Abstract
Breast fibroepithelial lesions are biphasic tumors which comprise the common benign fibroadenomas (FAs) and the rarer phyllodes tumors (PTs). This study analyzed 262 (42%) conventional FAs, 45 (7%) cellular FAs, and 321 (51%) benign PTs contributed by the International Fibroepithelial Consortium, using a previously curated 16 gene panel. Benign PTs were found to possess a higher number of mutations, and higher rates of cancer driver gene alterations than both groups of FAs, in particular MED12, TERT promoter, RARA, FLNA, SETD2, RB1, and EGFR. Cases with MED12 mutations were also more likely to have TERT promoter, RARA, SETD2, and EGFR. There were no significant differences detected between conventional FAs and cellular FAs, except for PIK3CA and MAP3K1. TERT promoter alterations were most optimal in discriminating between FAs and benign PTs. Our study affirms the role of sequencing and key mutations that may assist in refining diagnoses of these lesions.
Similar content being viewed by others
Introduction
Breast fibroepithelial lesions comprise a family of biphasic tumors that display variations in their biology and clinical management. They include the common benign fibroadenomas, as well as the rarer phyllodes tumors, which can be benign, borderline, or malignant [1, 2]. The pathogenesis of breast fibroepithelial lesions is not completely understood, with a paucity of studies outlining their developmental mechanisms [3, 4]. In recent years, there have been new findings from genomic studies of these tumors [5,6,7,8,9,10], with MED12 being the most frequently aberrant gene, where the majority of mutations occur in codon 44 of exon 2 [5, 11, 12]. Genomic features have also suggested that fibroadenomas may be non-obligate precursors of phyllodes tumors [6, 13].
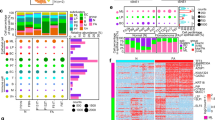
Histologically, fibroadenomas exhibit biphasic proliferation of stromal and epithelial compartments. The cellular variant of the fibroadenoma is characterized by mild to moderately increased stromal cellularity (Fig. 1). Phyllodes tumors also exhibit increased stromal cellularity but are accompanied by an exaggerated intracanalicular growth pattern with leaf-like stromal proliferations often referred to as fronds [14, 15]. Periductal stromal condensation may be observed. Between 60 and 75% of phyllodes tumors are diagnosed as benign, with borderline and malignant varieties at 15–20% and 10–20%, respectively [16, 17]. Phyllodes tumors may grow more rapidly than fibroadenomas on follow-up ultrasonography [18], but they cannot be reliably differentiated by imaging [19,20,21,22].
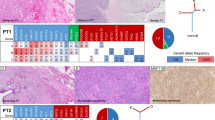
A Light microscopy image at low magnification displaying a conventional fibroadenoma with intracanalicular patterns. B Cellular fibroadenoma shows slightly increased cellularity. C Benign phyllodes tumor with prominent fronds at ×50 magnification. D Change in diagnoses after genomic analysis. FA: conventional fibroadenoma, CFA: cellular fibroadenoma, BPT: benign phyllodes tumor.
On histology, the differential diagnosis between cellular fibroadenomas and benign phyllodes tumors can prove challenging [3]. This was highlighted in an interobserver study involving ten breast pathologists evaluating 21 histologically challenging fibroepithelial lesions. Individual diagnoses varied from fibroadenoma to borderline phyllodes tumor in 9 (43%) of 21 cases. Of note, diagnoses were split equally (5/5) or nearly equally (6/4) between cellular fibroadenoma and benign phyllodes tumor in four cases [23].
While proliferative parameters such as mitotic activity and Ki67 labeling index have been applied in evaluating cellular fibroepithelial lesions, no consistent thresholds can be used for individual cases. This may restrict the value of such an approach to differentiating lesions within the fibroepithelial spectrum [24,25,26,27]. We have previously described a 5-gene RT-PCR signature that can distinguish between fibroadenoma and phyllodes tumor on core needle biopsies. However, the study evaluated mostly conventional fibroadenomas and further investigations into cellular fibroadenomas are needed [28]. We recently published the genomic characterization of an international series of breast fibroepithelial lesions [29]. Here, we delve deeper into specific comparisons among conventional fibroadenomas, cellular fibroadenomas, and benign phyllodes tumors derived from the cohort, to determine key mutations that may assist in refining their diagnoses.
Materials and methods
Formalin-fixed paraffin-embedded (FFPE) samples for a total of 628 adult fibroepithelial tumors [262 (42%) conventional fibroadenomas, 45 (7%) cellular fibroadenomas, and 321 (51%) benign phyllodes tumors] were obtained from institutions within the International Fibroepithelial Consortium in our initial study cohort.
DNA extraction and targeted next generation sequencing
One representative paraffin block of the tumor was selected from each case and eight sections of FFPE tissue, each measuring ten microns thick, were cut from the selected blocks. Genomic DNA was then extracted with the QIAamp DNA FFPE extraction kit (Qiagen, Valencia, CA, USA). The DNA yield and quality were determined using the PicoGreen® dsDNA quantitation assay (Thermofisher, Waltham, MA, USA).
The extracted DNA was then subjected to ultra-deep amplicon-based sequencing using the Illumina HiSeq 4000 platform. The assay encompasses a concise curated panel of 16 genes (Table 1) that was established based on the information derived from our group’s earlier studies [5, 6]. Sequencing was performed at a depth of at least 100× of the target regions for all samples.
Low-pass copy number variation (CNV) analysis
In total, 45 FFPE tissues comprising 15 conventional fibroadenomas, 15 cellular fibroadenomas, and 15 benign phyllodes tumors were selected for low-pass whole genome sequencing (WGS) to a depth of 5–10×. An additional ten blood samples were chosen as a panel of normal for this experiment. Sequencing was performed on the Illumina HiSeq platform with a paired end configuration of 150 base pairs.
Biocomputational analysis
Biocomputational analysis was done according to methods described in our previous study [29]. The FastQC software package (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) was utilized to perform quality checking of the FASTQ sequence files. After removing adapters, trimmed paired reads were mapped to hg19 (hs37d5) [30] using BWA-MEM (v0.7.15-r1140) [31] and sorted/indexed with samtools [32]. FreeBayes (v1.1.0-4-gb6041c6, settings: -m 30 -q 30 -F 0.01 -u) [33] and wANNOVA [34] were applied to call and annotate variants, respectively. Annotated variants were first filtered according to a few criteria: (1) only variants with more than 100X coverage and variant allele frequency of at least 5% were retained; (2) minor allele frequency of the variant in the normal population must be zero; (3) synonymous and non-exonic variants were also excluded, and (4) variants in dbSNP [35] were excluded, unless they are in COSMIC [36] or ClinVar [37, 38]. Manual curation of variants passing these filters was done on the Integrative Genomics Viewer [39] to remove any sequencing artefacts. R (R Foundation for Statistical Computing, Vienna, Austria) and the statistical software SPSS for Windows, version 25 (SPSS, Inc., Chicago, IL, USA) were used for statistical analysis. The relationship between the variant frequencies and the different FEL subgroups was analyzed using the χ2 and Fisher’s exact tests. A p value of < 0.05 was considered to be significant.
For the low-pass CNV, sequencing reads were aligned to human reference genome NCBI GRC Build 37 using BWA-MEM followed by duplicate marking and base-score recalibration using GATK version 4.14 for post alignment data processing.
ichorCNA was used as the method of estimating and accessing copy number alterations in this experiment. Firstly, a panel of normal was generated using the chosen germline samples which were identically processed and sequenced as the FFPE tissues to improve accuracy for estimating copy number alteration while correcting for systematic biases. This was performed with the help of an Rscript included with ichorCNA package. Once a panel of normal was generated copy number alterations were then identified using a bin size of 1 Mb. Downstream processes and illustrations were then analyzed using R.
Measuring sensitivity and specificity of the assay
Mutations of fibroadenomas and benign phyllodes tumors were compared to evaluate the assay’s performance. We classified the status of true- and false-positive and true- and false-negative categories based on the comparison groups. We calculated per cent sensitivity as 100 × [true positive/(true positive + false negative)], and specificity as 100 × [true negative/(true negative + false positive)]. Receiver-operating characteristic curves (ROC) and areas under the curve (AUC) were plotted to derive the mutation that could give optimal discrimination.
Histological review of cases
After genomic analysis, cases with available haematoxylin and eosin (H&E) slides and found to harbor unusual genotypes such as presence of cancer driver mutations (TP53, RB1, NF1, PTEN, PIK3CA, EGFR, BCOR, ERBB4, MAP3K1, IGF1R), or those with more than two mutations in non-cancer driver genes were histologically reviewed.
Results
Clinicopathological correlations
Our study cohort initially consisted of 628 cases of conventional fibroadenomas, cellular fibroadenomas, and benign phyllodes tumors. Ethnicity data were available for 605 cases. There were 517 (82%), 88 (14%), and 23 (4%) cases of Asian, Caucasian, or unknown descent, respectively. Three cases were re-diagnosed after histological review as borderline/malignant phyllodes tumors, thus excluding them from this study (Fig. 1). The following calculations for mutation rates are thereby based on the remaining 625 cases.
One hundred and fifty-eight (25%) cases showed no mutations, with 213 (34%), 150 (24%), 66 (11%), 24 (4%), 9 (1%), and 5 (1%) showing 1, 2, 3, 4, 5, or 6 mutations accordingly (Table 2). Benign phyllodes tumors harbored more mutations (mean 1.79), compared to cellular fibroadenomas (mean 1.18) and conventional fibroadenomas (mean 1.03) (p < 0.001). No significant differences were found between fibroadenomas and cellular fibroadenomas in terms of the number of mutations (p = 0.247). Benign phyllodes tumors were more likely to have a higher number of alterations in cancer driver genes than both groups of fibroadenomas (p = 0.008).
Fibroadenomas and benign phyllodes tumors possess distinct mutations
MED12 mutations were observed in 334 (53%) cases, followed by the TERT promoter, KMT2D, and RARA with aberrations in 122 (20%), 89 (14%), and 82 (13%) cases, respectively (Table 3 and Fig. 2). Benign phyllodes tumors showed the highest MED12 mutation rate with 199/322 (62%) cases harboring alterations in this gene, compared to 113/258 (44%) conventional fibroadenomas and 22/45 (49%) cellular fibroadenomas (p < 0.001). Further analysis revealed significant differences in mutation rates for six other genes in benign phyllodes tumors compared to conventional and cellular fibroadenomas (Table 3). These were TERT promoter (32 vs 6 vs 4%, p < 0.001), RARA (17 vs 8 vs 13%, p = 0.001), FLNA (13 vs 6 vs 4%, p = 0.002), SETD2 (12 vs 4 vs 7%, p < 0.001), RB1 (3 vs 1 vs 0%, p = 0.025), and EGFR (5 vs 2 vs 4%, p = 0.027). Conventional and cellular fibroadenomas did not differ significantly in their mutation spectrum except for PIK3CA (2 vs 9%, p = 0.011) and MAP3K1 (1 vs 4%, p = 0.047). Across our study cohort, we found MED12 mutations to be significantly associated with mutations in TERT (p < 0.001), RARA (p < 0.001), SETD2 (p = 0.012), and EGFR (p = 0.022) (Table 4).
Adjunctive value of genomic analysis in refining diagnosis
Histological review was performed for a subset of cases after genomic analysis of our initial study cohort of 628 fibroepithelial lesions, based on the criteria above. Numbers of cases here for each disease entity are reflective of what was known before the review. Out of 128 cases that underwent review, 6 (4.7%) had their diagnosis changed (Figs. 1 and 3).
A FEB941: this case was designated a fibroadenoma, but on histological review, the tumor comprised sheets of spindle cells areas of necrosis, consistent with a borderline/malignant phyllodes tumor. B FEB551: originally labeled as a fibroadenoma, histological review showed well-developed stromal fronds with mild stromal hypercellularity, consistent with a benign phyllodes tumor. C FEB589: case that was initially labeled as a benign phyllodes tumor, showed phyllodal architecture with leaf-like stromal fronds. D FEB589: higher magnification showed moderate to marked stromal atypia, readily found mitoses (arrows) and mild to moderate stromal hypercellularity, indicating borderline grade. E FEB1476: initially diagnosed as a benign phyllodes tumor, histological review favored borderline grade as the tumor borders were irregularly permeative along a relatively long front.
Of these six cases, three were revised from the original diagnosis of conventional fibroadenoma to benign phyllodes tumor. The first (FEB551) had missense mutations in MED12, FLNA, RARA, and ERBB4. The second (FEB851) had a frameshift mutation in KMT2D and a missense mutation in ERBB4, while the other (FEB858) had a missense mutation in MED12, a nonsense mutation in RARA, and a missense mutation in NF1.
Two cases of benign phyllodes tumor were re-diagnosed to borderline phyllodes tumor. The first (FEB589) harbored a promoter mutation in TERT and a missense mutation in TP53, while the other (FEB1476) displayed missense mutations in MED12, FLNA, and IGF1R. The last case (FEB941) was reclassified from conventional fibroadenoma to a spindle cell tumor with necrosis consistent with borderline or malignant phyllodes tumor. It disclosed mutations in MED12, TERT promoter, RARA, and EGFR.
Copy number variation analysis
CNVs were noted across all three subtypes in chromosomes 1–3, 7, 9–12, 15–16, 18–22, and X; and recurrent chromosomal alterations (occurring in at least two cases) were presented in this study (see Supplementary material, Table S1 and Fig. S1). A high proportion of fibroadenomas, cellular fibroadenomas, and benign phyllodes tumors had amplifications in 7q11.21-q21.11 (15/15 vs 15/15 vs 15/15), 19p13.3-p12 (15/15 vs 15/15 vs 14/15), and 19q13.11-q13.43 (15/15 vs 14/15 vs 14/15); gains in certain chromosomal segments such as 10q23.33-q25.1 (15/15 vs 15/15 vs 15/15), 20p13-p12.3 (15/15 vs 15/15 vs 13/15), and 21q21.1-q22.2 (12/15 vs 14/15 vs 11/15); and chromosomal deletions such as 7q36.1-q36.3 (14/15 vs 14/15 vs 15/15), 10p15.3-p14 (15/15 vs 15/15 vs 15/15), 18p11.32-q21.2 (15/15 vs 13/15 vs 14/15), and 19q11-q13.11 (14/15 vs 15/15 vs 14/15). We did not observe any statistically significant differences between conventional fibroadenoma and cellular fibroadenoma; and when comparing conventional fibroadenoma, cellular fibroadenoma, and benign phyllodes tumors.
Discussion
A diagnosis of fibroadenoma is usually straightforward. However, uncertainty arises in cellular fibroadenomas that possess overlapping features with phyllodes tumors [3]. The extent of leaf-like fronds and stromal cellularity that favor a benign phyllodes tumor over a cellular fibroadenoma remain subjective, and substantial interobserver differences exist. For instance, stromal atypia at the benign end of the fibroepithelial spectrum where benign phyllodes tumors and cellular fibroadenomas both lie is usually mild. Stromal cellularity and mitotic activity of benign phyllodes tumors also overlap with those of cellular fibroadenomas [40, 41]. As a result, distinguishing cellular fibroadenomas from benign phyllodes tumors can pose significant difficulties, even for experienced breast pathologists [23]. In this study, we determined if there were molecular differences between fibroadenomas (in particular cellular fibroadenomas) and benign phyllodes tumors that can be adjunctive aids in overcoming the limitations posed by histological examination.
We observed MED12 mutations across all three groups of lesions, with the majority (72%) occurring in codon 44 (see Supplementary material, Table S2). MED12 was noted to be involved in Wnt/β-catenin signaling, as β-catenin binds with the MED12 subunit in Mediator to activate transcription [42]. This pathway was found to be dysregulated in uterine leiomyomas which reported identical missense MED12 mutations as fibroadenomas [43]. Thus, the MED12 gene may be similarly affected in benign fibroepithelial lesions to promote cell proliferation and differentiation via Wnt signaling, which is known to be a fundamental growth control pathway [42].
According to our previous gene expression profiling study on ten fibroadenomas, genes upregulated in MED12-mutant fibroadenomas were associated with ER+ breast cancers and estrogen stimulus in ER+ breast cancer cells [5], and MED12 could enhance the function of estrogen receptor alpha (ERα) [44]. KMT2D which encodes the epigenetic regulator lysine (K)-specific methyltransferase 2D could directly interact with ERα, causing its recruitment and activation [45]. The retinoic acid receptor alpha which is encoded by RARA acts as a transcription factor and is capable of binding with ERα [46]. Activating mutations in PIK3CA have been reported to be frequent in ER-positive breast cancer [45]. Meanwhile, ER-EGFR signaling was discovered to be reciprocal: EGFR signaling promotes the activation of ER, and ER signaling promotes the activation of EGFR. Estrogen may induce activation of MAPK and PI3K which are associated with cell proliferation, angiogenesis, and tumor metastasis in non-small cell lung cancer [47]. In terms of immunohistochemical studies, epithelial expression of ER was higher in benign PTs than in borderline and malignant PTs [48], whereas Tan et al. observed significantly higher ER expression in fibroadenomas than in phyllodes tumors [49]. Thus, these results suggest potential involvement of estrogen signaling in the pathogenesis of benign tumors. Luo et al. observed that MED12 could bind to the EGFR promoter and result in EGFR transcription in ovarian cancers [50]. EGFR plays a role in elevating tumor cell survival, motility, and development through the Ras-Raf-Mek-Erk (Ras-MAPK) pathway [51], and its mutations have been described in breast, lung, and colorectal cancers [52,53,54]. Interestingly, activating mutations such as L858R confer sensitivity to tyrosine kinase inhibitors (TKIs) gefitinib and erlotinib in non-small cell lung cancer, as mutant EGFR binds the inhibitors more tightly than wild-type EGFR [55]. However, we did not detect this mutation in our current cohort of fibroadenomas and benign phyllodes tumors.
From our results, benign fibroepithelial tumors generally had a low incidence of cancer driver mutations, reaffirming their benign nature. The ten cancer driver genes (TP53, RB1, NF1, PTEN, PIK3CA, EGFR, BCOR, ERBB4, MAP3K1, and IGF1R) had the lowest incidence of mutations amongst the 16 genes (Table 4). Among the three subcategories of benign fibroepithelial tumors, benign phyllodes tumors also had a higher proportion of tumors having four or more mutations (33/322, 10%) compared to cellular fibroadenomas and conventional fibroadenomas (1/45, 2% and 4/258, 2%, respectively) (Table 2), indicating that they were more genetically complex. However, a statistical difference in mutation rate was found only in two out of the ten cancer driver genes (RB1 and EGFR) when comparing benign phyllodes tumors with fibroadenomas. Taken together, this shows that benign phyllodes tumors on the whole are similar to fibroadenomas (eight out of the ten cancer driver genes showed no significant differences in mutation frequencies), yet at the same time, the higher rate of mutations in RB1 and EGFR seen in benign phyllodes tumors suggests a potential for malignant progression compared to fibroadenomas as both alterations are detected in phyllodes tumors of higher grades [56]. Sixteen conventional fibroadenomas in our cohort harbored TERT promoter –124 C > T (chr5:1,295,228C > T) and –146 C > T (chr5:1,295,250C > T) mutations (15/16, 94% and 1/16, 6%, respectively), while two cellular fibroadenomas had alterations in the former (see Supplementary material, Table S3). Yoshida et al. similarly reported TERT promoter mutations in fibroadenomas although their proportion was less (4/58, 7%) [57], while four other studies did not detect such alterations among their fibroadenoma samples [8, 58,59,60]. When comparing benign phyllodes tumors and cellular fibroadenomas, mutations in TERT promoter gene showed the greatest difference in mutation frequency (32 vs 4%) (Table 3), while its sensitivity and specificity were 32% and 96%, respectively (AUC 0.641, p = 0.002) (Table 5 and Fig. 4). This shows that the TERT promoter gene mutation is probably the most useful in helping to subclassify cellular fibroepithelial tumors that border on cellular fibroadenomas and benign phyllodes tumors. This may also be useful in small biopsies where there is only partial sampling of the tumor and the full architectural features of a phyllodes tumor are not appreciated. Other mutations had AUC values close to the reference line of 0.5 (no real predictability) and p values which were not statistically significant, indicating that their utility for diagnostic discrimination may not be optimal. This was similarly observed when comparing both conventional and cellular fibroadenomas vs benign phyllodes tumors, as well as when comparing conventional fibroadenomas with their cellular counterpart.
Most sequencing studies on fibroadenomas and benign phyllodes tumors involved targeted profiling of MED12 and TERT promoter [11, 12, 57, 58, 61,62,63,64,65,66,67,68,69], while few had done multi-gene assays or exome sequencing [5,6,7,8, 59, 60, 70, 71]. To the best of our knowledge, our study has the largest number of fibroadenomas and cellular fibroadenomas profiled. Although most studies reported a lack of alterations in cancer driver genes in the benign group of fibroepithelial lesions, some studies did reveal such mutations. Our previous exome sequencing of eight fibroadenomas showed a singular RB1 mutation [5], while other investigators found mutant TP53 [72] and PIK3CA [73]. A single case of a pediatric fibroadenoma harboring PIK3CA missense mutation, a juvenile fibroadenoma with mutant MAP3K1, and a pediatric benign fibroepithelial neoplasm with PTEN and BRCA1 mutations, have been reported [59]. Profiling heterogeneous lesions demonstrated TP53 mutation in an FA-like area of a malignant PT, and a PIK3CA mutation in an FA-like area of a borderline PT [74]. It is not yet known whether presence of such mutations could signal possible malignant transformation of fibroadenomas, predict future progression into phyllodes tumors, or indicate likely recurrence of benign phyllodes tumors. However, it is critical to distinguish benign phyllodes tumors from fibroadenomas, since the former has a likelihood for grade progression upon recurrence and its malignant forms can metastasize, although such occurrences are extremely low [14, 15, 75,76,77,78,79,80,81,82,83]. Investigating genetic alterations of paired primary tumors and their recurrent/metastatic lesions could be an area for further exploration to unravel their molecular mechanisms.
When we histologically reviewed fibroadenomas and benign phyllodes tumors with cancer driver mutations, only 4.7% (6/128) of tumors had their original diagnoses revised, with three fibroadenomas reclassified as benign phyllodes tumors, two benign phyllodes tumors reclassified as borderline, and one fibroadenoma reclassified as a borderline/malignant phyllodes tumor. When we reviewed benign phyllodes tumors with mutations in more than two non-cancer driving genes, there were no tumors that were reclassified. While histological assessment remains reliable in classifying these lesions, the possibility of morphological heterogeneity underpinned by the molecular alterations has to be considered. While central review of all histological sections may be useful, these cases were appraised and diagnosed by international pathologist collaborators based on current histological criteria, with findings from this study according additional genomic insights that can help refine light microscopic interpretation in participating institutions.
Pareja et al. proposed that fibroepithelial tumor progression from normal mammary stroma toward phyllodes tumors, occurs either through the MED12-mutant pathway or the MED12-wild-type pathway. Fibroadenomas are the first step in the MED12-mutant pathway, after the stroma acquires mutations in MED12 exon 2 [84]. Our study, like others, confirm that MED12 mutations are not universally present in all breast fibroepithelial lesions [5, 6, 11, 12, 58, 61, 63, 65]. This provides corroborating evidence that phyllodes tumors can indeed develop through the MED12-wild-type pathway by acquisition of alterations in cancer driver genes.
In a study of 23 fibroadenomas using comparative genomic hybridization analysis, Ojopi et al. found that the most frequently overrepresented segments were 5p14, 5q34-qter, 13q32-qter, 10q25-qter, and 18q22 [85]. Amiel et al. detected genetic aberrations in chromosomes 4–6, 8–13, 16, 18, 19, 20, and 22 [86]; while Cavalli et al. and Ried et al. reported no alterations in the DNA copy number of 20 fibroadenomas and 13 fibroadenomas, respectively [87, 88]. Additionally, Xie et al. observed through whole exome sequencing that most of the 27 recurrent somatic CNVs were clustered in chromosomes 1, 4, 9, 11, 13, 15, 19, and were deletions [60].
As for benign phyllodes tumors, Lv et al. noted that chromosomal regions prevalently involved in copy number gains were 1p12-q21 (4/12), 18p11.2-q11.2 (3/12) and losses were most frequently found on chromosome regions 9q34 (3/12), 10q26 (3/12), 17q25 (3/12), 19q13.3 (3/12) [89]. Laé et al. reported losses in 6q (4/9), 10p (1/9), 16q (1/9), and 22q (1/9), and gains in 1q (1/9) [90]. We also observed losses in 19q11-q13.11 (14/15), 10p15.3-p14 (14/15), and 22q11.22-q12.1 (8/15), although our data showed gains in 9q33.3-q34.3 (7/15) instead.
These studies used different experimental techniques (comparative genomic hybridization and whole exome sequencing) compared to our low-pass WGS, thus the data are not entirely comparable. Our assessment of CNV did not show significant differences between the three tumor types (see Supplementary material, Table S1 and Fig. S1) in terms of chromosomal alterations too, hence somatic DNA profiling may be a better tool to differentiate them.
The strength of our study includes the large number of benign phyllodes tumors with representation from various countries. This helps to eliminate bias based on ethnic or geographic differences and adds confidence to our results. The limitations include the restricted panel of 16 genes that are studied which may not fully reveal the genetic differences between fibroadenomas and phyllodes tumors. Approaches such as WGS may help us gain greater insights into the full molecular spectrum of breast fibroepithelial tumors. WGS has already been performed in uterine leiomyomas, which share certain clinical and mutational characteristics with fibroadenomas [91]. In addition, the number of cellular fibroadenomas are disproportionately smaller than the number of benign phyllodes tumors and our results for cellular fibroadenomas may therefore be susceptible to bias. There is also a proportion of cases that did not have available H&E sections for histological review.
In summary, next generation sequencing using a targeted 16 gene panel revealed that benign phyllodes tumors have a higher mean number of mutations compared to fibroadenomas. Cellular fibroadenomas are genetically similar to conventional fibroadenomas. Most of the cancer driver genes studied showed no significant difference in mutation frequencies when comparing benign phyllodes tumors and fibroadenomas, but RB1 and EGFR were notably mutated in a higher proportion of benign phyllodes tumors than fibroadenomas. Benign phyllodes tumors possessed a significantly higher percentage of TERT promoter mutations compared to cellular fibroadenomas and this may be a useful adjunct when evaluating challenging cellular fibroepithelial tumors. Histologic criteria remain reliable in the classification of benign fibroepithelial tumors with only a small percentage of tumors being reclassified with the help of molecular findings. Further study of the genetics of benign fibroepithelial tumors beyond the scope of the 16 gene panel is required to help us better chart the genomic landscape of these tumors.
Data availability
The dataset generated is available in the NCBI Sequence Read Archive (SRA) under accession code PRJNA516727.
References
Tan PH, Sahin AA. Atlas of differential diagnosis in breast pathology. New York: Springer Science, Business Media LLC; 2017.
WHO Classification of Tumours Editorial Board. WHO classification of tumours of the breast. 5th ed. Lyon: IARC Press; 2019.
Krings G, Bean GR, Chen YY. Fibroepithelial lesions; the WHO spectrum. Semin Diagn Pathol. 2017;34:438–52.
Loke BN, Md Nasir ND, Thike AA, Lee JYH, Lee CS, Teh BT, et al. Genetics and genomics of breast fibroadenomas. J Clin Pathol. 2018;71:381–7.
Lim WK, Ong CK, Tan J, Thike AA, Ng CC, Rajasegaran V, et al. Exome sequencing identifies highly recurrent MED12 somatic mutations in breast fibroadenoma. Nat Genet. 2014;46:877–80.
Tan J, Ong CK, Lim WK, Ng CC, Thike AA, Ng LM, et al. Genomic landscapes of breast fibroepithelial tumors. Nat Genet. 2015;47:1341–5.
Cani AK, Hovelson DH, McDaniel AS, Sadis S, Haller MJ, Yadati V, et al. Next-Gen sequencing exposes frequent MED12 mutations and actionable therapeutic targets in phyllodes tumors. Mol Cancer Res. 2015;13:613–9.
Piscuoglio S, Ng CK, Murray M, Burke KA, Edelweiss M, Geyer FC, et al. Massively parallel sequencing of phyllodes tumours of the breast reveals actionable mutations, and TERT promoter hotspot mutations and TERT gene amplification as likely drivers of progression. J Pathol. 2016;238:508–18.
Liu SY, Joseph NM, Ravindranathan A, Stohr BA, Greenland NY, Vohra P, et al. Genomic profiling of malignant phyllodes tumors reveals aberrations in FGFR1 and PI-3 kinase/RAS signaling pathways and provides insights into intratumoral heterogeneity. Mod Pathol. 2016;29:1012–27.
Nozad S, Sheehan CE, Gay LM, Elvin JA, Vergilio JA, Suh J, et al. Comprehensive genomic profiling of malignant phyllodes tumors of the breast. Breast Cancer Res Treat. 2017;162:597–602.
Piscuoglio S, Murray M, Fusco N, Marchiò C, Loo FL, Martelotto LG, et al. MED12 somatic mutations in fibroadenomas and phyllodes tumors of the breast. Histopathology. 2015;67:719–29.
Mishima C, Kagara N, Tanei T, Naoi Y, Shimoda M, Shimomura A, et al. Mutational analysis of MED12 in fibroadenomas and phyllodes tumors of the breast by means of targeted next-generation sequencing. Breast Cancer Res Treat. 2015;152:305–12.
Piscuoglio S, Geyer FC, Burke KA, Murray MP, Ng CK, Mota A, et al. Massively parallel sequencing analysis of synchronous fibroepithelial lesions supports the concept of progression from fibroadenoma to phyllodes tumor. NPJ Breast Cancer. 2016;2:16035.
Tan PH. Fibroepithelial lesions revisited: implications for diagnosis and management. Mod Pathol. 2021;34:15–37.
Tan BY, Acs G, Apple SK, Badve S, Bleiweiss IJ, Brogi E, et al. Phyllodes tumours of the breast: a consensus review. Histopathology. 2016;68:5–21.
Karim RZ, Gerega SK, Yang YH, Spillane A, Carmalt H, Scolyer RA, et al. Phyllodes tumours of the breast: a clinicopathological analysis of 65 cases from a single institution. Breast. 2009;18:165–70.
Kim S, Kim JY, Kim DH, Jung WH, Koo JS. Analysis of phyllodes tumor recurrence according to the histologic grade. Breast Cancer Res Treat. 2013;141:353–63.
Gordon PB, Gagnon FA, Lazkowsky L. Solid breast masses diagnosed as fibroadenoma at fine-needle aspiration biopsy: acceptable rates of growth at long-term follow-up. Radiology. 2003;229:233–8.
Liberman L, Bonaccio E, Hamele-Bena D, Abramson AF, Cohen MA, Dershaw DD. Benign and malignant phyllodes tumors: mammographic and sonographic findings. Radiology. 1996;198:121–4.
Kalambo M, Adrada BE, Adeyefa MM, Krishnamurthy S, Hess K, Carkaci S, et al. Phyllodes tumor of the breast: ultrasound-pathology correlation. Am J Roentgenol. 2018;210:W173–9.
Chao TC, Lo YF, Chen SC, Chen MF. Sonographic features of phyllodes tumors of the breast. Ultrasound Obstet Gynecol. 2002;20:64–71.
Wurdinger S, Herzog AB, Fischer DR, Max C, Raabe G, Schneider A, et al. Differentiation of phyllodes breast tumors from fibroadenomas on MRI. Am J Roentgenol. 2005;185:1317–21.
Lawton TJ, Acs G, Argani P, Farshid G, Gilcrease M, Goldstein N, et al. Interobserver variability by pathologists in the distinction between cellular fibroadenomas and phyllodes tumors. Int J Surg Pathol. 2014;22:695–8.
Yasir S, Gamez R, Jenkins S, Visscher DW, Nassar A. Significant histologic features differentiating cellular fibroadenoma from phyllodes tumor on core needle biopsy specimens. Am J Clin Pathol. 2014;142:362–9.
Lee AH, Hodi Z, Ellis IO, Elston CW. Histological features useful in the distinction of phyllodes tumour and fibroadenoma on needle core biopsy of the breast. Histopathology. 2007;51:336–44.
Tsang AKH, Chan SK, Lam CCF, Lui PCW, Chau HHL, Tan PH, et al. Phyllodes tumours of the breast—differentiating features in core needle biopsy. Histopathology. 2011;59:600–8.
Jara-Lazaro AR, Akhilesh M, Thike AA, Lui PCW, Tse GMK, Tan PH. Predictors of phyllodes tumours on core biopsy specimens of fibroepithelial neoplasms. Histopathology. 2010;57:220–32.
Tan WJ, Cima I, Choudhury Y, Wei X, Lim JCT, Thike AA, et al. A five-gene reverse transcription-PCR assay for pre-operative classification of breast fibroepithelial lesions. Breast Cancer Res. 2016;18:31.
Md Nasir ND, Ng CCY, Rajasegaran V, Wong SF, Liu W, Ng GXP, et al. Genomic characterisation of breast fibroepithelial lesions in an international cohort. J Pathol. 2019;249:447–60.
1000 Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Garrison E, Marth G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907 (2012).
Yang H, Wang K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc. 2015;10:1556–66.
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001;29:308–11.
Forbes SA, Beare D, Gunasekaran P, Leung K, Bindal N, Boutselakis H, et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 2015;43:D805–11.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2016;44:D862–8.
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:D980–5.
Thorvaldsdottir H, Robinson JT, Mesirov JP. Integrative genomics viewer (IGP): high-performance genomics data visualization and exploration. Brief Bioinform. 2013;14:178–92.
Ross DS, Giri DD, Akram MM, Catalano JP, Olcese C, Zee KJV, et al. Fibroepithelial lesions in the breast of adolescent females: a clinicopathological study of 54 cases. Breast J. 2017;23:182–92.
Jacobs TW, Chen YY, Guinee DG Jr, Holden JA, Cha I, Bauermeister DE, et al. Fibroepithelial lesions with cellular stroma on breast core needle biopsy: are there predictors of outcome on surgical excision? Am J Clin Pathol. 2005;124:342–54.
Kim S, Xu X, Hecht A, Boyer TG. Mediator is a transducer of Wnt/beta-catenin signaling. J Biol Chem. 2006;281:14066–75.
Pérot G, Croce S, Ribeiro A, Lagarde P, Velasco V, Neuville A, et al. MED12 alterations in both human benign and malignant uterine soft tissue tumors. PLoS ONE. 2012;7:e40015.
Kang YK, Guermah M, Yuan CX, Roeder RG. The TRAP/Mediator coactivator complex interacts directly with estrogen receptors alpha and beta through the TRAP220 subunit and directly enhances estrogen receptor function in vitro. Proc Natl Acad Sci USA. 2002;99:2642–7.
Toska E, Osmanbeyoglu HU, Castel P, Chan C, Hendrickson RC, Elkabets M, et al. PI3K pathway regulates ER-dependent transcription in breast cancer through the epigenetic regulator KMT2D. Science. 2017;355:1324–30.
Ross-Innes CS, Stark R, Holmes KA, Schmidt D, Spyrou C, Russell R, et al. Cooperative interaction between retinoic acid receptor-alpha and estrogen receptor in breast cancer. Genes Dev. 2010;24:171–82.
Ding X, Li L, Tang C, Meng C, Xu W, Wei X, et al. Cytoplasmic expression of estrogen receptor β may predict poor outcome of EGFR‑TKI therapy in metastatic lung adenocarcinoma. Oncol Lett. 2018;16:2382–90.
Tse GMK, Lee CS, Kung FYL, Scolyer RA, Law BKB, Lau T, et al. Hormonal receptors expression in epithelial cells of mammary phyllodes tumors correlates with pathologic grade of the tumor. Am J Clin Pathol. 2002;118:522–6.
Tan WJ, Chan JY, Thike AA, Lim JCT, Md Nasir ND, Tan JSY, et al. MED12 protein expression in breast fibroepithelial lesions: correlation with mutation status and oestrogen receptor expression. J Clin Pathol. 2016;69:858–65.
Luo XL, Deng CC, Su XD, Wang F, Chen Z, Wu XP, et al. Loss of MED12 induces tumor dormancy in human epithelial ovarian cancer via downregulation of EGFR. Cancer Res. 2018;78:3532–43.
Nagaria TS, Shi C, Leduc C, Hoskin V, Sikdar S, Sangrar W, et al. Combined targeting of Raf and Mek synergistically inhibits tumorigenesis in triple negative breast cancer model systems. Oncotarget. 2017;8:80804–19.
Bemanian V, Sauer T, Touma J, Lindstedt BA, Chen Y, Ødegård HP, et al. The epidermal growth factor receptor (EGFR/HER-1) gatekeeper mutation T790M is present in European patients with early breast cancer. PLoS ONE. 2015;10:e0134398.
Bethune G, Bethune D, Ridgway N, Xu Z. Epidermal growth factor receptor (EGFR) in lung cancer: an overview and update. J Thorac Dis. 2010;2:48–51.
Kim N, Cho D, Kim H, Kim S, Cha YJ, Greulich H, et al. Colorectal adenocarcinoma-derived EGFR mutants are oncogenic and sensitive to EGFR-targeted monoclonal antibodies, cetuximab and panitumumab. Int J Cancer. 2020;146:2194–200.
Yun CH, Mengwasser KE, Toms AV, Woo MS, Greulich H, Wong KK, et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci USA. 2008;105:2070–5.
Cimino-Mathews A, Hicks JL, Sharma R, Vang R, Illei PB, De Marzo A, et al. A subset of malignant phyllodes tumors harbors alterations in the Rb/p16 pathway. Hum Pathol. 2013;44:2494–500.
Yoshida M, Ogawa R, Yoshida H, Maeshima A, Kanai Y, Kinoshita T. et al. TERT promoter mutations are frequent and show association with MED12 mutations in phyllodes tumors of the breast. Br J Cancer. 2015;113:1244–8.
Garcia-Dios DA, Levi D, Shah V, Gillett C, Simpson MA, Hanby A, et al. MED12, TERT promoter and RBM15 mutations in primary and recurrent phyllodes tumours. Br J Cancer. 2018;118:277–84.
Tay TKY, Guan P, Loke BN, Md Nasir ND, Rajasegaran V, Thike AA, et al. Molecular insights into paediatric breast fibroepithelial tumours. Histopathology. 2018;73:809–18.
Xie SN, Cai YJ, Ma B, Xu Y, Qian P, Zhou JD, et al. The genomic mutation spectrums of breast fibroadenomas in Chinese population by whole exome sequencing analysis. Cancer Med. 2019;8:2372–9.
Yoshida M, Sekine S, Ogawa R, Yoshida H, Maeshima A, Kanai Y, et al. Frequent MED12 mutations in phyllodes tumours of the breast. Br J Cancer. 2015;112:1703–8.
Nagasawa S, Maeda I, Fukuda T, Wu W, Hayami R, Kojima Y, et al. MED12 exon 2 mutations in phyllodes tumors of the breast. Cancer Med. 2015;4:1117–21.
Pfarr N, Kriegsmann M, Sinn P, Klauschen F, Endris V, Herpel E, et al. Distribution of MED12 mutations in fibroadenomas and phyllodes tumors of the breast—implications for tumor biology and pathological diagnosis. Genes Chromosomes Cancer. 2015;54:444–52.
Ng CC, Tan J, Ong CK, Lim WK, Rajasegaran V, Nasir ND, et al. MED12 is frequently mutated in breast phyllodes tumours: a study of 112 cases. J Clin Pathol. 2015;68:685–91.
Lien HC, Huang CS, Yang YW, Jeng YM. Mutational analysis of MED12 exon 2 in a spectrum of fibroepithelial tumours of the breast: implications for pathogenesis and histogenesis. Histopathology. 2016;68:433–41.
Yoon N, Bae GE, Kang SY, Choi MS, Hwang HW, Kim SW, et al. Frequency of MED12 mutations in phyllodes tumors: inverse correlation with histologic grade. Genes Chromosomes Cancer. 2016;55:495–504.
Lae M, Gardrat S, Rondeau S, Richardot C, Caly M, Chemlali W, et al. MED12 mutations in breast phyllodes tumors: evidence of temporal tumoral heterogeneity and identification of associated critical signaling pathways. Oncotarget. 2016;7:84428–38.
Darooei M, Khan F, Rehan M, Zubeda S, Jeyashanker E, Annapurna S, et al. MED12 somatic mutations encompassing exon 2 associated with benign breast fibroadenomas and not breast carcinoma in Indian women. J Cell Biochem. 2019;120:182–91.
Pareja F, Da Cruz Paula A, Murray MP, Hoang T, Gularte-Merida R, Brown D, et al. Recurrent MED12 exon 2 mutations in benign breast fibroepithelial lesions in adolescents and young adults. J Clin Pathol. 2019;72:258–62.
Kim JY, Yu JH, Nam SJ, Kim SW, Lee SK, Park WY, et al. Genetic and clinical characteristics of phyllodes tumors of the breast. Transl Oncol. 2018;11:18–23.
Sim Y, Ng GXP, Ng CCY, Rajasegaran V, Wong SF, Liu W, et al. A novel genomic panel as an adjunctive diagnostic tool for the characterization and profiling of breast Fibroepithelial lesions. BMC Med Genomics. 2019;12:142.
Millikan R, Hulka B, Thor A, Zhang Y, Edgerton S, Zhang X, et al. p53 mutations in benign breast tissue. J Clin Oncol. 1995;13:2293–300.
Vorkas PA, Poumpouridou N, Agelaki S, Kroupis C, Georgoulias V, Lianidou ES. PIK3CA hotspot mutation scanning by a novel and highly sensitive high-resolution small amplicon melting analysis method. J Mol Diagn. 2010;12:697–704.
Tan BY, Md Nasir ND, Chang HY, Ng CCY, Guan P, Nagarajan S, et al. Morphologic and genetic heterogeneity in breast fibroepithelial lesions-a comprehensive mapping study. Mod Pathol. 2020;33:1732–45.
Tan PH, Thike AA, Tan WJ, Thu MM, Busmanis I, Li H, et al. Predicting clinical behaviour of breast phyllodes tumours: a nomogram based on histological criteria and surgical margins. J Clin Pathol. 2012;65:69–76.
Jones AM, Mitter R, Springall R, Graham T, Winter E, Gillett C, et al. A comprehensive genetic profile of phyllodes tumours of the breast detects important mutations, intra-tumoral genetic heterogeneity and new genetic changes on recurrence. J Pathol. 2008;214:533–44.
Mitus J, Adamczyk A, Majchrzyk K, Kowalik A, Ryś J, Niemiec J. Comparison of mutation profile between primary phyllodes tumors of the breast and their paired local recurrences. Pol J Pathol. 2020;71:7–12.
Koh VCY, Thike AA, Tan PH. Distant metastases in phyllodes tumours of the breast: an overview. Appl Cancer Res. 2017;37:15.
Reinfuss M, Mituś J, Duda K, Stelmach A, Ryś J, Smolak K. The treatment and prognosis of patients with phyllodes tumor of the breast: an analysis of 170 cases. Cancer. 1996;77:910–6.
Mangi AA, Smith BL, Gadd MA, Tanabe KK, Ott MJ, Souba WW. Surgical management of phyllodes tumors. Arch Surg. 1999;134:487–92.
Chaney AW, Pollack A, McNeese MD, Zagars GK, Pisters PW, Pollock RE, et al. Primary treatment of cystosarcoma phyllodes of the breast. Cancer. 2000;89:1502–11.
Asoglu O, Ugurlu MM, Blanchard K, Grant CS, Reynolds C, Cha SS, et al. Risk factors for recurrence and death after primary surgical treatment of malignant phyllodes tumors. Ann Surg Oncol. 2004;11:1011–7.
Abdalla HM, Sakr MA. Predictive factors of local recurrence and survival following primary surgical treatment of phyllodes tumors of the breast. J Egypt Natl Canc Inst. 2006;18:125–33.
Pareja F, Geyer FC, Kumar R, Selenica P, Piscuoglio S, Ng CKY, et al. Phyllodes tumors with and without fibroadenoma-like areas display distinct genomic features and may evolve through distinct pathways. NPJ Breast Cancer. 2017;3:40.
Ojopi EP, Rogatto SR, Caldeira JR, Barbiéri-Neto J, Squire JA. Comparative genomic hybridization detects novel amplifications in fibroadenomas of the breast. Genes Chromosomes Cancer. 2001;30:25–31.
Amiel A, Kaufman Z, Goldstein E, Bruchim RB, Kidron D, Gaber E, et al. Application of comparative genomic hybridization in search for genetic aberrations in fibroadenomas of the breast. Cancer Genet Cytogenet. 2003;142:145–8.
Cavalli LR, Cornelio DA, Lima RS, Urban CA, Rone JD, Cavalli IJ, et al. Lack of DNA copy number alterations revealed with comparative genomic hybridization in fibroadenomas of the breast. Cancer Genet Cytogenet. 2004;153:173–6.
Ried T, Just KE, Holtgreve-Grez H, du Manoir S, Speicher MR, Schröck E, et al. Comparative genomic hybridization of formalin-fixed, paraffin-embedded breast tumors reveals different patterns of chromosomal gains and losses in fibroadenomas and diploid and aneuploid carcinomas. Cancer Res. 1995;55:5415–23.
Lv S, Niu Y, Wei L, Liu Q, Wang X, Chen Y. Chromosomal aberrations and genetic relations in benign, borderline and malignant phyllodes tumors of the breast: a comparative genomic hybridization study. Breast Cancer Res Treat. 2008;112:411–8.
Laé M, Vincent-Salomon A, Savignoni A, Huon I, Fréneaux P, Sigal-Zafrani B, et al. Phyllodes tumors of the breast segregate in two groups according to genetic criteria. Mod Pathol. 2007;20:435–44.
Mehine M, Kaasinen E, Makinen N, Katainen R, Kampjarvi K, Pitkanen E, et al. Characterization of uterine leiomyomas by whole-genome sequencing. N Engl J Med. 2013;369:43–53.
Acknowledgements
We extend our gratitude to the International Fibroepithelial Consortium for their contribution of samples and clinical data.
International Fibroepithelial Consortium
Norraha Abd Rahman11, S. M. Khodeza Nahar Begum12, Phaik Leng Cheah13, Chih Jung Chen14, Emmanuel Dela Fuente15, Aaron Han16, Oi Harada17, Naoki Kanomata18, Cheok Soon Lee19, Jonathan Yu Han Lee20, Mohammed Kamal21, Rieko Nishimura22, Yasuyo Ohi23, Elinor J. Sawyer24, Kean Hooi Teoh13, Alex Koon Ho Tsang25, Julia Yuen-Shan Tsang26, Gary M. K. Tse26, Rin Yamaguchi27
Funding
This study was funded by a SingHealth Duke-NUS Research Collaborative Grant (SDC/021/2015) and an SHF-Foundation Research Grant (SHF/PRISM018/2017), as awarded by the SingHealth Foundation and Duke-NUS; as well as a grant from the Singapore Ministry of Health’s National Medical Research Council (MOH-STAR18NOV-0001).
Author information
Authors and Affiliations
Consortia
Contributions
PHT, BTT, PT, GWCY, and BHB were responsible for study conceptualization and design, and project supervision. CCYN, NDMN, BNL, VR, WL, and JYL performed the experiments. KTEC, MAG, PM, BKTT, VKMT, CYW, WSY, GHH, KWO, and IFC provided resources and clinical data. CCYN, NDMN, BNL, VR, WL, JYL, PG, AHL, AAT, and TKYT collected, analyzed, and interpreted the data. CCYN, NDMN, BNL, PG, and AHL wrote the manuscript. All the authors revised and provided input to the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
PT, BTT, and PHT jointly hold patent applications for PCT/SG2015/050107 (Breast fibroadenoma susceptibility mutations and use thereof) and PCT/SG2015/050368 (Method and kit for pathologic grading of breast neoplasms). Other authors declare no competing interests.
Ethical approval
This study was approved by the SingHealth Centralized Institutional Review Board (CIRB Ref: 2018/2581), with a waiver of informed consent granted.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Members of the International Fibroepithelial Consortium are listed below Acknowledgements
Supplementary information
Rights and permissions
About this article
Cite this article
Ng, C.C.Y., Md Nasir, N.D., Loke, B.N. et al. Genetic differences between benign phyllodes tumors and fibroadenomas revealed through targeted next generation sequencing. Mod Pathol 34, 1320–1332 (2021). https://doi.org/10.1038/s41379-021-00787-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00787-w
This article is cited by
-
Epithelial to mesenchymal transition (EMT) in metaplastic breast cancer and phyllodes breast tumors
Medical Oncology (2023)
-
Discrimination between phyllodes tumor and fibro-adenoma: Does artificial intelligence-aided mammograms have an impact?
Egyptian Journal of Radiology and Nuclear Medicine (2022)
-
Fibroepithelial tumours of the breast—a review
Virchows Archiv (2022)
-
Artificial intelligence modelling in differentiating core biopsies of fibroadenoma from phyllodes tumor
Laboratory Investigation (2022)