critical regulator of multiciliogenesis: TAp73 acts downstream of E2F4/MCIDAS and regulates the expression of FOXJ1, RFX2, and RFX3 in pulmonary tissues [12, 14,15,16,17].
The FT of female reproductive tract consists of MCCs that possess hundreds of motile cilia beating in a wave-like manner which, along with musculature contraction, moves the oocyte or zygote towards the uterus [18,19,20]. Defects in ciliary functions may lead to ectopic pregnancies or infertility [19, 21]. In the male reproductive tract, MCCs in the EDs are involved in the transport of spermatozoa from testis to epididymis (Epi), their maturation and concentration [22,23,24,25].
MCCs in the brain can be found in a single layer of ependymal cells facing the ventricles and choroid plexus (CP). The CP epithelium, a specialized secretory epithelium that secretes cerebrospinal fluid, arises from monociliated progenitors in the roof plate around embryonic day (E) 12 and undergoes multiciliate differentiation to form multiple primary cilia [26, 27]. Ependymal cells in mice are specified around E16 and form multiple motile cilia on the apical surface after birth to facilitate cerebrospinal fluid movement [28, 29]. Defects in the ependymal and CP lineages are implicated in aging, hydrocephalus, and brain tumors [30, 31].
In this study, we detected robust TAp73 expression in MCCs in diverse tissues. In reproductive ducts, TAp73 loss leads to a profound reduction of multiciliogenesis and suppression of TAp73-dependent transcriptional network activity. However, MCCs in the brain maintain robust multiciliogenesis activity despite TAp73 loss. Molecular studies revealed alterations in miR-34/449 family members in diverse MCCs from TAp73 mutant mice: decreased levels of TAp73 target miR34bc concurrent with increased expression of miR449. In the brain, miR449 is highly expressed in the CP and experiences significant upregulation following TAp73 deletion. In addition, brain ventricles but no other multiciliated tissues from TAp73 mutant animals exhibit increased expression of E2F4, which in turn is capable of eliciting robust transcriptional response from miR449 genomic loci, suggesting that miR449 plays a crucial role in brain multiciliogenesis in collaboration with TAp73. Indeed, miR449 loss alone results in ciliary reduction in the CP, whereas loss of both TAp73 and miR449 leads to a dramatic reduction of multiciliogenesis in the CP and severe hydrocephalus. Therefore, the molecular network governing MCC fate is subjected to tissue-specific feedback modulation by transcriptional and post-transcriptional mechanisms.
Materials and methods
Animals
TAp73 mutant mice with a targeted deletion of exons 2 and 3 of the Trp73 gene were a generous gift from Dr. Tak Mak (Princess Margaret Cancer Centre, Toronto, Canada) [32]. miR449 mutants were previously described [33]. Both strains were maintained in C57Bl/6 background (n8) at the animal facility of the European Neuroscience Institute Goettingen, Germany in full compliance with institutional guidelines. The study was approved by the Animal Care Committee of the University Medical Centre Goettingen and the authorities of Lower Saxony under the number 16/2069.
Human samples
Human Epi samples were procured with informed consent from two patients. All experimental procedures were approved and performed in accordance with the requirements set forth by the Ethics Committee of the University Medical Centre Goettingen (application number: 18/2/16).
Histology and immunostaining
Paraformaldehyde-fixed, paraffin-embedded tissues were treated with heat-induced epitope retrieval using Rodent Decloaker (RD913 L, Biocare Medical, Pacheco, CA, USA). For immunohistochemistry, endogenous peroxidase activity was quenched with 3% H2O2 for 10 min. Tissue sections were blocked with 10% fetal calf serum in phosphate-buffered saline with 0.1% Triton X-100, and subsequently incubated with primary antibodies (list of antibodies is provided in Supplementary Table 1). Biotinylated secondary antibodies were applied for 1 h at room temperature (list of antibodies is provided in Supplementary Table 2), after which avidin enzyme complex and substrate/chromogen were used for color development (Vector laboratories, Burlingame, CA, USA). Stained tissue sections were counterstained with hematoxylin. For immunofluorescence, sections were stained with fluorescently labeled secondary antibodies (list of antibodies is provided in Supplementary Table 2) for 1 h at room temperature. Nuclei were counterstained with 4′,6-diamidin-2-phenylindol (DAPI). Histology of tissue sections was assessed by using hematoxylin (Merck, Darmstadt, Germany) and eosin (Carl Roth, Karlsruhe, Germany) staining.
Electron microscopy
Transmission electron microscopy (TEM) was performed as previously described [12]. Briefly, murine tissue samples were fixed by immersion using 2% glutaraldehyde in 0.1 M cacodylate buffer (Science Services, München, Germany) at pH 7.4 overnight at 4 °C. Postfixation was performed using 1% osmium tetroxide diluted in 0.1 M cacodylate buffer. After pre-embedding staining with 1% uranyl acetate, tissue samples were dehydrated and embedded in Agar 100 (Plano, Wetzler, Germany). Thin tissue sections (100 nm) were examined using a Philips CM 120 BioTwin transmission electron microscope (Philips Inc., Eindhoven, The Netherlands) and images were taken with a TemCam F416 CMOS camera (TVIPS, Gauting, Germany).
Quantification of cilia markers
Cilia were quantified using the ImageJ software [34]. Briefly, the region of interest was selected and a threshold was set to exclude unspecific background signals. The Analyze Particles tool was used to measure the area of the ciliary staining. Values were normalized to the length of the epithelia measured.
Western blot
Samples were homogenized in RIPA buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 9.5 mM EDTA, 1% Triton X-100, 0.1% SDS, 1% sodium deoxycholate) supplemented with urea (2.7 M) and protease inhibitors (Complete Mini EDTA-free, Roche, Basel, Switzerland). Equal amounts of protein extracts were separated by SDS-polyacrylamide gels prior to transfer onto a nitrocellulose membrane and incubated with primary antibodies (list of antibodies is provided in Supplementary Table 1). The membrane was washed and incubated for 1 h with horse radish peroxidase-conjugated secondary antibodies (list of antibodies is provided in Supplementary Table 2) followed by chemiluminescence detection. β-Actin or heat shock cognate 71 kDa protein (HSC70) was used as protein loading controls.
RNA extraction, quantitative PCR, small RNA sequencing, and RNAscope
Tissue samples were snap-frozen in liquid nitrogen and total RNA was isolated by Extrazol (7BioScience, Hartheim, Germany)/Chloroform extraction followed by 80% ethanol precipitation at −20 °C. For cDNA synthesis, 1 μg of total RNA was incubated with the M-MuLV reverse transcriptase and a mix of random nonameric and polyA tail primers at 42 °C for 1 h in a total volume of 50 µl. All reactions were set up in triplicate with self-made SYBR Green quantitative PCR (qPCR) Mix (Tris-HCl [75 mM], (NH4)2SO4 [20 mM], Tween-20 [0.01% v/v], MgCl2 [3 mM], Triton X-100 [0.25% v/v], SYBR Green I (1:40 000), dNTPs [0.2 mM], and Taq polymerase [20 U/ml]) using 250 nM of each gene-specific primer (list of primers is provided in Supplementary Table 3). Standard curve method was used to assess relative transcript content. Transcript of interests was normalized to the reference transcript of ribosomal phosphoprotein P0 (Rplp0 or 36b4) and normalized to the mean value of control samples. The results for each sample were obtained by averaging transcript levels of technical triplicates. No RT controls and dilution curves as well as melting curves and gel electrophoresis assessment of amplicons were performed for all primer combinations. For miR449a, miR34b, and miR34c quantification, the TaqMan miRNA Assay (Applied Biosystems, Thermo Fisher Scientific, Waltham, MA, USA) was performed according to the manufacturer’s instructions with U6 snRNA as internal control.
Copy number in RNA samples was determined by qPCR using a murine TAp73 plasmid (MC219984, Origene, Rockville, USA) with a known copy number as standard curve. Copy number of the TAp73 plasmid was determined using the following formula: number of copies = (plasmid amount [ng] × 6.022 × 1023 [molecules/mole])/(plasmid length [bp] × 1 × 109 [ng/g] × 650 [g/mol])
The libraries for small RNA samples were prepared using the TruSeq Small RNA Library Prep Kit-Set A (24 rxns) (Set A: indexes 1–12; Cat N°: RS-200-001, Illumina, San Diego, CA, USA) using 1 µg of total RNA according to the manufacturer’s recommendations. Samples were sequenced on the Illumina HiSeq 4000 using a 50 bp single-end approach. Mapping, prediction of novel miRNAs, quality control, and differential expression (DE) analysis were carried out using Oasis2.0 (Oasis: online analysis of small RNA deep sequencing data) [35]. In brief, FASTQ files were trimmed with cutadapt 1.7.1 [36] removing TruSeq adapter sequences (TGGAATTCTCGGGTGCCAAGG) followed by removing sequences smaller than 15 or larger than 32 nucleotides. Trimmed FASTQ sequences were aligned to mouse small RNAs using STAR version 2.4.1d [37] with a mismatch of 5% of the sequence length and by utilizing the following databases: Mirbase version 21 for miRNAs; piRNAbank V.2 for piwiRNA; and Ensembl v84 for small nuclear RNA, small nucleolar RNA, and ribosomal RNA. Counts per small RNA were calculated using featureCounts v1.4.6 [38]. Novel miRNAs were searched using miRDeep2 version 2.0.0.5 [39]. DE of small RNA was determined by DESeq2 [40], where small RNAs were considered differentially expressed with an adjusted p-value < 0.05 and absolute log2 fold change > 1. The results of the DE analysis can be found in Supplementary Table S6, and the small RNA-seq data sets can be found in Gene Expression Omnibus with accession number GSE108385.
TAp73 (probe no. 475741), Mcidas (probe no. 510401-C2), Hes1 (probe no. 417701), and Hes5 (probe no. 400991-C2) were visualized using the RNAscope 2.5 HD Duplex Reagent Kit (#322430, Advanced Cell Diagnostics, Hayward, CA, USA) according to the manufacturer’s instructions.
Chromatin immunoprecipitation (ChIP)
Chromatin was harvested from Saos2 cells transiently overexpressing TAp73α, TAp73β, and the control vector pcDNA3.1. Saos2 cells were routinely tested negative for Mycoplasma. ChIP and qPCR were performed as previously described using gene-specific primers (sequence information is provided in Supplementary Table 4) [12]. Enrichment levels were determined as the number of PCR products for each gene relative to total input.
Luciferase assay
The luciferase assay was performed as previously described [12]. Briefly, Saos2 cells were transfected with pcDNA3.1 empty vector, or pcDNA3.1 vector carrying E2F4 or MCIDAS, or both E2F4 and MCIDAS vectors. Moreover, a firefly luciferase reporter construct containing the putative three wild type E2F-binding sequences of miR449 genomic region (wild type, or “WT”), or the same sequences lacking the strongest predicted E2F-binding motif (mutant, or “Mut”) was transfected (sequence information is provided in Supplementary Table 5). In addition, a Renilla TK luciferase vector was co-transfected. At 24 h after transfection, cells were harvested and the luciferase activities were measured using the dual luciferase assay. Firefly luciferase activities were determined relative to those of Renilla TK luciferase vector and normalized to the mean value of samples from the control vector. Luciferase assays were performed as technical triplicates on every biological replicate.
Video microscopy
Murine FT and testis connected to the Epi were dissected and transferred to Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, Thermo Fisher Scientific, Waltham, MA, USA). To image spermatozoa, the Epi was separated from testis and vas deferens and an incision was made at distal end to release the spermatozoa. Spermatozoa as well as the peristaltic contraction of the FT were imaged with an inverse microscope.
Imaging of cilia-generated bead flow and cilia beating in the brain ventricular system
Mice brains were dissected and transferred to DMEM 21063 (Gibco, Thermo Fisher Scientific). Coronal slices containing the lateral ventricle, ventral third ventricle, aqueduct, and fourth ventricle were prepared by using a coronal adult brain matrix (ASI Instruments, Warren, MI, USA). The ventral third ventricle was processed further as previously described [41]. Tissue explant was placed in DMEM containing fluorescent latex beads (Fluoresbrite Multifluorescent 1.0 micron Microspheres, Polysciences, Warrington, PA, USA). Movement of fluorescent beads along the ventricular wall and within ventricular lumen was observed by fluorescence microscopy using a DMR (Leica, Wetzlar, Germany) upright microscope with an epifluorescence lamp. Ciliary beating was observed by differential interference contrast microscopy using the same set-up. Bead movement was recorded using a high-speed camera (Cascade II-512, Photometrics, Tucson, AZ, USA) operated by MultiRecorder Software (developed by Johannes Schröder_Schetlig) and analyzed using ImageJ software [34].
Statistical analysis
One-tailed, unpaired Student’s t-test assuming normal distribution, and equal variances were used to calculate statistical significance for pairwise comparisons. Luciferase assay statistics were assessed using one-way ANOVA assuming normal distribution followed by Dunnett’s multiple comparison tests. The following indications of significance were used: *P < 0.05, **P < 0.01, ***P < 0.001. N values represent biological replicates. Error bars indicate standard error of the mean (SEM).
Results
TAp73 is expressed in diverse multiciliated epithelia
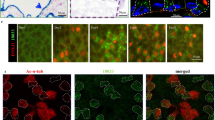
We and others previously showed that TAp73 expressed in respiratory epithelia controls multiciliogenesis [12, 13]. However, little is known about the expression and function of TAp73 in other MCCs. To address this, we performed immunostaining and in situ hybridization and demonstrated that in addition to the testis [42, 43], TAp73 is expressed in EDs, FTs, and ependymal and CP epithelial cells in the brain (Fig. 1a–f; Supplementary Fig. 1). qPCR and western blot analyses showed that, among different multiciliated epithelia, FTs and EDs exhibit higher levels of TAp73 expression than testis or brain (Fig. 1g–i). Taken together, these results demonstrate robust TAp73 expression in different MCC types.
TAp73 is expressed in diverse multiciliated epithelia. a Schematic illustration of efferent ducts (EDs, red arrows) that connect testis and epididymis (Epi). Blue dotted lines indicate regions used for histological, protein, and RNA analyses. b Representative images of the expression of P73 (green) and the axonemal marker acetylated alpha-tubulin (Ac-α-TUB, red) in the human ED. White bracket circles delineate P73 nuclear staining. DAPI staining (blue) marks nuclei. c Schematic illustration of the fallopian tube (FT) that connects ovary and uterus. Blue dotted line illustrates the region used for immunofluorescence analysis including fimbriae (red arrow). d Expression of P73 in human FT. Upper panel depicts a magnification of the boxed region in the lower panel. Arrowheads mark P73+ cells. Images were retrieved from Human Protein Atlas (https://www.proteinatlas.org/ENSG00000078900-TP73/tissue/fallopian+tube). e Schematic illustration of murine brain ventricles (red arrows). Blue dotted lines indicate the position of coronal brain slices used in protein and RNA analyses. f Expression of TAp73 in lateral and fourth ventricles of wild type (WT) adult mice. Red dotted lines demarcate ependymal cells lining brain ventricles. Notice that both ependymal and choroid plexus (CP) epithelial cells express TAp73. g Quantitative PCR analysis of TAp73 expression in the testis, EDs, FTs, and brain ventricles from WT adult mice. Expression levels are shown in copy number. Data from a single experiment are shown (testis, n = 3; FT and ventricle, n = 4; ED, n = 6). h Western blot analysis of the expression of TAp73 and ΔNp73 in testis, Epi, FT, and brain ventricle from WT adult mice. Heat shock cognate 71 kDa protein (HSC70) serves as a loading control. Data are representative of two independent experiments. i Quantitation of the signal intensity of TAp73 bands relative to that of HSC70 (h) is shown. All data are presented as mean ± SEM with *P < 0.05
TAp73 is crucial for the molecular circuit of multiciliogenesis in efferent ducts
Loss of TAp73 leads to male infertility that has been attributed to defective germ cell maintenance during spermatogenesis [42, 43]. Interestingly, we detected spermatozoa in testis from TAp73 knockout (KO) mice, although at a markedly reduced level (Supplementary Fig. 2a, b). Despite normal morphology and mobility of these cells, no mature spermatozoa were detected in the Epi of these mice (Fig. 2a; Supplementary Video 1a–d), suggesting that additional defects may contribute to infertility. The multiciliated epithelium of the EDs contributes to gamete transport by facilitating testicular fluid circulation, fluid reabsorption, and spermatozoa concentration [22, 24, 25], all essential aspects of male fertility [9, 44, 45]. Indeed, though no gross morphological difference was observed in EDs between control and TAp73 KO animals (Fig. 2a), immunofluorescent staining of the cilia components acetylated alpha-tubulin (Ac-α-TUB) and dynein axonemal intermediate chain 1 (DNAI1) showed a dramatic reduction in the number and length of cilia in the EDs from TAp73 KO mice (Fig. 2b, c). In contrast to the abundant long cilia of WT cells, mutant MCCs generated far fewer cilia as observed by TEM (Fig. 2d; Supplementary Fig. 2c), resembling the loss of airway cilia in these animals [12]. Consistent with its role as transcriptional regulator, ChIP followed by qPCR revealed significant enrichment of TAp73 in genomic loci of FOXJ1 [12] and dynein axonemal light intermediate chain 1 (DNALI1) and DNAI1, both encoding axonemal dyneins (Fig. 2e; Supplementary Fig. 3). Accordingly, expression of Dnali1, Foxj1, Rfx2, and Rfx3 was reduced or almost completely lost in male reproductive ducts from TAp73 KO animals (Fig. 2f, g; Supplementary Fig. 2d). Together, our data indicate that TAp73 directs Dnali1 and Dnai1 in addition to known critical nodes including Foxj1, Rfx2, and Rfx3 to mediate multiciliogenesis in EDs (Fig. 7a, b). Thus, these additional defects in the multiciliated epithelium of the EDs may contribute to male infertility in TAp73 KO mice.
TAp73 controls multiciliogenesis in the male reproductive tract. a Representative images of hematoxylin and eosin (H&E) staining of Epi sections from WT and TAp73 knockout (KO) animals. Bracket lines demarcate the border of EDs and Epi. Notice the lack of mature spermatozoa in cauda Epi from TAp73 KO mice (arrowheads). b Representative images of the expression of Ac-α-TUB (green) and axonemal dynein DNAI1 (red) in EDs from WT and TAp73 KO mice. DAPI staining (blue) labels nuclei. Boxed regions are magnified in the bottom panel. Note that TAp73 KO mice have less cilia that also exhibit reduced length (white bars). c Quantitation of Ac-α-TUB and DNAI1 signals normalized to epithelial length. Data from a single experiment are shown (WT, n = 6 images from 3 animals; TAp73 KO, n = 11 images from 4 animals). d Representative photomicrographs of transmission electron microscopy (TEM) in EDs from WT and TAp73 KO mice. Dotted lines mark apical region of the cells. Notice the abundant cilia (white arrows) and clustered basal bodies (white arrowhead) docked to the apical surface of WT cells, whereas mutant cells exhibit fewer cilia (red arrow). Interspersed microvilli are marked with asterisks. e Chromatin immunoprecipitation was performed for Saos2 cells transfected with TAp73α, TAp73β, and empty vector. Binding of TAp73α and TAp73β to genomic regions of FOXJ1, axonemal dyneins DNALI1 and DNAI1 was evaluated by quantitative PCR and compared to vector control (n = 3 for each antibody/gene pair, except for DNALI1 [n = 4], genomic regions examined are illustrated in Supplementary Fig. 3; [78]). f Semiquantitative PCR analysis of Dnali1, Foxj1, Rfx2, and Rfx3 expression in EDs from WT and TAp73 KO mice. Data from a single experiment are shown (WT, n = 4 for Dnali1, Foxj1, and Rfx3, n = 3 for Rfx2; TAp73 KO, n = 3). g Immunoblot analysis of the expression of TAp73, ΔNp73, DNAI1, and DNALI1 in Epi from WT and TAp73 KO animals. HSC70 serves as a loading control. Representative result of three independent experiments is displayed. All data are presented as mean ± SEM and relative to the WT group with *P < 0.05, **P < 0.01, ***P < 0.001
TAp73-driven transcriptional network regulates multiciliogenesis in fallopian tubes
Though infertility in TAp73 KO females is thought to arise from defects of oocyte development and release from the ovary [32, 46], it remains unclear whether TAp73 loss affects the multiciliated epithelium of the FT, thereby possibly influencing ova transport. Despite normal tubal morphology, analysis of Ac-α-TUB and DNAI1 expression showed reduced cilia coverage of the oviduct epithelium (Fig. 3a–c). Consistently, TEM demonstrated reduced cilia and mislocated basal bodies in FTs from TAp73 KO mice (Fig. 3d; Supplementary Fig. 4a). Transcript levels of Dnali1, Foxj1, and Rfx2, but not Rfx3, were reduced in TAp73 KO FTs (Fig. 3e), which were accompanied by declined protein expression of FOXJ1, DNAI1, DNALI1 (all expressed in the human FTs, Supplementary Fig. 4b), and gamma-tubulin (γ-TUB, basal body marker) (Fig. 3f), though to a lesser degree when compared to the decrease in multiciliogenesis activity observed in TAp73-deficient EDs. Further, smooth muscle contraction pattern in FTs is similar between control and TAp73 KO animals (Supplementary Video 2a, b). Taken together, our data indicate that TAp73 loss leads to reduced multiciliogenesis in the oviducts (Fig. 7a, c).
TAp73 controls multiciliogenesis in the oviducts. a Representative H&E staining of FTs from WT and TAp73 KO animals. b The expression of Ac-α-TUB (green) and DNAI1 (red) in FTs from WT and TAp73 KO mice. DAPI staining (blue) labels nuclei. Boxed regions are magnified in the bottom panel. In contrast to multiciliated epithelia in WT mice, TAp73 KO mice exhibit FT segments devoid of cilia (arrowheads). c Quantitation of Ac-α-TUB and DNAI1 signals normalized to epithelial length. Data from a single experiment are shown (WT, n = 6 images from 4 mice; TAp73 KO, n = 6 images from 3 mice). d Representative TEM photomicrographs of FTs from WT and TAp73 KO animals. Dotted lines mark apical region of the cells. Notice the presence of abundant cilia (black arrows) and basal bodies (white arrowhead) docked at the apical surface of WT cells, whereas mutant cells display fewer cilia (red arrows). Interspersed microvilli are marked with asterisks. e Semiquantitative PCR analysis of Dnali1, Foxj1, Rfx2, and Rfx3 expression in oviducts from WT and TAp73 KO mice. Data from a single experiment are shown (n = 3). f Immunoblot analysis of the expression of TAp73, ΔNp73, FOXJ1, DNAI1, DNALI1, and gamma-tubulin (γ-TUB) in oviducts from WT and TAp73 KO animals. HSC70 serves as a loading control. Data are representative of three independent experiments. All data are presented as mean ± SEM and relative to the WT group with *P < 0.05, **P < 0.01
Ciliary function in the brain is intact in the absence of TAp73
Given TAp73 expression in ependymal and CP epithelial cells, we further evaluated TAp73 expression during embryonic brain development. Immunofluorescent studies showed that proliferative progenitors (KI-67+) are present in hindbrain roof plate at E14.5, whereas post-mitotic cells expressing aquaporin 1 (AQP1) [31, 47] are detected in CP epithelium (KI-67−/AQP1+) (Fig. 4a). Notably, a portion of the roof plate exists between the progenitors and CP epithelium that remains undifferentiated after cell cycle exit (KI-67−/AQP1−) (Fig. 4a). In contrast to progenitors with a solitary primary cilium, the “transition” zone is comprised of MCCs that exhibit TAp73 expression (Fig. 4b).
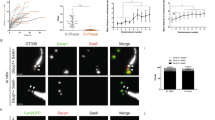
TAp73 is dispensable for brain multiciliogenesis. a The expression of KI-67, Aquaporin 1 (AQP1, green), and ADP-ribosylation factor-like 13b (ARL13B, red) in WT hindbrain roof plate/CP at E14.5. Notice that KI-67+ roof plate progenitors, and AQP1+ CP epithelial cells are spatially separated. ARL13B labels monociliated roof pate progenitors and multiciliated CP epithelial cells. White lines demarcate roof pate epithelium (KI-67+/AQP1−, arrows), CP epithelium (KI-67−/AQP1+, asterisks), and “transition zone” (KI-67−/AQP1−, arrowheads) in which MCCs appear. Dotted lines mark apical cell surface with cilia. DAPI staining (blue) labels nuclei. b Expression of TAp73 (green, red), AQP1 (green), and ARL13B (red) in WT hindbrain roof plate/CP at E14.5. Dotted lines mark apical cell surface of roof plate (TAp73−, arrow) and transition zone (TAp73+, arrowhead). White lines mark transition zone (TAp73+/AQP1−, arrowhead) and CP epithelium (TAp73+/AQP1+, asterisk). DAPI staining (blue) labels nuclei. c Representative images of TAp73 expression of ependymal and CP epithelial cells in hindbrain and lateral ventricle from WT and TAp73 KO. Red dotted lines mark ventricles lined with ependymal cells. Note that p73 expression is lost in TAp73 KO mice. d Expression of the cilia marker ARL13B (red) in CP epithelial cells from WT and TAp73 KO. White arrowheads mark cilia on cell surface. DAPI staining (blue) labels nuclei. Quantitation of average cilia length is shown in the lower panel. Data from a single experiment are shown (WT, n = 12 cells [hindbrain] and 9 cells [lateral ventricle] from 2 mice; TAp73 KO, n = 17 cells [hindbrain] and 15 cells [lateral ventricle] from 3 mice). e Immunoblot analysis of Ac-α-TUB in brain ventricles from WT and TAp73 KO animals. β-Actin serves as a loading control. Data are representative of two independent experiments. f Semiquantitative PCR of Dnali1, Foxj1, Rfx2, and Rfx3 in brain ventricles from WT and TAp73 KO. Data from a single experiment are shown (WT, n = 3; TAp73 KO, n = 4). g Movement of fluorescent beads along the ventricular system in WT and TAp73 KO mice. Images of maximum intensity projections of representative movies of the lateral and the ventral third ventricles are shown (WT, n = 2; TAp73 KO, n = 3; TAp73 heterozygous, n = 1). Red arrows mark the direction of bead flow. Bracket lines depict ependymal layer lining the lateral ventricles. Refer to Supplementary Video S3a, b for examples of recording of ciliary beating. All data are presented as mean ± SEM and relative to the WT group with *P < 0.05, **P < 0.01
The expression of TAp73 in ependymal and CP epithelial cells, along with recent studies demonstrating the role of E2F4/MCIDAS in multiciliogenesis of ependymal cells [7, 48, 49], led us to examine the role of TAp73 in MCCs in the brain. Immunostainings confirmed the loss of TAp73 expression in ependymal cells and the CP from TAp73 KO mice (Fig. 4c), whereas morphological analysis revealed no apparent defect in these cells (Supplementary Fig. 5a). We performed immunostainings for the cilia markers ADP-ribosylation factor-like 13b (ARL13B) [50], Ac-α-TUB, and DNAI1 in the fourth and lateral ventricles. In contrast to FTs and EDs, MCCs in ependyma and CP from TAp73 KO animals are similar to those of WT mice (Fig. 4d, e; Supplementary Fig. 5b–d). No significant difference was observed in the expression of markers for epithelial differentiation of CP between control and TAp73 KO animals (Supplementary Fig. 6a–d). RT-qPCR analysis demonstrated similar expression levels of Dnali1 and Foxj1, whereas increased Rfx2 and Rfx3 mRNA levels were observed in brain ventricles from TAp73 KO mice (Fig. 4f). Consistently, ciliary beating and bead flow in the cerebrospinal fluid appeared unaffected by TAp73 loss (Fig. 4g; Supplementary Video 3a, b). Taken together, these results indicate that, unlike EDs, FTs, and the airways [12], the differentiation and function of MCCs in the brain remain intact despite TAp73 loss.
TAp73 regulates miR-34/449 family members in diverse MCCs
Functional MCCs in the brain from TAp73 KO mice suggest that other ciliogenic factors may rescue brain multiciliogenesis in the absence of TAp73. TAp73 influences post-transcriptional mechanisms via regulation of miRNAs [12]. Analysis of small RNA species from brain ventricles in TAp73 KO mice revealed reduced miR34bc levels, along with a strong induction of the miR449a and miR449c that work together with the miR-34 cluster to regulate multiciliogenesis in different tissues across species (Fig. 5a, c; Supplementary Table 6) [11, 33, 51,52,53]. In the brain, miR449a is predominantly detected in the CP [54, 55], where its expression undergoes greater than tenfold increase upon TAp73 loss, whereas miR34bc levels strongly decline (Fig. 5b, c). Although miR34b levels were downregulated also in the trachea from TAp73 KO (Supplementary Fig. 7a), miR449a induction was less pronounced and more variable in FTs and EDs (Fig. 5d). Altogether, these results reveal a conserved reaction from the miR-34/449 family following TAp73 loss in diverse multiciliated epithelia.
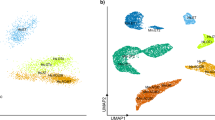
TAp73 loss leads to changes in miR-34/449 family and E2F4/MCIDAS circuit in the brain. a Hierarchical clustering of differentially expressed miRNAs in brain ventricles from WT and TAp73 KO mice (WT, n = 3; TAp73 KO, n = 4, one-way ANOVA, FDR < 0.05, fold change is shown). log2 values for miRNAs are plotted on the right. b In situ hybridization analysis of the expression of miR449 in WT roof plate/CP at E14.5 (http://www.eurexpress.org/ee/) [79]). Semiquantitative PCR analysis of miR449a, miR34b, and miR34c in brain ventricles (c), miR449a expression in EDs, FTs, and trachea (d), and E2f4 and Mcidas levels in brain ventricles (e) from WT and TAp73 KO mice. Data from a single experiment are shown (WT: ED, n = 3; FT, n = 7; trachea, n = 4; ventricle, n = 3. TAp73 KO: ED, n = 4; FT, n = 8; trachea, n = 4; ventricle, n = 4). f Immunoblot of E2F4 in WT and TAp73 KO ventricles. β-Actin serves as a loading control. Representative result of three independent experiments. g The luciferase assay of miR449 regulatory regions containing E2F-binding motifs. Three consensus E2F-binding sites in miR449 locus (http://jaspar.binf.ku.dk/) were placed in front of a luciferase cassette. A deletion mutant (Mut) that lacks the strongest consensus site was also created (Supplementary Table 5). WT or Mut luciferase vector was then co-transfected with empty vector (control), or vectors expressing E2F4, MCIDAS, or both. Fold changes in luciferase activities relative to those of control vector are shown. Data from five independent experiments are shown. All data are presented as mean ± SEM and relative to the WT group with *P < 0.05, **P < 0.01
In an effort to understand miR449 upregulation, we analyzed potential changes in the RB-E2F pathway known to regulate miR449 levels [56, 57]. However, expression of E2f1, E2f3, Cdkn1a, and Cdkn1b in brain ventricles were comparable between WT and mutant animals (Supplementary Fig. 8a, b), indicative of a RB-E2F pathway unaffected by TAp73 loss in brain MCCs. Interestingly, transcript and protein levels of the other E2F family member E2F4, which is a potent inducer of multiciliogenesis [6,7,8, 48, 58, 59], were markedly increased in TAp73 KO ventricles, despite only a mild increase of its cofactor Mcidas (Fig. 5e, f). In contrast, E2F4 levels in FTs and EDs were unaltered and even downregulated in tracheae (Supplementary Fig. 7b–e). Therefore, increased E2F4 levels concurrent with a miR449 increase are restricted to the brain in TAp73 KO mice.
To assess potential E2F4 contribution to miR449 elevation, we used the genomic region of miR449 containing three putative E2F-binding sites in a reporter-based assay. Indeed, E2F4 in combination with MCIDAS elicited a strong transcriptional response from the miR449 locus, a reaction almost abolished by mutating the strongest out of three E2F consensus motifs (Fig. 5g; Supplementary Table 5). Together, these results indicate that increased E2F4/MCIDAS activity may stimulate miR449 expression in TAp73 KO brains.
TAp73 collaborates with miR449 in brain multiciliogenesis
Our data suggest that miR449 upregulation may compensate at least partially for TAp73 loss to maintain brain multiciliogenesis. To address this, we generated mice with a deletion of the miR449 cluster in addition to TAp73. Strikingly, TAp73−/−;miR449−/− (TAp73xmiR449 KO) mice developed severe hydrocephalus, a defect not present in TAp73 KO or miR449 KO mice (Fig. 6a; Supplementary Fig. 9a). Since defective ependymal and CP cilia contribute to the development of hydrocephalus [60,61,62], we next assessed ciliation in the ventricles of TAp73xmiR449 KO mice. Analysis of the expression of ARL13B in CP epithelium revealed a decrease in cilia number and length in the absence of miR449, whereas a more pronounced reduction in cilia was observed in TAp73xmiR449 KO mice (Fig. 6b, c; Supplementary Fig. 9b). TEM studies also revealed mildly disorganized apical docking of basal bodies in ependymal cells in TAp73 KO and TAp73xmiR449 KO mice (Fig. 6d; Supplementary Fig. 9c); however, Ac-α-TUB content was similar in ependymal cells among WT and TAp73xmiR449 KO animals (Supplementary Fig. 9d). Consistently, ciliary beating and bead flow over ventricles appeared unaffected in TAp73xmiR449 KO animals (Supplementary Fig. 9e; Supplementary Video 3a, c). Furthermore, expression of cytokeratins, AQP1, and OTX2 in CP epithelial cells was similar among WT, miR449 KO, and TAp73xmiR449 KO animals (Supplementary Fig. 10a–c). Despite the role of Notch signaling in CP development and tumorigenesis [31, 63], RNAscope studies revealed similar expression of Notch targets Hes1 and Hes5, multiciliogenesis regulators Mcidas and Foxj1 in the roof plate of WT, miR449 KO, and TAp73xmiR449 KO embryos at E14.5 (Supplementary Fig. 11). In summary, additional loss of miR449 in TAp73 KO mice strongly impairs ciliogenesis in the CP, and slightly affects ependymal cilia, which is consistent with its prominent expression in the CP [54] (Fig. 5b). Thus, our data indicate that miR449 collaborates with TAp73 to drive multiciliogenesis in the brain.
TAp73 functions through miR449 in brain multiciliogenesis. a Coronal brain slices from WT, TAp73 KO, miR449 KO, and TAp73xmiR449 KO mice. Note that TAp73xmiR449 KO mice display enlarged lateral ventricles. b ARL13B (red) expression in CP epithelial cells of the fourth and lateral ventricles from WT, miR449 KO, and TAp73xmiR449 KO animals. White arrowheads mark cilia on cell surface. DAPI staining (blue) labels nuclei. c Quantitation of average cilia length of CP epithelial cells shown in (b). Data from a single experiment are shown (WT, n = 4 cells [hindbrain, lateral ventricle] from 2 mice; miR449 KO, n = 18 cells [hindbrain] and 14 cells [lateral ventricle] from 4 mice; TAp73xmiR449 KO, n = 8 cells [hindbrain, lateral ventricle] from 3 mice). d Representative TEM photomicrographs of ependymal cells in WT, TAp73 KO, and TAp73xmiR449 KO mice. Dotted lines mark apical region of the cells. Notice that WT cells possess cilia (white arrow) and basal bodies (white arrowhead) docked to the apical surface, whereas mutant cells have a similar number of cilia (red arrows) but disorganized basal bodies (red arrowheads) located further away from the apical surface. Interspersed microvilli are marked with asterisks. e Representative staining of Ac-α-TUB (green) in tracheae from WT and TAp73xmiR449 KO mice. DAPI staining (blue) labels nuclei. Note that mutants harbor less and shorter cilia (arrowhead) than WTs. f Quantitation of Ac-α-TUB signals normalized to epithelial length is shown. Data from a single experiment are shown (n = 4 samples/genotype). g Representative TEM photomicrographs of tracheae from WT and TAp73xmiR449 KO mice. Dotted lines mark apical region of the cells. Notice the abundant cilia (black arrows) and clustered basal bodies (white arrowhead) docked to apical surface in WT cells, whereas mutant cells exhibit fewer cilia (red arrow). Interspersed microvilli are marked with asterisks. All data are presented as mean ± SEM and relative to the WT group with **P < 0.01, ***P < 0.001
As miR449 was induced upon TAp73 deletion in further multiciliated tissues, we analyzed tracheae and EDs in TAp73xmiR449 KO mice. Immunostainings and TEM consistently revealed a dramatic decrease in cilia coverage and an increase in defective basal body docking in trachea from TAp73xmiR449 KO animals compared to WT animals (Fig. 6e–g; Supplementary Fig. 12a), a phenotype bearing resemblance to our previous findings in the airways of TAp73 KO animals [12]. Likewise, loss of miR449 did not further enhance MCC reduction in TAp73-deficient EDs (Supplementary Fig. 12b, c). Thus, additional deletion of the miR449 cluster fails to exacerbate ciliary defects in trachea and EDs in the absence of TAp73.
Overall, our data indicate that TAp73 utilizes the unique topology of its transcriptional circuit to communicate with the miR-34/449 family and other crucial regulators of multiciliogenesis e.g., E2F4/MCIDAS to regulate brain multiciliogenesis (Fig. 7a, d, e).
Schematic diagram of the molecular circuits of TAp73-driven multiciliogenesis in diverse tissues. a TAp73-dependent transcriptional network, including dyneins, miR34bc, Foxj1, Rfx2, and Rfx3 factors, critically regulates multiciliogenesis in various ciliated epithelia downstream of E2f4/Mcidas. In the EDs (b) and FTs (c) TAp73 KO impairs multiciliogenesis concurrent with male and female infertility. d TAp73 is not essential for multiciliogenesis in the brain; however, TAp73 loss leads to upregulation of pro-ciliogenic E2f4 and its target miR449. e Further removal of miR449 in TAp73 KO animals leads to reduced number and length of CP cilia and severe hydrocephalus, indicating that miR449 and TAp73 complement each other to support brain ciliogenesis
Discussion
TAp73 activates a plethora of ciliogenic effectors to drive multiciliogenesis in the airways [12, 13]. The current study examines the role of TAp73-driven molecular circuit in MCCs of reproductive tracts and the brain. Our results revealed a profound reduction of cilia in EDs and FTs from TAp73-deficient mice, as well as diminished Foxj1, Rfx2, and Rfx3 expression. These molecular and cellular changes in MCCs are reminiscent of our previous findings in respiratory epithelia of these mice, suggesting that male and female infertility associated with TAp73 loss could be in part related to the observed cilia loss. The expression of the axonemal dyneins Dnai1 and Dnali1, both of which exhibit TAp73 binding in their genomic loci, was also significantly reduced in EDs and FTs from mutant animals, indicating that they are part of the TAp73-directed multiciliogenesis program in reproductive tracts.
Consistent with previous reports, we found partial degradation of the germinal epithelium and reduced sperm cell production in TAp73 KO mice [42, 43]. The EDs are comprised of MCCs, which are required for fluid circulation and reabsorption, thereby, facilitating the transport of spermatozoa to their storage and maturation in the Epi [22, 24, 25]. Despite the presence of flagellated spermatozoa in testis, lack of spermatozoa in Epi of TAp73 KO mice indicates that defective multiciliogenesis may contribute to male sterility. Indeed, disruption of transcriptional and post-transcriptional regulators of multiciliogenesis has been shown to cause infertility in mice and humans [3, 64, 65], whereas fertility issues have been reported in female primary ciliary dyskinesia patients [20, 21]. Importantly, TAp73 is downregulated as women age [66], and certain single nucleotide polymorphisms in TP73 are associated with female patients over 35 years of age seeking in vitro fertilization [67, 68]. Hence, the integrity of MCCs is critical for reproductive health. Further studies using tissue-specific deletion of TAp73 in MCCs of EDs and oviducts are necessary to delineate its role in reproductive motile cilia maintenance and fertility.
In the brain, TAp73 expression is initiated at the onset of multiciliated differentiation of ependymal and CP epithelial cells. However, our data indicate that TAp73 is dispensable for the generation of cilia in the brain, although it is plausible that TAp73 loss results in more subtle defects such as polarity and cilia orientation [69, 70]. In contrast to the dynamic TAp73-dependent program in the airways and reproductive tracts, expression of Foxj1, Rfx2, and Rfx3 in the brain remains mostly unaltered in the absence of TAp73, suggesting that other effectors maintain the activity of the molecular circuit to support MCC differentiation.
Previous studies revealed robust expression of GemC1 and E2f/Mcidas, all of which are capable of transcriptional activation of Foxj1, TAp73 itself, and many other ciliogenic effectors e.g., Rfx2 and Rfx3, in MCCs of the brain [4, 6, 8, 48, 71]. Indeed, E2F4/MCIDAS expression is upregulated in the brain but not in other multiciliated tissues upon TAp73 loss, and therefore may facilitate brain multiciliogenesis. In agreement, loss of either Mcidas or GemC1, both transcriptional activators of TAp73, leads to defect in MCC differentiation and hydrocephalus [3, 6].
Although it is less clear how TAp73 loss results in enhanced E2F/MCIDAS activity in the brain, a quick look downstream of TAp73 provides some clues: reduced expression of the TAp73 target miR34bc is concurrent with an induction of miR449 in the absence of TAp73. Interestingly, expression of Cdkn1a/p21, Cdkn1b/p27, E2f1, and E2f3 in brain ventricles remain unchanged following TAp73 loss, suggesting that TAp73 loss regulates E2F and miR449 activity independently of the conserved RB-E2F1 axis. miR449 induction is commonly observed in miR-34-deficient MCCs, whereas ablation of the entire miR-34/449 family severely impairs multiciliogenesis in diverse tissues [33, 65, 72]. The redundancy might explain why a loss of the miR449 cluster failed to worsen ciliary defects outside the brain in TAp73 KO animals. In contrast, miR449 cluster is prominently expressed in MCCs in the brain, where its loss alone is sufficient to impact multiciliogenesis, indicating a unique role for miR449 in brain MCCs. miR449 is known to inhibit the Notch pathway to relieve the suppression of MCC fate determination; however, Notch pathway activity as well as the expression of Mcidas and Foxj1 in the roof plate and the CP remains unchanged after miR449 loss. Thus, it is plausible that miR449 may indirectly increase E2F/MCIDAS activity in MCCs of the brain independent of Notch inhibition and Mcidas derepression. Conversely, transcriptional activation of miR449 by E2F/MCIDAS complexes may complete the feedback loop to keep the molecular circuit fully engaged in the absence of TAp73.
This interpretation posits that the crosstalk between miR449 and E2F/MCIDAS serves as a crucial backup circuit for TAp73-driven multiciliogenesis network in the brain. Indeed, combined deletion of TAp73 and miR449 results in disruption of multiciliogenesis in the brain and hydrocephalus, defects distinct from those associated with loss of either TAp73 or miR449 alone, or complete loss of the miR-34/449 family [33, 73]. Strikingly, in TAp73-deficient MCCs outside the brain that exhibit less prominent increase in miR449 and no increase in E2F4 levels, further deletion of miR449 cluster fails to exacerbate multiciliogenesis defects caused by TAp73 loss, indicating that TAp73 functions at least partially through miR449 to support MCCs in the brain. Recent studies also demonstrated the role of TAp73-driven miR34a expression in neuronal development [74]. Therefore, interaction of TAp73 with miR-34/449 family members is crucial for normal brain functions.
Nonetheless, detailed studies are necessary to clarify the interaction between miR449 and E2F4/MCIDAS pathway in MCCs in the brain, but also to address miR449 regulation in TAp73-deficient MCCs outside the brain.
Unlike TAp73 mutant animals, p73 KO mice lacking both TAp73 and ΔNp73 exhibit hydrocephalus, defective ependymal cell maturation and aqueduct stenosis, suggesting a potential role for ΔNp73 in ependymal cells [75, 76]. Given the abnormal apical localization of basal bodies in ependymal cells along with ciliary defects in the CP from TAp73xmiR449 KO mice, it is conceivable that ΔNp73 may regulate miR449 expression indirectly in these cells. In support of this notion, miR449 is highly expressed in the CP whereby its loss alone leads to ciliary defects, whereas ΔNp73 deletion also results in defects in the CP [77]. Further analysis of the multiciliogenesis network and miR449 expression in MCCs in the brains of p73 KO and ΔNp73 KO animals are necessary to resolve these questions.
References
Spassky N, Meunier A. The development and functions of multiciliated epithelia. Nat Rev Mol Cell Biol. 2017;18:423–36.
Choksi SP, Lauter G, Swoboda P, Roy S. Switching on cilia: transcriptional networks regulating ciliogenesis. Development. 2014;141:1427–41.
Terré B, Piergiovanni G, Segura-Bayona S, Gil-Gómez G, Youssef SA, Attolini CS-O, et al. GEMC1 is a critical regulator of multiciliated cell differentiation. EMBO J. 2016;35:942–60.
Arbi M, Pefani D-E, Kyrousi C, Lalioti M-E, Kalogeropoulou A, Papanastasiou AD, et al. GemC1 controls multiciliogenesis in the airway epithelium. EMBO Rep. 2016;17:400–13.
Zhou F, Narasimhan V, Shboul M, Chong YL, Reversade B, Roy S. Gmnc is a master regulator of the multiciliated cell differentiation program. Curr Biol. 2015;25:3267–73.
Boon M, Wallmeier J, Ma L, Loges NT, Jaspers M, Olbrich H, et al. MCIDAS mutations result in a mucociliary clearance disorder with reduced generation of multiple motile cilia. Nat Commun. 2014;5:4418.
Ma L, Quigley I, Omran H, Kintner C. Multicilin drives centriole biogenesis via E2f proteins. Genes Dev. 2014;28:1461–71.
Stubbs JL, Vladar EK, Axelrod JD, Kintner C. Multicilin promotes centriole assembly and ciliogenesis during multiciliate cell differentiation. Nat Cell Biol. 2012;14:140–7.
Danielian PS, Hess RA, Lees JA. E2f4 and E2f5 are essential for the development of the male reproductive system. Cell Cycle. 2016;15:250–60.
Danielian PS, Bender Kim CF, Caron AM, Vasile E, Bronson RT, Lees JA. E2f4 is required for normal development of the airway epithelium. Dev Biol. 2007;305:564–76.
Marcet B, Chevalier B, Luxardi G, Coraux C, Zaragosi L-E, Cibois M, et al. Control of vertebrate multiciliogenesis by miR-449 through direct repression of the Delta/Notch pathway. Nat Cell Biol. 2011;13:693–9.
Nemajerova A, Kramer D, Siller SS, Herr C, Shomroni O, Pena T, et al. TAp73 is a central transcriptional regulator of airway multiciliogenesis. Genes Dev. 2016;30:1300–12.
Marshall CB, Mays DJ, Beeler JS, Rosenbluth JM, Boyd KL, Santos Guasch GL, et al. p73 is required for multiciliogenesis and regulates the Foxj1-associated gene network. Cell Rep. 2016;14:2289–300.
Blatt EN, Yan XH, Wuerffel MK, Hamilos DL, Brody SL. Forkhead transcription factor HFH-4 expression is temporally related to ciliogenesis. Am J Respir Cell Mol Biol. 1999;21:168–76.
Brody SL, Yan XH, Wuerffel MK, Song SK, Shapiro SD. Ciliogenesis and left-right axis defects in forkhead factor HFH-4-null mice. Am J Respir Cell Mol Biol. 2000;23:45–51.
Chen J, Knowles HJ, Hebert JL, Hackett BP. Mutation of the mouse hepatocyte nuclear factor/forkhead homologue 4 gene results in an absence of cilia and random left-right asymmetry. J Clin Investig. 1998;102:1077–82.
Yu X, Ng CP, Habacher H, Roy S. Foxj1 transcription factors are master regulators of the motile ciliogenic program. Nat Genet. 2008;40:1445–53.
Crow J, Amso NN, Lewin J, Shaw RW. Morphology and ultrastructure of fallopian tube epithelium at different stages of the menstrual cycle and menopause. Hum Reprod. 1994;9:2224–33.
Lyons RA, Saridogan E, Djahanbakhch O. The reproductive significance of human Fallopian tube cilia. Hum Reprod Update. 2006;12:363–72.
Raidt J, Werner C, Menchen T, Dougherty GW, Olbrich H, Loges NT, et al. Ciliary function and motor protein composition of human fallopian tubes. Hum Reprod. 2015;30:2871–80.
Vanaken GJ, Bassinet L, Boon M, Mani R, Honoré I, Papon J-F, et al. Infertility in an adult cohort with primary ciliary dyskinesia: phenotype–gene association. Eur Respir J. 2017;50:1700314.
Ilio KY, Hess RA. Structure and function of the ductuli efferentes: a review. Microsc Res Tech. 1994;29:432–67.
Lambot M-AH, Mendive F, Laurent P, Van Schoore G, Noël J-C, Vanderhaeghen P, et al. Three-dimensional reconstruction of efferent ducts in wild-type and Lgr4 knock-out mice. Anat Rec. 2009;292:595–603.
Hess RA. The efferent ductules: structure and functions [Internet]. In: Robaire B, Hinton BT, editors. The epididymis: from molecules to clinical practice. Boston, MA: Springer US; 2002. [cited 2017 Jan 19]. p. 49–80. Available from: http://link.springer.com/10.1007/978-1-4615-0679-9_4.
Hess RA. Small tubules, surprising discoveries: from efferent ductules in the turkey to the discovery that estrogen receptor alpha is essential for fertility in the male. Anim Reprod. 2015;12:7–23.
Lun MP, Johnson MB, Broadbelt KG, Watanabe M, Kang Y-J, Chau KF, et al. Spatially heterogeneous choroid plexus transcriptomes encode positional identity and contribute to regional CSF production. J Neurosci. 2015;35:4903–16.
Silva-Vargas V, Maldonado-Soto A, Mizrak D, Codega P, Doetsch F. Age-dependent niche signals from the choroid plexus regulate adult neural stem cells. Cell Stem Cell. 2016;19:643–52.
Spassky N, Merkle FT, Flames N, Tramontin AD, García-Verdugo JM, Alvarez-Buylla A. Adult ependymal cells are postmitotic and are derived from radial glial cells during embryogenesis. J Neurosci J Soc Neurosci. 2005;25:10–8.
Lun MP, Monuki ES, Lehtinen MK. Development and functions of the choroid plexus-cerebrospinal fluid system. Nat Rev Neurosci. 2015;16:445–57.
Del Bigio MR. Ependymal cells: biology and pathology. Acta Neuropathol. 2010;119:55–73.
Li L, Grausam KB, Wang J, Lun MP, Ohli J, Lidov HGW, et al. Sonic hedgehog promotes proliferation of Notch-dependent monociliated choroid plexus tumour cells. Nat Cell Biol. 2016;18:418–30.
Tomasini R, Tsuchihara K, Wilhelm M, Fujitani M, Rufini A, Cheung CC, et al. TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes Dev. 2008;22:2677–91.
Song R, Walentek P, Sponer N, Klimke A, Lee JS, Dixon G, et al. miR-34/449 miRNAs are required for motile ciliogenesis by repressing cp110. Nature. 2014;510:115–20.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Capece V, Garcia Vizcaino JC, Vidal R, Rahman R-U, Pena Centeno T, Shomroni O, et al. Oasis: online analysis of small RNA deep sequencing data. Bioinformatics. 2015;31:2205–7.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Friedländer MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012;40:37–52.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Faubel R, Westendorf C, Bodenschatz E, Eichele G. Cilia-based flow network in the brain ventricles. Science. 2016;353:176–8.
Holembowski L, Kramer D, Riedel D, Sordella R, Nemajerova A, Dobbelstein M, et al. TAp73 is essential for germ cell adhesion and maturation in testis. J Cell Biol. 2014;204:1173–90.
Inoue S, Tomasini R, Rufini A, Elia AJ, Agostini M, Amelio I, et al. TAp73 is required for spermatogenesis and the maintenance of male fertility. Proc Natl Acad Sci USA. 2014;111:1843–8.
Dacheux J-L, Dacheux F. New insights into epididymal function in relation to sperm maturation. Reproduction. 2013;147:R27–42.
Mendive F, Laurent P, Van Schoore G, Skarnes W, Pochet R, Vassart G. Defective postnatal development of the male reproductive tract in LGR4 knockout mice. Dev Biol. 2006;290:421–34.
Santos Guasch GL, Beeler JS, Marshall CB, Shaver TM, Sheng Q, Johnson KN, et al. p73 is required for ovarian follicle development and regulates a gene network involved in cell-to-cell adhesion. iScience. 2018;8:236–49.
Huang X, Ketova T, Fleming JT, Wang H, Dey SK, Litingtung Y, et al. Sonic hedgehog signaling regulates a novel epithelial progenitor domain of the hindbrain choroid plexus. Development. 2009;136:2535–43.
Kyrousi C, Lalioti M-E, Skavatsou E, Lygerou Z, Taraviras S. Mcidas and GemC1/Lynkeas specify embryonic radial glial cells. Neurogenesis. 2016;3:e1172747.
Mori M, Hazan R, Danielian PS, Mahoney JE, Li H, Lu J, et al. Cytoplasmic E2f4 forms organizing centres for initiation of centriole amplification during multiciliogenesis. Nat Commun. 2017;8:15857.
Caspary T, Larkins CE, Anderson KV. The graded response to sonic hedgehog depends on cilia architecture. Dev Cell. 2007;12:767–78.
Lizé M, Herr C, Klimke A, Bals R, Dobbelstein M. MicroRNA-449a levels increase by several orders of magnitude during mucociliary differentiation of airway epithelia. Cell Cycle. 2010;9:4579–83.
Marcet B, Chevalier B, Coraux C, Kodjabachian L, Barbry P. MicroRNA-based silencing of Delta/Notch signaling promotes multiple cilia formation. Cell Cycle. 2011;10:2858–64.
Otto T, Candido SV, Pilarz MS, Sicinska E, Bronson RT, Bowden M, et al. Cell cycle-targeting microRNAs promote differentiation by enforcing cell-cycle exit. Proc Natl Acad Sci USA. 2017;114:10660–5.
Redshaw N, Wheeler G, Hajihosseini MK, Dalmay T. microRNA-449 is a putative regulator of choroid plexus development and function. Brain Res. 2009;1250:20–6.
Shu P, Wu C, Liu W, Ruan X, Liu C, Hou L, et al. The spatiotemporal expression pattern of MicroRNA in the developing mouse nervous system. J Biol Chem. 2018;294:3444–53. jbc.RA118.004390
Yang X, Feng M, Jiang X, Wu Z, Li Z, Aau M, et al. miR-449a and miR-449b are direct transcriptional targets of E2F1 and negatively regulate pRb-E2F1 activity through a feedback loop by targeting CDK6 and CDC25A. Genes Dev. 2009;23:2388–93.
Lizé M, Pilarski S, Dobbelstein M. E2F1-inducible microRNA 449a/b suppresses cell proliferation and promotes apoptosis. Cell Death Differ. 2010;17:452–8.
Kyrousi C, Arbi M, Pilz G-A, Pefani D-E, Lalioti M-E, Ninkovic J, et al. Mcidas and GemC1 are key regulators for the generation of multiciliated ependymal cells in the adult neurogenic niche. Development. 2015;142:3661–74.
Kim S, Ma L, Shokhirev MN, Quigley I, Kintner C. Multicilin and activated E2f4 induce multiciliated cell differentiation in primary fibroblasts. Sci Rep. 2018;8:12369.
Ibanez-Tallon I. Dysfunction of axonemal dynein heavy chain Mdnah5 inhibits ependymal flow and reveals a novel mechanism for hydrocephalus formation. Hum Mol Genet. 2004;13:2133–41.
Banizs B, Pike MM, Millican CL, Ferguson WB, Komlosi P, Sheetz J, et al. Dysfunctional cilia lead to altered ependyma and choroid plexus function, and result in the formation of hydrocephalus. Development. 2005;132:5329–39.
Banizs B, Komlosi P, Bevensee MO, Schwiebert EM, Bell PD, Yoder BK. Altered pH(i) regulation and Na(+)/HCO3(−) transporter activity in choroid plexus of cilia-defective Tg737(orpk) mutant mouse. Am J Physiol Cell Physiol. 2007;292:C1409–1416.
Bill BR, Balciunas D, McCarra JA, Young ED, Xiong T, Spahn AM, et al. Development and Notch signaling requirements of the zebrafish choroid plexus. PLoS ONE 2008;3:e3114.
Amirav I, Wallmeier J, Loges NT, Menchen T, Pennekamp P, Mussaffi H, et al. Systematic analysis of CCNO variants in a defined population: implications for clinical phenotype and differential diagnosis. Hum Mutat. 2016;37:396–405.
Yuan S, Liu Y, Peng H, Tang C, Hennig GW, Wang Z, et al. Motile cilia of the male reproductive system require miR-34/miR-449 for development and function to generate luminal turbulence. Proc Natl Acad Sci USA. 2019;116:201817018.
Guglielmino MR, Santonocito M, Vento M, Ragusa M, Barbagallo D, Borzì P, et al. TAp73 is downregulated in oocytes from women of advanced reproductive age. Cell Cycle. 2011;10:3253–6.
Hu W, Zheng T, Wang J. Regulation of fertility by the p53 family members. Genes Cancer. 2011;2:420–30.
Feng Z, Zhang C, Kang H-J, Sun Y, Wang H, Naqvi A, et al. Regulation of female reproduction by p53 and its family members. FASEB J Publ Fed Am Soc Exp Biol. 2011;25:2245–55.
Fujitani M, Sato R, Yamashita T. Loss of p73 in ependymal cells during the perinatal period leads to aqueductal stenosis. Sci Rep. 2017;7:12007.
Gonzalez-Cano L, Fuertes-Alvarez S, Robledinos-Anton N, Bizy A, Villena-Cortes A, Fariñas I, et al. p73 is required for ependymal cell maturation and neurogenic SVZ cytoarchitecture. Dev Neurobiol. 2016;76:730–47.
Pefani D-E, Dimaki M, Spella M, Karantzelis N, Mitsiki E, Kyrousi C, et al. Idas, a novel phylogenetically conserved geminin-related protein, binds to geminin and is required for cell cycle progression. J Biol Chem. 2011;286:23234–46.
Bao J, Li D, Wang L, Wu J, Hu Y, Wang Z, et al. MicroRNA-449 and MicroRNA-34b/c function redundantly in murine testes by targeting E2F transcription factor-retinoblastoma protein (E2F-pRb) pathway. J Biol Chem. 2012;287:21686–98.
Fededa JP, Esk C, Mierzwa B, Stanyte R, Yuan S, Zheng H, et al. MicroRNA‐34/449 controls mitotic spindle orientation during mammalian cortex development. EMBO J. 2016;35:2386–98.
Agostini M, Tucci P, Killick R, Candi E, Sayan BS, Rivetti di Val Cervo P, et al. Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci USA. 2011;108:21093–8.
Medina-Bolívar C, González-Arnay E, Talos F, González-Gómez M, Moll UM, Meyer G. Cortical hypoplasia and ventriculomegaly of p73-deficient mice: developmental and adult analysis: p73 in developing and adult cortex. J Comp Neurol. 2014;522:2663–79.
Yang A, Walker N, Bronson R, Kaghad M, Oosterwegel M, Bonnin J, et al. p73-deficient mice have neurological, pheromonal and inflammatory defects but lack spontaneous tumours. Nature. 2000;404:99–103.
Tissir F, Ravni A, Achouri Y, Riethmacher D, Meyer G, Goffinet AM. DeltaNp73 regulates neuronal survival in vivo. Proc Natl Acad Sci USA. 2009;106:16871–6.
Koeppel M, van Heeringen SJ, Kramer D, Smeenk L, Janssen-Megens E, Hartmann M, et al. Crosstalk between c-Jun and TAp73alpha/beta contributes to the apoptosis-survival balance. Nucleic Acids Res. 2011;39:6069–85.
Diez-Roux G, Banfi S, Sultan M, Geffers L, Anand S, Rozado D, et al. A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol. 2011;9:e1000582.
Acknowledgements
We thank Tak Mak for providing TAp73 KO mice, Gerd Hasenfuß for support, Matthias Dobbelstein for hosting, and Karola Metze and Verena Siol for assistance. ML is supported by the Deutsche Forschungsgemeinschaft (DFG Li 2405); HZ by the New York Institute of Technology, Sanford Research, Matthew Larson Foundation, Institutional Development Award from the National Institute of General Medical Sciences under grant numbers 5P20GM103548, 1P20GM103620-01A1, and the National Cancer Institute (R01CA220551); FB by Wilhelm-Sander-Stiftung (2016.041.1); AKG by the Max Planck Society. We thank Heymut Omran’s group for introduction to cilia microscopy and Travis Stracker for disclosure of nonpublished data.
Author information
Authors and Affiliations
Contributions
MeW and TE characterized cilia defects and gene expression and generated figures. MeW and MaW validated TAp73 targets by WB and ChIP. EE and FB contributed IF analysis of human epididymis. EE performed cilia quantification on tracheae. DR performed electron microscopy analysis. CW and SB maintained mice. CW performed RNA isolations and qPCRs. LV-H and SB contributed to western blot analysis of different tissues. ZH performed RNAscope analysis for TAp73 on diverse tissues. KBG, JZ, LL, and HZ contributed brain analyses. A-KG analyzed ex vivo ciliary beating. OS analyzed small RNA sequencing data. SA contributed to interpretation and supported the group. ML developed the project, interpreted the data, designed and coordinated the experiments to complete this study. MeW, TE, HZ, and ML were major contributors to manuscript preparation.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Melino
Supplementary information
Rights and permissions
About this article
Cite this article
Wildung, M., Esser, T.U., Grausam, K.B. et al. Transcription factor TAp73 and microRNA-449 complement each other to support multiciliogenesis. Cell Death Differ 26, 2740–2757 (2019). https://doi.org/10.1038/s41418-019-0332-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0332-7
This article is cited by
-
CDC42 governs normal oviduct multiciliogenesis through activating AKT to ensure timely embryo transport
Cell Death & Disease (2022)
-
Disruption of GMNC-MCIDAS multiciliogenesis program is critical in choroid plexus carcinoma development
Cell Death & Differentiation (2022)
-
The p53 family member p73 in the regulation of cell stress response
Biology Direct (2021)