Abstract
Breast fibroepithelial lesions (FELs) encompass the common fibroadenoma (FA) and relatively rare phyllodes tumour (PT); the latter entity is usually classified as benign, borderline or malignant. Intratumoural heterogeneity is frequently present in these tumours, making accurate histologic evaluation challenging. Despite their rarity, PTs are an important clinical problem due to their propensity for recurrence and, in the case of malignant PT, metastasis. Surgical excision is the mainstay of management. Recent work has uncovered myriad genetic alterations in breast FELs. In this study, exome sequencing was performed on seven cases of morphologically heterogeneous breast FELs, including FAs, PTs of all grades, and a case of metaplastic spindle cell carcinoma arising in PT, in order to elucidate their intratumoural genetic repertoire. Gene mutations identified encompassed cell signalling, tumour suppressor, DNA repair and cell cycle regulating pathways. Mutations common to multiple tumour regions generally showed higher variant allele frequency. Frequent mutations included MED12, TP53, RARA and PIK3CA. Histological observations of increased cellular density and pleomorphism correlated with mutational burden. Phylogenetic analyses revealed disparate pathways of possible tumour progression. In summary, histological heterogeneity correlated with genetic changes in breast FELs.
Similar content being viewed by others
Introduction
Breast fibroepithelial lesions (FELs), comprising fibroadenomas (FAs) and phyllodes tumours (PTs), represent a distinct group of biphasic breast tumours characterized by the intimate participation of epithelium and stroma [1]. FAs are benign, while PTs range from histologically bland to frankly malignant. Although constituting 0.5–1.5% of breast tumours in Western series [2], PTs have been documented to be more common in Asian populations, with reported incidences of up to 6.9% [3, 4], when compared with the frequency of breast cancers. Malignant PTs, behaviourally akin to sarcomas, have the potential for significant morbidity and mortality, in the form of recurrences and distant metastases [5].
Current PT grading, as proposed by the WHO Working Group [1], is based on histological integration of multiple parameters on a semi-quantitative basis, which despite predictive utility across cohorts, cannot accurately ascertain clinical behaviour in an individual patient. This is aside from significant interobserver variability even amongst experienced pathologists [6, 7]. A nomogram has been developed by our group, which incorporates histological parameters and status of surgical margins as an individualized risk-assessment tool [8]. This nomogram has been validated across diverse populations [9, 10]. Nonetheless, long-standing issues pertaining to the diagnosis and management of PT remain [11], which relate to difficulties in pathological diagnosis and the lack of robust molecular predictors of outcome.
While the accepted treatment of PT is complete surgical excision, tumour recurrence remains an important clinical problem [5]. Recurrence rates in a series of PTs were 10.9%, 14.4% and 29.6% for benign, borderline and malignant tumours, respectively [8]. The role of adjuvant therapy is controversial and is currently not a routine recommendation.
Intratumoural heterogeneity is a notable feature of PT. Malignant PTs may harbour heterologous stromal elements with heterogeneous histological and immunohistochemical attributes, which can hold prognostic and therapeutic importance. Array- comparative genomic hybridization (CGH) analysis of paired primary and recurrent tumours showed acquisition of new genetic changes in the progression from benign to malignant PT phenotypes, which allude to the presence of sub-clones not easily identifiable by histological evaluation alone [12].
Despite myriad biological markers that have been studied in PTs, many showing an association with grade, their use in defining grade and determining clinical behaviour in specific cases remains limited at this time [13,14,15,16,17,18,19,20,21]. The role of each tumour compartment, including morphologically heterogeneous components, as well as their molecular characteristics in lesional evolution and malignant transformation, is currently not well-defined [12, 22, 23], although small series have revealed novel genomic aberrations [24].
In this study, we comprehensively mapped and analyzed seven cases of breast FELs, in order to elucidate the genetic alterations in morphologically heterogeneous tumour foci.
Methods
This study was approved by the institutional review board of Singapore General Hospital (SGH), Singhealth, Singapore. Seven cases of breast FELs (including FAs and PTs) that demonstrated morphologic heterogeneity were retrieved from the archives of the Department of Anatomical Pathology, SGH, spanning 2006 and 2015. Clinical information was obtained from institutional electronic case records. All cases were re-reviewed, with diagnoses and tumour grades independently confirmed by two pathologists using standard criteria [1]. Regions were selected based on the presence of morphological heterogeneity.
Macrodissection
Eight serial unstained formalin-fixed paraffin-embedded (FFPE) sections (10 μm thick) from each region were cut on a microtome (Leica) per case, mounted onto glass slides and stored at −20 °C until required. A corresponding haematoxylin and eosin (HE)-stained slide was reviewed and marked by a pathologist for regions of interest (ROIs), comprising tumour and adjacent normal areas. The reverse side of the unstained slide was then drawn with a marker, guided by the HE slide as a reference. Tissue samples within the marked areas were then scraped and collected into a 1.5 mL microcentrifuge tube using disposable sterile surgical blades.
Exome sequencing
Matched normal tissue and tumour ROIs were selected from each tumour for whole-exome sequencing to profile protein-coding regions of the genomes. Genomic DNA was extracted using a Qiagen kit on FFPE tissue. FFPE genomic DNA yield and quality were determined using PicoGreen fluorometric analysis and an in-house developed multiplex PCR system. Only samples that passed quality control checks for quality and minimum quantity (>200 ng) were pursued for exome sequencing. Extracted genomic DNA was subjected to fragmentation with the Covaris S2 system following recommended settings. Sequencing library preparation was performed with sequencing adaptor ligation using the Kappa hyper prep kit (Kappa Biosystems). Enrichment of coding sequences was performed by employing the SureSelectXT Human All Exon v5 (51 Mb) kit (Agilent Technologies) according to the manufacturer’s recommended protocol. Exome-enriched libraries were then sequenced on Illumina’s HiSeq 3000/4000 sequencing platform to generate 150-bp paired-end reads.
Paired reads for all samples were mapped to hg19 using BWA-MEM [25]. Raw mappings were processed for marking duplicates and local realignment using Picard tools (http://broadinstitute.github.io/picard/) and GATK [26]. Qualities of the mapping were checked using Qualimap [27, 28]. Somatic mutations were called using consensus approaches with MuSE [29], MuTect2 [30], Scalpel [31], Strelka [32] and VarScan2 [33, 34] and subsequently annotated using wANNOVAR [35], where synonymous variants excluded. Confident non-synonymous variants were those satisfying all the following criteria:
-
(1)
Reported by at least two variant callers.
-
(2)
Not in in dbSNP [36] unless it is in COSMIC [37] or ClinVar [38, 39] databases.
-
(3)
Minor allele frequency in the population is no more than 0.1%.
Discrete-characters Wagner parsimony method (pars) in the PHYLIP [40] package was used to generate the phylogenetic trees using the following settings:
-
(1)
“Outgroup root” is the control normal sample.
-
(2)
“Randomize input order of species” is set to “Yes” (seed = 9999).
Results
Case 1 (PT1)
A 52-year-old Chinese female with a left breast tumour underwent a left simple mastectomy following a diagnosis of high-grade sarcoma on trucut biopsy. Histology showed a malignant PT measuring 13.5 cm in maximum dimension. The tumour comprised an area of relatively banal, leaf-like stromal fronds morphologically reminiscent of a benign PT, adjacent to highly cellular sheets and fascicles of spindled cells displaying brisk mitotic activity (31/10hpf) with focal infarction and hyalinization. The background displayed a myxoid quality. Skeletal muscle invasion was present, as well as skin ulceration. No malignant heterologous element was identified.
Exome sequencing was performed on the region morphologically akin to a “benign” PT, as well as two regions of frankly malignant PT, one of which was adjacent to the “benign PT” region, and the other at an area of stromal permeation:
-
A—malignant PT component (area of stromal permeation).
-
B—malignant PT component.
-
C—more classic, “benign PT”-appearing component.
All three regions shared mutations in ADAMTS17, MFSD10, EZH1, FAM13C, NCOR2, KMT2D, ATP1A3, DDAH2, TRIM72, RORC and ATF7. In addition, regions A and B, which were histologically malignant, had common TMEM8A and FOSL2 mutations not found in region C (Fig. 1).
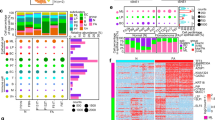
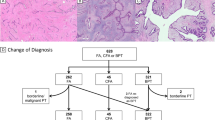
Light microscopy images at low magnification show malignant PT regions (a, b) of case PT1, and a more “benign PT” appearing component (c); and of case PT2’s borderline PT (d) diagnosed in 2007, and spindle cell metaplastic carcinoma (e) in 2013. The latter shows diffuse cytoplasmic MNF116 immunoreactivity. The tables reflect their corresponding mutations and variant allele frequencies (darker shades of red reflect higher VAF, closer to 100%; darker shades of blue reflect lower VAF, closer to 0%). Phylogenetic trees indicate genes mutated in all samples (blue), those mutated in at least two samples (green), and those mutated in one sample (red).
Mutations shared across multiple tumour samples of PT1 had comparable and higher variant allele frequency since they were likely to be early events in tumourigenesis. The mutation pattern supported a linear progression model in this tumour.
No recurrences were documented at the time of this study (5 years from last surgery).
Case 2 (PT2)
A 65-year-old Chinese female first presented with a right breast lump in 2007, which revealed histological features in keeping with a borderline PT, exhibiting mild-to-moderate stromal nuclear pleomorphism, readily discernible mitoses (7/10hpf), and focal stromal overgrowth. Tumour borders could not be accurately assessed, due to tumour extension to cauterized margins. However, where assessable, focally infiltrative margins were observed. There were no features of concomitant or concurrent metaplastic carcinoma, morphologically or immunohistochemically.
The patient was lost to follow-up for 6 years until 2013, when she presented with a right breast mass over the previous excision site. The tumour measured 2.5 cm in maximum dimension and comprised intersecting fascicles of plump spindled cells with frequent mitoses (13/10hpf), irregular permeative margins, and rare, diminutive, intratumoural epithelial nests. Foci of necrosis were present. The tumour cells showed diffuse cytoplasmic MNF116 and SMA immunoreactivity with patchy calponin, smooth muscle myosin heavy chain and p63 staining, and non-specific cytoplasmic beta-catenin positivity. No significant CK7, CK14, 34-beta-E12 or CD34 staining was seen. ER, PR and c-erb-B2 were negative. The overall immunomorphological findings were regarded as most in keeping with a spindle cell metaplastic carcinoma, with immunopositivity for smooth muscle markers possibly reflecting myoepithelial differentiation.
Exome sequencing was performed on a representative section from each tumour:
-
D—borderline PT (2007).
-
E—spindle cell metaplastic carcinoma (2013).
Alterations in MED12, CMTM5, DBF4B, C3orf67, STARD9, PIN1, NF1 and GIT2 were shared by both regions, which were obtained from tumours temporally separated by 6 years (Fig. 1).
The mutation pattern supported a linear progression model in this tumour.
The patient was subsequently lost to follow-up.
Case 3 (PT3)
A 59-year-old Chinese female presented with four right-sided breast nodules which were excised. Three of the nodules, ranging in size from 0.6 to 2.3 cm, showed features of hyalinized FAs, while the remaining nodule exhibited features of a borderline PT, measuring 5.8 cm in maximum dimension. The PT revealed leaf-like stromal fronds, mild stromal atypia, stromal hypercellularity, focal stromal overgrowth and 11 mitoses/10hpf. Focally permeative tumour margins were observed.
Exome sequencing was performed on a representative region of the PT and the largest FA, respectively:
-
F—borderline PT.
-
G—hyalinised FA.
No common genetic alterations were found between the two regions. Among numerous mutations, region F notably demonstrated mutations in MED12 and RB1, and region G in KMT2D (Fig. 2).
The lack of shared gene mutations among the regions suggested a parallel progression model in this tumour.
No recurrences were documented at the time of this study (4 years from last surgery).
Case 4 (PT4)
A 69-year-old Chinese female underwent a right simple mastectomy for a right breast mass measuring 8 cm in maximum dimension. A prior core biopsy had rendered a diagnosis of a spindle cell neoplasm. The tumour in the mastectomy contained foci morphologically reminiscent of FA, as well as areas of stromal hypercellularity, overgrowth and focally permeative margins, consistent with a borderline PT. The stromal cells were highlighted by CD34 and vimentin, while showing no significant AE1/3 and calponin staining.
Exome sequencing was performed on an FA-like area, as well as two areas of PT stroma:
-
H—FA-like area.
-
I—PT stroma.
-
J—PT stroma.
All three regions demonstrated genetic alterations in ZNF86, AMPD1, SLC23A1, ZNF630, NR1D2, FBRS, ZBTB7A, SLC22A6, TNXB, CFAP20, CD3EAP and PPP1R3F. Regions I and J, in addition, exhibited common NRAS, SLC13A2, BCCIP, C8orf58, BCCIP, ERBB4 and KLHL23 mutations, which were not found in the histologically dissimilar region H. Region H was also noted to exhibit PIK3CA mutation (Fig. 2).
The mutation pattern supported a linear progression model in this tumour. Regions I and J appeared genetically “closer” in tumourigenesis as compared with region H, which paralleled the morphologic findings.
At the same setting, a left mastectomy was performed for a synchronous left invasive ductal carcinoma. The patient presented 3 years later with biopsy-proven metastatic carcinoma involving the liver, porta hepatis, spleen and periaortic region, and died shortly thereafter.
Case 5 (PT5)
A 38-year-old Malay female underwent wide excision of a left breast mass measuring 5 cm in maximum dimension. Histology revealed a heterogeneous appearance, with relatively banal foci morphologically reminiscent of FA, as well as areas of marked stromal cellularity, pronounced nuclear pleomorphism, and irregular margins, in keeping with malignant PT. In addition, a few areas showed stromal disposition around benign epithelial elements in a fashion evocative of periductal stromal tumour (PST).
Exome sequencing was performed on five areas: two FA-like foci, two PST-like foci, and one area of frankly malignant PT stroma:
-
K—FA-like area.
-
L—FA-like area.
-
M—PST-like area.
-
N—PST-like area.
-
O—malignant PT area.
All five regions shared PER3 mutations. Regions L, M, N and O had common mutations in ADAMTS16, SLC5A4 and TP53. In addition, regions M, N and O possessed common KLHDC2, MGAT3, NADK2, PDGFRB and UBR4 mutations, in contradistinction to regions K and L, which appeared histologically banal and lacked these mutations (Fig. 3).
Regions M, N and O shared mutations across multiple regions with comparable and higher variant allele frequency, suggestive of a linear progression model. However, the paucity of shared mutations for regions K and L suggested parallel progression in relation to the other three regions.
No recurrences were documented at the time of this study (4 years from last surgery).
Case 6 (PT6)
A 69-year-old Chinese female with a left breast tumour underwent left simple mastectomy. No information on any prior core biopsy was available. On gross examination, three separate tumour nodules in close proximity were seen, measuring 16, 2 and 2 cm, respectively in maximum dimensions. The patient passed away a year after the diagnosis.
The dominant tumour exhibited marked histologic heterogeneity of the stromal component, ranging from relatively uniform, small, rounded cells in a chondromyxoid background, to pleomorphic spindled cells focally arranged in a vague herringbone pattern, as well as groups of plump cells with rhabdoid morphology. Several multinucleated giant cells were present. Frequent mitoses, up to 47 per 10 high power fields (hpfs), were identified. The tumour border was permeative, with focal invasion into underlying skeletal muscle. In places, epithelium-lined fronds were identified; infarction was present, though with preservation of “ghost” leaf-like architecture.
Diffuse staining of tumour cells for CAM5.2 was observed in some areas, with similar but less intense staining noted with MNF116. EMA, AE1/3, CK5/6, CK7, CK14 and p63 were negative. No significant CD34, CD31, D2-40, Factor VIII or S100 staining was seen. The rhabdoid cells were focally highlighted by desmin and less strongly by SMA, while myogenin and myoD1 showed no significant staining.
The second and third tumour nodules showed similar histological appearances, comprising malignant spindled cells arranged in a vague storiform fashion, with occasional rhabdoid cells.
Exome sequencing was performed on three histologically heterogeneous areas of the dominant tumour nodule, and on one region each of the two smaller tumour nodules:
-
P—dominant tumour (spindled cells within chondromyxoid stroma).
-
Q—dominant tumour (rhabdoid cells).
-
R—dominant tumour (rhabdoid cells, with keratin positivity).
-
S—second tumour nodule.
-
T—third tumour nodule.
All five regions shared myriad mutations, including MED12, ANK1, SPTY2D1, CCDC151, CDK4, KDM6A, SYNDIG1L, GPRIN3, WDFY4, SLC2A10, SLC22A23 and CDH7. Regions Q, R, S and T shared mutations in TP53 and SVEP1, while regions P and Q had shared mutations in COL3A1 and IL22RA1 (Fig. 3).
Of note, region R, which stained positive for cytokeratin markers on immunohistochemical interrogation, was found to harbour mutations in EGFR and KRTAP5-5 (keratin-associated protein 5-5).
As alluded to above, mutations shared across multiple tumour regions possessed higher variant allele frequency, which was expected since they likely represented early events of tumourigenesis.
A linear branching phylogenetic tree supported a linear progression model in this tumour.
Case 7 (PT7)
A Chinese female with a history of multiple FELs first presented in 1992, aged 20, with a left-sided FA. She subsequently underwent excision biopsies for a left-sided FA in 2007, a left PT in 2015 and a left FA in 2015.
The PT, which measured 14 cm in maximum dimension, showed borderline features, including variable stromal cellularity with peri-epithelial accentuation, mitoses up to 11/10hpf, and focally permeative borders with adipocytic entrapment. Stromal atypia was mild. No significant stromal overgrowth or malignant heterologous stromal element was identified. Tumour abutted the excision margin. The background showed focal myxoid oedema and haemorrhage.
The FAs showed banal features.
Exome sequencing was performed on a selected region from each of the five tumours:
-
U, V and W (1992, 1995, 2007 lesions)—FAs.
-
X—borderline PT.
-
Y—FA.
No recurrences were documented at the time of this study (3 years from last surgery).
No common genes were mutated across all regions. MED12 and RARA mutations were found in both regions X and Y (Fig. 4).
The starburst phylogenetic tree pattern suggested a parallel progression model in this tumour.
In summary, out of 25 regions analyzed, the most common recurrently mutated genes were MED12 and TP53 (Table 1, Fig. 5).
MED12 was mutated in ten regions (one FA, three borderline PTs, five malignant PTs and one metaplastic carcinoma arising in the background of previous PT).
TP53 mutation was observed in eight regions (one FA-like area within a malignant PT, two PST-like areas within a malignant PT and five malignant PTs).
Of note, RARA mutation was found in two regions (one FA and one borderline PT), while PIK3CA mutation was elucidated in one region (one FA-like area within a borderline PT).
Discussion
Multi-region exome sequencing identified a diverse set of mutations among morphologically distinct lesions. Borderline and malignant PTs disclosed cancer-related aberrations which were not seen in their FA counterparts, such as RB1 (in PT3) and ERBB4 (in PT4). However, one FA region in PT5 had a similar missense TP53 G73T mutation as its neighbouring malignant PT, and one FA-like area in PT4 harboured a PIK3CA mutation, concurring with our recent finding [41] that it is not impossible for FAs to also harbour such mutations.
Other somatic alterations could have contributed to the mesenchymal histology in driving differentiation and proliferation. PDGFRB was mutated in the malignant PT regions of PT1 and PT5, but not in the patients’ benign PT and FA components, respectively. The gene encodes for platelet-derived growth factor receptor B, and activating mutations were found in gastrointestinal stromal tumours, while breast and lung cancer cells acquire PDGFR expression following epithelial-to-mesenchymal transition (EMT), promoting metastasis [42]. Kim et al. and Pareja et al. reported PDGFRB mutation in a malignant PT (1/13, 8% and 1/11, 9% respectively) [43, 44], while Moon et al. observed PDGFRA and PDGFRB alterations in a set of malignant PTs [45]. In contrast, Carvalho et al. found no activating PDGFRA mutations in 19 PTs (13 benign and 6 malignant) [46]. It may be interesting to study the clinical relevance of this aberration, as targeted treatment against the PDGFR pathway was noted to suppress PT growth in vivo [45].
Another recurrently mutated gene was NRAS, observed in malignant PT (case PT6), in the benign PT component of malignant PT (case PT1), and borderline PT (case PT4). It encodes for the GTPase NRas protein, and its mutation results in an active GTP state. It activates PI3K signalling and MAPK pathway, leading to cell growth, differentiation and survival [47]. Our finding is in line with that of Pareja et al and Jardim et al, who observed NRAS mutation in malignant PTs (1/11, 9% and 1/1, 100% respectively) [44, 47].
The spindle cell metaplastic carcinoma (region E) of case PT2 harboured OR1A1 and insulin-like growth factor-1 receptor (IGF1R) mutations, which were absent in a prior borderline PT. OR1A1 belongs to the class A G-protein-coupled receptor. While its function has yet to be fully characterized, other olfactory receptor (OR) genes such OR2W3 and OR2B6 were found to be correlated to breast cancer progression [48]. Another member of the olfactory subfamily, prostate-specific G-protein-coupled receptor, could also activate NF-κB through the AKT pathway and induce EMT [49]. Activation of IGF1R could result in downstream activation of two signalling pathways, namely, IRS-1/PI3K/Akt and Ras/Raf/ERK pathways. Both pathways regulate transcription factors of ZEB, Snail and Twist families, which are key markers in causing EMT [50]. Whether metaplastic carcinoma progresses from a fibroepithelial tumour remains an area for further investigation, as malignancy can occur in the epithelial compartment of these tumours [51, 52]. A case study by Muller et al., in which a benign PT first recurred as a malignant PT, and later on as a malignant PT with coexisting spindle cell metaplastic breast carcinoma in the same location over a 7-year period, suggests that this phenomenon may occur [53].
Both borderline PT (region D) and spindle cell metaplastic carcinoma (region E) displayed similar MED12 p.G44D and PIN1 p.K95N alterations, which may suggest a relationship between the two tumours. PIN1 is overexpressed in multiple cancers, including prostate, breast, lung, ovary and cervical tumours [54], reported to upregulate >50 oncogenes or proliferation-promoting factors and inhibit >20 tumour suppressors [55]. PIN1 also promotes EMT by increasing the transcriptional activity of STAT3 in MCF-7 breast cancer cell lines, augmenting invasion and metastasis of cancer cells by activating NF-κB, p53, β-catenin and BRD [56].
Identical nucleotide and protein changes were observed for those alterations harboured across all regions of respective cases, except cases PT3 and PT7, which did not have common mutations. This may indicate clonal relatedness between tumours, despite being of different histological phenotypes (see Supplementary material, Table S1). FAs had historically been regarded as polyclonic stromal and epithelial proliferations [57], while PTs possessed monoclonal stromal growth [58]. However, monoclonality in FA stroma and polyclonality in PT stroma were reported in one PCR-based clonality assay study [59]. Although previously considered to be de novo lesions, evidence has accumulated that at least some PTs may derive from pre-existing FAs [59, 60]. Clonal analysis of PTs and FAs that developed sequentially in the same patient revealed that not only were all lesions monoclonal, but that they also contained the same inactivated allele [61]. An exome sequencing analysis of paired longitudinally-acquired tumour samples from patients with initial FAs and subsequent PT recurrences by Tan et al. showed results suggestive of linear progression in two out of five cases [62].
Lae et al., in a CGH study of 30 PT cases, revealed recurrent chromosome imbalances in 55, 91 and 100% of benign, borderline and malignant PTs, respectively [63]. In that study, two distinct patterns of genomic imbalance in PTs were found: none or a few chromosomal changes corresponding to benign tumours, and numerous recurrent chromosomal changes, in particular 1q gain and 13q loss, that were seen in malignant ones. CGH findings segregating PTs into benign and malignant groups were also reported by Lv et al. [64] and Jones et al. [12]. In the latter study, areas of PTs that appeared histologically similar were found to be genomically heterogeneous on CGH analysis. mRNA expression profiling of 23 PTs found 162 genes to be upregulated in borderline and malignant PTs [65]. Ang et al. elucidated 29 genes that segregated 21 PTs into appropriate histologic grades using Affymetrix U133Plus 2.0 GeneChips [18]. Studies addressing the role of promoter methylation found methylation status to be associated with high-grade PT histology [12, 66, 67]. In a gene expression study of 75 breast fibroepithelial tumours by Vidal et al., the majority of FAs, benign and borderline PTs were identified as normal breast-like, while malignant PTs were more likely to be basal-like and claudin-low by intrinsic breast cancer subtyping [68].
Whole-genome profiling of 53 borderline and malignant PTs by Lae et al., using the Human CNV370 BeadChip microarray containing 370,000 SNP markers, identified a high number of copy number variations, including gains of 7p and 8q, losses of 13q (encompassing the RB1 gene) and 10p, losses in 9p21.3 and the presence of amplifications, especially involving EGFR, as markers of potentially malignant tumours [69]. Liu et al., in a genomic profiling study of ten malignant PTs with and without heterologous sarcomatous differentiation, found genomic aberrations in FGFR/EGFR PI-3 kinase and RAS pathways in 80% of cases, as well as TERT aberrations in 70% of the tumours [24]. Significant intratumoural heterogeneity with evidence of divergent evolution was revealed by copy number analysis in that study.
Highly recurrent somatic mutations in MED12 (Mediator complex subunit 12) were initially discovered in FAs [70]; these mutations were subsequently reported in both FAs and PTs by multiple groups [71,72,73,74,75], with reported frequencies of MED12 mutations in PTs ranging from 43 to 77% [62, 75, 76]. Piscuoglio et al. discovered identical MED12 mutations in multiple ipsilateral synchronous FELs (FAs, PTs) in the same patient, suggestive of clonal relatedness in different lesions [77]. Subsequently, Pareja et al. demonstrated that PTs with FA-like areas contained highly recurrent mutations in exon 2 of MED12, while PTs without FA-like areas were more likely to be MED12 wild type with more genetic alterations in cancer-related genes, such as EGFR [44]. These results suggested two distinct pathways of PT development: an FA-dependent, MED12-mutated pathway of progressive genetic alterations in cancer genes (such as TERT), and a MED12-independent pathway of de novo PT histogenesis following early cancer gene aberrations. This hypothesis corroborates the observation of improved disease-free survival in patients whose PTs harboured MED12 mutations as compared with those without [76].
Tan et al. found RARA mutations to be present in both FAs and PTs, while PTs exhibited mutations in FLNA, SETD2 and KMT2D, as well as additional mutations in cancer-associated genes including NF1, RB1, TP53, PIK3CA, ERBB4 and EGFR [62]. Tsang et al., in an analysis of TERT protein expression in 207 PTs, found high stromal TERT and positive margins to correlate with worse recurrence-free survival in PTs [78]. Nozad et al., who subjected 24 malignant PTs to comprehensive genomic profiling, found TP53, TERT, NF1, MED12, CDKN2A/B and MLL2 to be the most commonly mutated genes [79]. Deep sequencing performed on 17 PTs in a study by Kim et al. found MED12 and TERT to be the most common genetic alterations [43]. All 5 of 12 malignant PTs with local recurrence harboured RARA mutations, while 2 malignant PTs with lung metastases revealed similar BRCA2, SETD2 and TSC1 genetic changes.
In our study, gene mutations identified encompassed cell signalling, tumour suppressor, DNA repair and cell cycle regulating pathways. In general, mutations shared across multiple tumour regions showed higher variant allele frequency, which is consonant with their early role in tumourigenesis. Light microscopic morphological attributes of increasing cellular density and pleomorphism correlated with greater mutational burdens. A high prevalence of MED12 mutation, as well as recurrent mutations in TP53, RARA and PIK3CA, was noted. KMT2D, NRAS, PDGFRB and RB1 mutations were also identified. Phylogenetic analyses revealed likely disparate pathways of tumour progression.
An important limitation of this study is that we did not specifically analyze the stromal and epithelial components separately. It had been postulated that the epithelium in FELs, although histologically banal, is more than an innocuous bystander in lesional evolution. Epithelial–stromal interactions (ESI) between the ductular epithelium and adjacent fibrovascular stroma of the breast are required for normal mammary development, as well as development and progression of neoplasia. Immunohistochemical studies and molecular analysis comparing allelic imbalances highlight the importance of ESI in PTs [80,81,82]. ESIs may play a role in the acquisition of proliferative characteristics and malignant progression of tumour stroma. Dietrich et al. elucidated clonal abnormalities in both the epithelium and stroma of PTs [83], while Kuijper et al. also demonstrated monoclonality in PT epithelium [59]. These findings suggest the possible presence of a putative precursor cell with epithelial–stromal plasticity that may be the origin of diverse lesional components. Although laser microdissection of tumour epithelium was initially attempted in this study, the sequencing data generated was unsuitable for use, due to likely inadvertent contamination by tumour stroma.
Surgical excision is the current mainstay of therapy for PTs. Tumour recurrence remains an important clinical problem. Marked intratumoural genetic heterogeneity, which may not be readily apparent clinically or histologically, can contribute to under-recognition of subclones that may progress and recur. On the other hand, there likely also exists a group of PTs with benign biology that are overtreated [22]. The role of adjuvant therapy is currently unclear beyond that of palliation [84,85,86]. In a retrospective study of 36 malignant PTs profiled using gene sequencing, gene copy number analysis, whole-genome expression, and protein expression, there was consistent overexpression of genes involved in angiogenesis, including VEGFA, Angiopoietin-2, VCAM1, PDGFRA and PTTG1, in addition to EGFR, PIK3CA, TP53 and RB1 mutations [87]. In addition, three PTs exhibited above-threshold (>5%) PD-L1 expression, suggesting that further investigations into the roles of anti-angiogenic and immunomodulatory agents in the management of PTs may be warranted.
Our findings affirm the complex genetic makeup of breast FELs, with the burden of genetic aberrations appearing to parallel light microscopic features of malignancy. Larger studies to determine their prognostic and therapeutic implications may underpin risk-stratified management and guide novel directed therapies.
References
WHO Classification of Tumours Editorial Board. WHO classification of tumours of the breast. 5th ed. Lyon: IARC Press; 2019.
Guerrero MA, Ballard BR, Grau AM. Malignant phyllodes tumor of the breast: review of the literature and case report of stromal overgrowth. Surg Oncol. 2003;12:27–37.
Chua CL, Thomas A, Ng BK. Cystosarcoma phyllodes–Asian variations. Aust N Z J Surg. 1988;58:301–5.
Tan PH, Jayabaskar T, Chuah KL, Lee HY, Tan Y, Hilmy M, et al. Phyllodes tumors of the breast: the role of pathologic parameters. Am J Clin Pathol. 2005;123:529–40.
Tan BY, Acs G, Apple SK, Badve S, Bleiweiss IJ, Brogi E, et al. Phyllodes tumours of the breast: a consensus review. Histopathology. 2016;68:5–21.
Tan BY, Tan PH. A diagnostic approach to fibroepithelial breast lesions. Surg Pathol Clin. 2018;11:17–42.
Dessauvagie BF, Lee AHS, Meehan K, Nijhawan A, Tan PH, Thomas J, et al. Interobserver variation in the diagnosis of fibroepithelial lesions of the breast: a multicentre audit by digital pathology. J Clin Pathol. 2018;71:672–9.
Tan PH, Thike AA, Tan WJ, Thu MM, Busmanis I, Li H, et al. Predicting clinical behaviour of breast phyllodes tumours: a nomogram based on histological criteria and surgical margins. J Clin Pathol. 2012;65:69–76.
Nishimura R, Tan PH, Thike AA, Tan MH, Taira N, Li HH, et al. Utility of the Singapore nomogram for predicting recurrence-free survival in Japanese women with breast phyllodes tumours. J Clin Pathol. 2014;67:748–50.
Cristando C, Li HH, Almekinders M, Tan PH, Brogi E, Murray M. Validation of the Singapore nomogram for outcome prediction in a US-based population of women with breast phyllodes tumors (PT). Mod Pathol. 2017;30:36A.
Slodkowska E, Nofech-Mozes S, Xu B, Parra-Herran C, Lu FI, Raphael S, et al. Fibroepithelial lesions of the breast: a comprehensive morphological and outcome analysis of a large series. Mod Pathol. 2018;31:1073–84.
Jones AM, Mitter R, Springall R, Graham T, Winter E, Gillett C, et al. A comprehensive genetic profile of phyllodes tumours of the breast detects important mutations, intra-tumoral genetic heterogeneity and new genetic changes on recurrence. J Pathol. 2008;214:533–44.
Tsang JYS, Ni YB, Ng EK, Shin VY, Mak KF, Go EM, et al. MicroRNAs are differentially deregulated in mammary malignant phyllodes tumour. Histopathology. 2015;67:294–305.
Vilela MHT, de Almeida FM, de Paula GM, Ribeiro NB, Cirqueira MB, Silva AL, et al. Utility of Ki-67, CD10, CD34, p53, CD117, and mast cell content in the differential diagnosis of cellular fibroadenomas and in the classification of phyllodes tumors of the breast. Int J Surg Pathol. 2014;22:485–91.
Ho SK, Thike AA, Cheok PY, Tse GM, Tan PH. Phyllodes tumours of the breast: the role of CD34, vascular endothelial growth factor and β-catenin in histological grading and clinical outcome. Histopathology. 2013;63:393–406.
Kim GE, Kim JH, Lee KH, Choi YD, Lee JS, Lee JH, et al. Stromal matrix metalloproteinase-14 expression correlates with the grade and biological behavior of mammary phyllodes tumors. Appl Immunohistochem Mol Morphol. 2012;20:298–303.
Kwon JE, Jung WH, Koo JS. Molecules involved in epithelial-mesenchymal transition and epithelial-stromal interaction in phyllodes tumors: implications for histologic grade and prognosis. Tumour Biol. 2012;33:787–98.
Ang MK, Ooi AS, Thike AA, Tan P, Zhang Z, Dykema K, et al. Molecular classification of breast phyllodes tumors: validation of the histologic grading scheme and insights into malignant progression. Breast Cancer Res Treat. 2011;129:319–29.
Tsai WC, Jin JS, Yu JC, Sheu LF. CD10, actin, and vimentin expression in breast phyllodes tumors correlates with tumor grades of the WHO grading system. Int J Surg Pathol. 2006;14:127–31.
Chen CM, Chen CJ, Chang CL, Shyu JS, Hsieh HF, Harn HJ. CD34, CD117, and actin expression in phyllodes tumor of the breast. J Surg Res. 2000;94:84–91.
Tan WJ, Thike AA, Bay BH, Tan PH. Immunohistochemical expression of homeoproteins Six1 and Pax3 in breast phyllodes tumours correlates with histological grade and clinical outcome. Histopathology. 2014;64:807–17.
Karim RZ, O’Toole SA, Scolyer RA, Cooper CL, Chan B, Selinger C, et al. Recent insights into the molecular pathogenesis of mammary phyllodes tumours. J Clin Pathol. 2013;66:496–505.
McGranahan N, Swanton C. Biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell. 2015;27:15–26.
Liu SY, Joseph NM, Ravindranathan A, Stohr BA, Greenland NY, Vohra P, et al. Genomic profiling of malignant phyllodes tumors reveals aberrations in FGFR1 and PI-3 kinase/RAS signaling pathways and provides insights into intratumoral heterogeneity. Mod Pathol. 2016;29:1012–27.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26:589–95.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
García-Alcalde F, Okonechnikov K, Carbonell J, Cruz LM, Götz S, Tarazona S, et al. Qualimap: evaluating next-generation sequencing alignment data. Bioinformatics. 2012;28:2678–9.
Okonechnikov K, Conesa A, García-Alcalde F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics. 2016;32:292–4.
Fan Y, Xi L, Hughes DST, Zhang J, Zhang J, Futreal PA, et al. MuSE: accounting for tumor heterogeneity using a sample-specific error model improves sensitivity and specificity in mutation calling from sequencing data. Genome Biol. 2016;17:178.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31:213–9.
Fang H, Bergmann EA, Arora K, Vacic V, Zody MC, Iossifov I, et al. Indel variant analysis of short-read sequencing data with Scalpel. Nat Protoc. 2016;11:2529–48.
Saunders CT, Wong WSW, Swamy S, Becq J, Murray LJ, Cheetham RK. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics. 2012;28:1811–7.
Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009;25:2283–5.
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568–76.
Yang H, Wang K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc. 2015;10:1556–66.
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001;29:308–11.
Forbes SA, Beare D, Gunasekaran P, Leung K, Bindal N, Boutselakis H, et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 2015;43:D805–11.
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:D980–5.
Landrum MJ, Lee JM, Benson M, Brown G, Chao C, Chitipiralla S, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2016;44:D862–8.
Baum BR. PHYLIP: Phylogeny Inference Package. Version 3.2. Joel Felsenstein. Q Rev Biol. 1989;64:539–41.
Md Nasir ND, Ng CCY, Rajasegaran V, Wong SF, Liu W, Ng GX, et al. Genomic characterisation of breast fibroepithelial lesions in an international cohort. J Pathol. 2019;249:447–60.
Steller EJ, Raats DA, Koster J, Rutten B, Govaert KM, Emmink BL, et al. PDGFRB promotes liver metastasis formation of mesenchymal-like colorectal tumor cells. Neoplasia. 2013;15:204–17.
Kim JY, Yu JH, Nam SJ, Kim SW, Lee SK, Park WY, et al. Genetic and clinical characteristics of phyllodes tumors of the breast. Transl Oncol. 2018;11:18–23.
Pareja F, Geyer FC, Kumar R, Selenica P, Piscuoglio S, Ng CKY, et al. Phyllodes tumors with and without fibroadenoma-like areas display distinct genomic features and may evolve through distinct pathways. NPJ Breast Cancer. 2017;3:40.
Moon HG, Yun J, Hong BS, Lee E, Lee HB, Han W, et al. Molecular characterization of human malignant phyllodes tumors reveals potential targeted approaches [abstract]. Cancer Res. 2018;78:4.
Carvalho S, e Silva AO, Milanezi F, Ricardo S, Leitão D, Amendoeira I, et al. c-KIT and PDGFRA in breast phyllodes tumours: overexpression without mutations? J Clin Pathol. 2004;57:1075–9.
Jardim DL, Conley A, Subbiah V. Comprehensive characterization of malignant phyllodes tumor by whole genomic and proteomic analysis: biological implications for targeted therapy opportunities. Orphanet J Rare Dis. 2013;8:112.
Masjedi S, Zwiebel LJ, Giorgio TD. Olfactory receptor gene abundance in invasive breast carcinoma. Sci Rep. 2019;9:1–12.
Rodriguez M, Luo W, Weng J, Zeng L, Yi Z, Siwko S, et al. PSGR promotes prostatic intraepithelial neoplasia and prostate cancer xenograft growth through NF-κB. Oncogenesis. 2014;3:e114.
Li H, Batth IS, Qu X, Xu L, Song N, Wang R, et al. IGF-IR signaling in epithelial to mesenchymal transition and targeting IGF-IR therapy: overview and new insights. Mol Cancer. 2017;16:6.
Sin EI, Wong CY, Yong WS, Ong KW, Madhukumar P, Tan VK, et al. Breast carcinoma and phyllodes tumour: a case series. J Clin Pathol. 2016;69:364–9.
Widya RL, Rodrigues MF, Truong PT, Watson PH, Weir LM, Knowling MA, et al. Malignant epithelial transformation in phyllodes tumor: a population-based case series. Cureus. 2017;9:e1815.
Muller KE, Tafe LJ, de Abreu FB, Peterson JD, Wells WA, Barth RJ, et al. Benign phyllodes tumor of the breast recurring as a malignant phyllodes tumor and spindle cell metaplastic carcinoma. Hum Pathol. 2015;46:327–33.
Lu Z, Hunter T. Prolyl isomerase Pin1 in cancer. Cell Res. 2014;24:1033–49.
Zhou XZ, Lu KP. The isomerase PIN1 controls numerous cancer-driving pathways and is a unique drug target. Nat Rev Cancer. 2016;16:463–78.
Chen Y, Wu YR, Yang HY, Li XZ, Jie MM, Hu CJ, et al. Prolyl isomerase Pin1: a promoter of cancer and a target for therapy. Cell Death Dis. 2018;9:883.
Noguchi S, Motomura K, Inaji H, Imaoka S, Koyama H. Clonal analysis of fibroadenoma and phyllodes tumor of the breast. Cancer Res. 1993;53:4071–4.
Parker SJ, Harries SA. Phyllodes tumours. Postgrad Med J. 2001;77:428–35.
Kuijper A, Buerger H, Simon R, Schaefer KL, Croonen A, Boecker W, et al. Analysis of the progression of fibroepithelial tumours of the breast by PCR-based clonality assay. J Pathol. 2002;197:575–81.
Hodges KB, Abdul-Karim FW, Wang M, Lopez-Beltran A, Montironi R, Easley S, et al. Evidence for transformation of fibroadenoma of the breast to malignant phyllodes tumor. Appl Immunohistochem Mol Morphol. 2009;17:345–50.
Noguchi S, Yokouchi H, Aihara T, Motomura K, Inaji H, Imaoka S, et al. Progression of fibroadenoma to phyllodes tumor demonstrated by clonal analysis. Cancer. 1995;76:1779–85.
Tan J, Ong CK, Lim WK, Ng CC, Thike AA, Ng LM, et al. Genomic landscapes of breast fibroepithelial tumors. Nat Genet. 2015;47:1341–5.
Laé M, Vincent-Salomon A, Savignoni A, Huon I, Fréneaux P, Sigal-Zafrani B, et al. Phyllodes tumors of the breast segregate in two groups according to genetic criteria. Mod Pathol. 2007;20:435–44.
Lv S, Niu Y, Wei L, Liu Q, Wang X, Chen Y. Chromosomal aberrations and genetic relations in benign, borderline and malignant phyllodes tumors of the breast: a comparative genomic hybridization study. Breast Cancer Res Treat. 2008;112:411–8.
Jones AM, Mitter R, Poulsom R, Gillett C, Hanby AM, Tomlinson IP, et al. mRNA expression profiling of phyllodes tumours of the breast: identification of genes important in the development of borderline and malignant phyllodes tumours. J Pathol. 2008;216:408–17.
Huang KT, Dobrovic A, Yan M, Karim RZ, Lee CS, Lakhani SR, et al. DNA methylation profiling of phyllodes and fibroadenoma tumours of the breast. Breast Cancer Res Treat. 2010;124:555–65.
Kim JH, Choi YD, Lee JS, Lee JH, Nam JH, Choi C, et al. Borderline and malignant phyllodes tumors display similar promoter methylation profiles. Virchows Arch. 2009;455:469–75.
Vidal M, Peg V, Galván P, Tres A, Cortés J, Ramón Y, et al. Gene expression-based classifications of fibroadenomas and phyllodes tumours of the breast. Mol Oncol. 2015;9:1081–90.
Laé M, La Rosa P, Mandel J, Reyal F, Hupé P, Terrier P, et al. Whole-genome profiling helps to classify phyllodes tumours of the breast. J Clin Pathol. 2016;69:1081–7.
Lim WK, Ong CK, Tan J, Thike AA, Ng CC, Rajasegaran V, et al. Exome sequencing identifies highly recurrent MED12 somatic mutations in breast fibroadenoma. Nat Genet. 2014;46:877–80.
Cani AK, Hovelson DH, McDaniel AS, Sadis S, Haller MJ, Yadati V, et al. Next-gen sequencing exposes frequent MED12 mutations and actionable therapeutic targets in phyllodes tumors. Mol Cancer Res. 2015;13:613–9.
Nagasawa S, Maeda I, Fukuda T, Wu W, Hayami R, Kojima Y, et al. MED12 exon 2 mutations in phyllodes tumors of the breast. Cancer Med. 2015;4:1117–21.
Pfarr N, Kriegsmann M, Sinn P, Klauschen F, Endris V, Herpel E, et al. Distribution of MED12 mutations in fibroadenomas and phyllodes tumors of the breast-implications for tumor biology and pathological diagnosis. Genes Chromosomes Cancer. 2015;54:444–52.
Piscuoglio S, Murray M, Fusco N, Marchiò C, Loo FL, Martelotto LG, et al. MED12 somatic mutations in fibroadenomas and phyllodes tumours of the breast. Histopathology. 2015;67:719–29.
Yoshida M, Sekine S, Ogawa R, Yoshida H, Maeshima A1, Kanai Y, et al. Frequent MED12 mutations in phyllodes tumours of the breast. Br J Cancer. 2015;112:1703–8.
Ng CCY, Tan J, Ong CK, Rajasegaran V, Nasir ND, Lim JC, et al. MED12 is frequently mutated in breast phyllodes tumours: a study of 112 cases. J Clin Pathol. 2015;68:685–91.
Piscuoglio S, Ng CK, Murray M, Burke KA, Edelweiss M, Geyer FC, et al. Massively parallel sequencing of phyllodes tumours of the breast reveals actionable mutations, and TERT promoter hotspot mutations and TERT gene amplification as likely drivers of progression. J Pathol. 2016;238:508–18.
Tsang JYS, Hui YK, Lee MA, Lacambra M, Ni YB, Cheung SY, et al. Association of clinicopathological features and prognosis of TERT alterations in phyllodes tumor of breast. Sci Rep. 2018;8:3881.
Nozad S, Sheehan CE, Gay LM, Elvin JA, Vergilio JA, Suh J, et al. Comprehensive genomic profiling of malignant phyllodes tumors of the breast. Breast Cancer Res Treat. 2017;162:597–602.
Sawyer EJ, Hanby AM, Ellis P, Lakhani SR, Ellis IO, Boyle S, et al. Molecular analysis of phyllodes tumors reveals distinct changes in the epithelial and stromal components. Am J Pathol. 2000;156:1093–8.
Dacic S, Kounelis S, Kouri E, Jones MW. Immunohistochemical profile of cystosarcoma phyllodes of the breast: a study of 23 cases. Breast J. 2002;8:376–81.
Tan PH, Jayabaskar T, Yip G, Tan Y, Hilmy M, Selvarajan S, et al. p53 and c-kit (CD117) protein expression as prognostic indicators in breast phyllodes tumors: a tissue microarray study. Mod Pathol. 2005;18:1527–34.
Dietrich CU, Pandis N, Rizou H, Petersson C, Bardi G, Qvist H, et al. Cytogenetic findings in phyllodes tumors of the breast: karyotypic complexity differentiates between malignant and benign tumors. Hum Pathol. 1997;28:1379–82.
Chaney AW, Pollack A, McNeese MD, Zagars GK, Pisters PW, Pollock RE, et al. Primary treatment of cystosarcoma phyllodes of the breast. Cancer. 2000;89:1502–11.
Khosravi-Shahi P. Management of non metastatic phyllodes tumors of the breast: review of the literature. Surg Oncol. 2011;20:e143–8.
Strode M, Khoury T, Mangieri C, Takabe K. Update on the diagnosis and management of malignant phyllodes tumors of the breast. Breast. 2017;33:91–6.
Gatalica Z, Vranic S, Ghazalpour A, Xiu J, Ocal IT, McGill J, et al. Multiplatform molecular profiling identifies potentially targetable biomarkers in malignant phyllodes tumors of the breast. Oncotarget. 2016;7:1707–16.
Acknowledgements
This study was funded by the National Cancer Centre Research Fund and Oncology Academic Clinical Program (NCCRF-OACPCCS-YR2015-AUG-4). It was presented in part at the 2017 United States and Canadian Academy of Pathology Annual Scientific Meeting held in San Antonio, Texas. We thank our surgical colleagues who contributed the cases.
Author information
Authors and Affiliations
Contributions
Study conceptualization, design and project supervision: PHT, BTT and BYT. Performed experiments: NDMN, CCYN, VR, PG, SN, JYL and JQL. Data collection, analysis and interpretation: BYT, NDMN, HYC, CCYN, VR, PG, SN, JYL, JQL and AAT. Writing of paper: BYT, NDMN, HYC, CCYN and PG. Revised and provided inputs to paper: all authors.
Corresponding authors
Ethics declarations
Conflict of interest
BTT and PHT jointly hold patent applications for PCT/SG2015/050107 (Breast fibroadenoma susceptibility mutations and use thereof) and PCT/SG2015/050368 (Method and kit for pathologic grading of breast neoplasms). Other authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Tan, B.Y., Md Nasir, N.D., Chang, H.Y. et al. Morphologic and genetic heterogeneity in breast fibroepithelial lesions—a comprehensive mapping study. Mod Pathol 33, 1732–1745 (2020). https://doi.org/10.1038/s41379-020-0533-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-020-0533-0
This article is cited by
-
Artificial intelligence modelling in differentiating core biopsies of fibroadenoma from phyllodes tumor
Laboratory Investigation (2022)
-
Fibroepithelial lesions revisited: implications for diagnosis and management
Modern Pathology (2021)
-
Genetic differences between benign phyllodes tumors and fibroadenomas revealed through targeted next generation sequencing
Modern Pathology (2021)