Abstract
The involvement of Wnt signaling in human lung cancer remains unclear. This study investigated the role of Wnt11 in neuroendocrine (NE) differentiation, cell proliferation, and epithelial-to-mesenchymal transition (EMT) in human small-cell lung cancer (SCLC). Immunohistochemical staining of resected specimens showed that Wnt11 was expressed at higher levels in SCLCs than in non-SCLCs; 58.8% of SCLC, 5.2% of adenocarcinoma (ADC), and 23.5% of squamous cell carcinoma tissues stained positive for Wnt11. A positive relationship was observed between Achaete-scute complex homolog 1 (Ascl1) and Wnt11 expression in SCLC cell lines, and this was supported by transcriptome data from SCLC tissue. The expression of Wnt11 and some NE markers increased after the transfection of ASCL1 into the A549 ADC cell line. Knockdown of Ascl1 downregulated Wnt11 expression in SCLC cell lines. Ascl1 regulated Wnt11 expression via lysine H3K27 acetylation at the enhancer region of the WNT11 gene. Wnt11 controlled NE differentiation, cell proliferation, and E-cadherin expression under the regulation of Ascl1 in SCLC cell lines. The phosphorylation of AKT and p38 mitogen-activated protein kinase markedly increased after transfection of WNT11 into the SBC3 SCLC cell line, which suggests that Wnt11 promotes cell proliferation in SCLC cell lines. Ascl1 plays an important role in regulating the Wnt signaling pathway and is one of the driver molecules of Wnt11 in human SCLC. Ascl1 and Wnt11 may employ a cooperative mechanism to control the biology of SCLC. The present results indicate the therapeutic potential of targeting the Ascl1–Wnt11 signaling axis and support the clinical utility of Wnt11 as a biological marker in SCLC.
Similar content being viewed by others
Introduction
Lung cancer is a leading cause of cancer-related mortality worldwide. Small-cell lung cancer (SCLC) accounts for 10–15% of all lung cancer cases and is genetically considered to be one of the most aggressive malignant neuroendocrine (NE) tumors. Despite a high response rate to first-line treatment, most cases show rapid growth and metastasis and acquire multidrug resistance, resulting in a 5-year survival rate of 5–10% [1]. In recent years, the clinical focus has been on personalized therapy, as represented by the use of epidermal growth factor tyrosine kinase inhibitors and anaplastic lymphoma kinase inhibitors to treat non-small-cell lung cancer (NSCLC) [2, 3]. Furthermore, personalized therapies have been employed for lung adenocarcinoma (ADC) patients with rare mutant oncogenes, such as ROS1 and RET fusion genes [4, 5]. In contrast, novel target molecules of therapies for SCLC are yet to be developed. The molecular mechanisms of small-cell carcinogenesis have not yet been elucidated, and thus, advances in research on novel therapeutic modalities are expected [6].
A characteristic feature of SCLC is the expression of NE phenotypes, which is considered to correlate with tumor malignancy, including rapid growth, a high metastatic rate at an early stage, and the acquisition of multidrug resistance [7,8,9]. Achaete-scute complex 1 (ASCL1), a proneural basic helix–loop–helix factor, is strongly expressed in SCLC [10]. Osada et al. reported that the knockdown of Ascl1 induced growth inhibition and apoptosis in SCLC cell lines [11, 12]. Borromeo et al. showed that Ascl1 played a more pivotal role in tumorigenesis than Neurod1 in human SCLC cell lines and mouse models of SCLC [13]. Our previous findings suggest that SCLC exhibited an epithelial-to-mesenchymal transition (EMT)-like phenotype, which may be related to inactivated Notch signaling [14]. Notch signaling is an important cell signaling system, and the interaction between Notch receptors and ligands induces several genes, such as hairy and enhancer of split-1 (HES1), CCND1, MYC, and AKT [15]. The Notch1–Hes1 pathway represses NE differentiation by decreasing the expression of NE-promoting transcription factors, including ASCL1 and insulinoma-associated protein 1 (INSM1) [16,17,18,19,20]. Among lung cancers, SCLC was shown to be positive for Ascl1 and/or Insm1, but negative for Notch1, whereas NSCLC was negative for Ascl1 and/or Insm1, but positive for Notch1 [19].
Wnt signaling regulates a number of developmental processes, including cell fate specification, proliferation, polarity, and migration [21]. Wnt molecules trigger gene transcription via at least three signaling pathways: the canonical or β-catenin-dependent and two noncanonical pathways, comprising the planar cell polarity (PCP) and Wnt/Ca2+ pathways. Increasing evidence has shown that the Wnt pathway is one of the main signaling pathways involved in carcinogenesis in many organs. The aberrant activation of this pathway underlies several debilitating diseases, and lung cancer is not an exception [21, 22].
Most of the 19 Wnt ligands and 10 receptors, frizzled (FZD) proteins, which have been identified in mammalian cells, may also be present in human lung cells [23]. Mammalian Wnt11 is a 354-amino acid protein with a predicted 24-amino acid signal peptide, and 22 core cysteine residues that are found in all the Wnt family members, two predicted sites for lipid modification (palmitate at C80 and palmitoleic acid at S215), and five predicted sites for Asn-linked glycosylation [24]. Wnt11 is known for its role during development; it is required for convergent extension movements during gastrulation and kidney morphogenesis [25]. Wnt11 has also been implicated in the carcinogenesis of some organs, such as the stomach [26], kidneys [26], colon [27], and mammary gland [28]. Uysal-Onganer et al. reported that Wnt11 promoted NE differentiation, survival, and migration in prostate cancer [29]. In NSCLC, Bartis et al. identified Wnt11 as a regulator of cadherin expression and demonstrated its role in cellular adhesion [30].
We hypothesized that Wnt11 may contribute to the biology of SCLC. To test this hypothesis, this study investigated the following: (1) the frequency of Wnt11 expression in resected lung cancer tumors and cell lines, (2) the relationship between Ascl1 and Wnt11 in SCLC, and (3) the molecular and biological functions of Wnt11 in human SCLC. The results obtained showed that Wnt11 was more strongly expressed in SCLC than in NSCLC, and also that it regulated NE differentiation, cell proliferation, and E-cadherin expression in an Ascl1-dependent manner in SCLC.
Materials and methods
Cell lines
Seven SCLC cell lines (H69, H889, SBC1, H69AR, H1688, SBC3, and SBC5), three ADC cell lines (A549, H358, and H1975), and two SCC cell lines (H2170 and H226) were used in this study. H69, H889, H69AR, H1688, A549, H358, H1975, H2170, and H226 were purchased from ATCC (Manassas, VA), and SBC1, SBC3, and SBC5 from the Japan Collection of Research Bioresources Cell Bank (Osaka, Japan).
Tissue samples
Tissue samples of SCLCs (n = 17), ADCs (n = 19), and SCCs (n = 17) resected at the Department of Thoracic Surgery of Kumamoto University Hospital were obtained from 53 patients for experiments. A histological diagnosis of samples was made according to the criteria of the World Health Organization [31]. These sections were used for immunohistochemical (IHC) staining. This study followed the guidelines of the Ethics Committee of Kumamoto University.
Western blot (WB) analysis
Cells were prepared for a WB analysis as described previously [32]. A list of the primary antibodies used is shown in Table 1. Membranes were washed and incubated with the respective secondary antibodies conjugated with peroxidase (Amersham Pharmacia Biotech, Buckinghamshire, UK) for 1 h, and the immunocomplex was visualized using a chemiluminescence substrate (Amersham Pharmacia Biotech, UK).
Transcriptome data of SCLC tissues
The transcriptome data of SCLC tissue samples from the GSE60052 (n = 79) RNA-seq data set [33] were used to examine the relationship between the expression levels of ASCL1 and WNT11.
RNA-sequence (RNA-seq) analysis
An RNA-seq analysis was performed by the Liaison Laboratory Research Promotion Center (LILA) (Kumamoto University), as follows. Total RNA was isolated using an RNeasy Mini Kit (Qiagen, Germany). A 2100 Bioanalyzer (Agilent, Santa Clara, CA) was employed to detect the concentration and purity of total RNA. All samples with an RNA integrity number >7.5 were used for sequencing. NextSeq 500 (Illumina, San Diego, CA) was used for the analysis, and the data were converted into Fastq files. The quality control of data was performed using FastQC. Filtered reads were then used to map to the UCSC hg19 genome reference genome using HISAT2 2.1.0. Fragments per kilobase of exon per million reads mapped (FPKM) values were calculated using Cufflinks. Significant genes were extracted by cuffdiff (p < 0.05).
Transfection with siRNA
We purchased Wnt11 siRNA (sc-41120) and Ascl1 siRNA (sc-37692) from Santa Cruz Biotechnology (Santa Cruz, CA) and transfected them into cells at a concentration of 20 nmol/L using an electroporator (NEPA21 pulse generator; Nepa Gene, Chiba, Japan), as described in the manufacturer’s instructions. These were pools of three different siRNA duplexes, and the sequences for Wnt11 were as follows: sense; 5′-CUCUGGAAAUGAAGUGUAATT-3′, antisense; 5′-UUACACUUCAUUUCCAGAGTT-3′ (sc-41120A), sense; 5′-CCUGACUUCUGCAUGAAGATT-3′, antisense; 5′-UCUUCAUGCAGAAGUCAGGTT-3′ (sc-41120B), and sense; 5′-GGAUGUGGAAGCUACAGAATT-3′, antisense; 5′-UUCUGUAGCUUCCACAUCCTT-3′ (sc-41120C). The sequences for Ascl1 were as follows: sense; 5′-CCAACAAGAAGAUGAGUAATT-3′, antisense; 5′-UUACUCAUCUUCUUGUUGGTT-3′ (sc-37692A), sense; 5′-GAAGCGCUCAGAACAGUAUTT-3′, antisense; 5′-AUACUGUUCUGAGCGCUUCTT-3′ (sc-37692B), and sense; 5′-GUUCGGGGAUAUAUUAAGATT-3′, antisense; 5′-UCUUAAUAUAUCCCCGAACTT-3′. Control siRNA (Cat# sc-36869) was used as a control. Cells were harvested 48 h post transfection.
Plasmid construction and transfection
To construct pCMV-AC-GFP-WNT11, we replicated the WNT11 gene of a human Wnt11 cDNA ORF clone (OriGene, USA) and replaced it with WNT11. We generated pCMV6-acGFP-mock from a human WNT11 cDNA ORF clone by cleaving out WNT11. These plasmids were transfected into SBC3 and H1975 cells using Lipofectamine 3000 (Invitrogen) as described in the manufacturer’s instructions. After 48 h, 1 mg/mL of Neomycin (G418, Invitrogen) was added to the cells for 2 weeks, with the medium being exchanged every 3 days. Stably transfected resistant cell lines were cloned from the transfectants. Regarding ASCL1-transfected A549 cells, we used a pCAG-IRES-puro-FlagHA vector (a generous gift from Prof. H. Niwa; Kumamoto University) to establish stable lines.
Cell-counting assay
After 48 h of siRNA and control transfection, cells were stained with trypan blue and counted. When H69 and H889 cell lines were used, 2 × 105 cells were seeded on six-well plates. Every 2 days, cells were collected and counted, after which they were seeded into fresh medium and left at 37 °C in 5% CO2. The counting method was continued until day 6. The criteria for cellular integrity included trypan blue exclusion, an intact nucleus, and an intact cell membrane. Experiments were repeated three times separately to confirm reproducibility. Stably transfected resistant SBC3 and H1975 cell lines with pCMV-AC-GFP-WNT11 and the control vector were also compared in the same manner, but only until day 3.
IHC staining and evaluation
Formalin-fixed, paraffin-embedded specimens were cut into 4-μm-thick sections and mounted onto MAS-GP-coated slides (Matsunami Glass Ind, Osaka, Japan). After being deparaffinized and rehydrated, sections were heated using an autoclave in 0.01 mol/L citrate buffer (pH 7.0) for antigen retrieval. Sections were incubated with 0.3% H2O2 in absolute methanol for 30 min to block endogenous peroxidase activity. Sections were then incubated with skimmed milk for 30 min to block nonspecific staining. After this blocking step, sections were incubated with the primary antibodies shown in Table 1 at 4 °C overnight. This was followed by sequential 1-h incubations with the secondary antibodies (En Vision + System-HRP-Labeled Polymer, Dako) and visualization with the Liquid DAB + Substrate Chromogen System (Dako). All slides were counterstained with hematoxylin for 30 s before being dehydrated and mounted. We evaluated the IHC results based on staining intensity and the percentage of positively stained tumor cells. The percentage of positively stained tumor cells was divided into four groups: with no staining, <5% tumor cells reactive, 5–25% reactive, and >50% reactive. Staining intensity levels were divided into three groups: negative, weak, and strong. We designed a table to allocate IHC scores to each specimen. IHC scores were classified into three groups: negative (0), weak positive (1+), and positive (2+). We defined a 2+ IHC score as significantly positive. Scoring was performed simultaneously by two independent observers who were blinded to patient details.
Tumor xeno-transplantation experiment
Rag(−/−):Jak3(−/−), 8-week-old male mice (generous gift from Prof. S. Okada, Kumamoto University) were used. Two groups of mice were injected subcutaneously: one group was injected with 2 × 106 stably transfected cells with ASCL1, and the other group was injected with an equal number of the control cell population. After 4 weeks, subcutaneous tumors were removed and fixed. Samples were fixed with 10% formalin and embedded in paraffin. Tissue sets were stained with hematoxylin and eosin, and additional sections were used for IHC staining. All animal work was performed in accordance with the Institutional Animal Care and Use Committee guidelines.
Reverse transcription-polymerase chain reaction (RT-PCR)
Total RNA was isolated using an RNeasy Mini Kit (Qiagen, Germany) in accordance with the manufacturer’s instructions. cDNA was produced using a ReverTra Ace qPCR RT-kit (Toyobo, Osaka, Japan), according to the manufacturer’s instructions. A list of the primers used is shown in Table 2. Amplified DNA fragments were separated on a 1.5% agarose gel and identified after ethidium bromide staining.
Quantitative real-time PCR (qRT-PCR)
RNA isolation and cDNA synthesis were performed as described above. cDNA was subjected to quantitative SYBR Green real-time PCR using SYBR Premix Ex Taq II (Takara Bio). A list of the specific primers used is shown in Table 2. qRT-PCR was performed with LightCycler® Nano (Roche Diagnostic K.K.), using 40 cycles of a three-stage program under the following conditions: at 94 °C for 20 s, at 60 °C for 40 s, and at 72 °C for 15 s, as recommended by the manufacturer. The products were quantified during the initial exponential phase of amplification above the baseline. Data were obtained from triplicate reactions. The means and SDs of the copy number were normalized to the value for glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNA.
Chromatin immunoprecipitation (ChIP)–qPCR assay for H3K27Ac at the enhancer region of the WNT11 gene
H889, SBC1, ASCL1-transfected A549 cells (clones 1 and 2), and A549 mock cells were cross-linked with 1% formaldehyde at 37 °C for 5 min, washed with PBS containing 2% FBS, lysed with ChIP buffer (10 mM Tris-HCl, pH 8.0, 200 mM NaCl, 1 mM CaCl2, 0.5% NP-40, and protease inhibitor cocktail), and sonicated for 10 s × 3 times (50 s on ice) using a NR-50M (MicroTech Nition, Chiba, Japan). Cells were digested with 200 U/mL micrococcal nuclease (MNase) (NEB) at 37 °C for 40 min and treated with 10 mM EDTA to stop the reaction. After the addition of an equal volume of RIPA buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 2 mM EDTA, 1% NP-40, 0.5% sodium deoxycholate, and 0.1% SDS), cells were resonicated for 10 s × 10 times (50 s on ice) using NR-50M. After centrifugation, the soluble chromatin fraction was immunoprecipitated at 4 °C overnight with Dynabeads Protein A/G with an anti-H3K27Ac antibody (Active Motif; MABI 0309, San Diego, CA). A normal mouse IgG antibody (Santa Cruz Biotechnology; sc-2025) was used as the control. Immunoprecipitates were washed with high-salt ChIP buffer (500 mM NaCl) four times and TE buffer (10 mM Tris-HCl, pH 8.0 and 1 mM EDTA) twice. Bound chromatin and 25 μL of input DNA were suspended in 47.5 and 22.5 μL of elution buffer (50 mM Tris-HCl, pH 8.0, 10 mM EDTA, 1% SDS, and 250 mM NaCl), respectively, and vortexed. After the addition of 5 μL of 5 M NaCl, the solutions were incubated at 65 °C for 4 h and then treated with 25 μg/mL RNase A (Sigma-Aldrich) at 37 °C for 30 min and 0.1 mg/mL proteinase K (Roche) at 50 °C for 1 h to reverse formaldehyde cross-linking. Eluted DNA was purified with a Wizard® SV Gel and PCR Clean-Up system (Promega). In ChIP assays, quantitative real-time PCR was performed on input and ChIP DNA samples. The primer sequences used were as follows: gene: forward/reverse primer (5′ to 3′).
WNT11: AATGCTCAGCACGTAATCGG/CGGCCACCTGACAATTCCTC, and control: CTTTGTGCCAGGTCACCTCCTTC/ AGGGAAGCCCTAATGTCAGGGTG.
Cell invasion assay for WNT11-transfected SBC3 cells
Cell invasion assay activity was measured with the Corning® BioCoatTM Matrigel® Invasion Chamber (#354480, Corning, New York) according to the manufacturer’s protocol. In brief, WNT11-transfected SBC3 or mock cells were inoculated on the coated Matrigel of the culture inserts (upper chamber) at a density of 5 × 104 cells per 500 µL of serum-free medium. The lower chamber contained the usual serum-containing medium as a chemoattractant. At the end of the 22-h incubation at 37 °C and 5% CO2 atmosphere, cells on the upper surface of the filter were completely removed with cotton swabs. The invaded cells that remained on the lower surface of the filter were stained with Diff-Quick (Sysmex, Hyogo, Japan). The numbers of stained cells in five randomly selected microscopic fields (×100) per filter were counted and the average number was calculated. Experiments were repeated three separate times.
Canonical Wnt pathway inhibitory experiment
We purchased IWR-1-endo (CAS 1127442-82-3) from Cayman Chemicals (Ann Arbor, MI). At 10 µM, IWR-1-endo inhibits the Wnt-induced accumulation of β-catenin, leading to the proteasomal degradation of this protein through a destruction complex, which consists of Apc, Axin2, Ck1, and GSK3β. IWR-1-endo stabilizes the destruction complex and blocks a cell-based Wnt/β-catenin pathway reporter response [34]. We treated the H69 and H889 cell lines in accordance with the manufacturer’s instructions. In brief, IWR-1-endo was dissolved in DMSO at 10 µM. We seeded cells at 5 × 105 cells/well on a six-well plate. We supplied IWR-1-endo or DMSO alone, and after 24 h at 37 °C in 5% CO2, collected the cells.
Statistical analysis
Spearman’s correlation coefficient (ρ) was calculated for the correlation analysis. Cell-counting data were expressed as the means ± standard deviation of triplicate measurements. Differences in mean values between the groups were analyzed by a two-tailed statistical analysis using the Student’s t-test. GraphPad Prism version 7.04 (San Diego, CA) was used for the statistical analysis. p values <0.05 were considered to be significant.
Results
Wnt11 is overexpressed in SCLC tissues and cell lines and positively correlates with Ascl1 expression
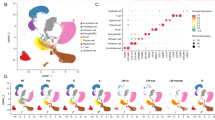
To the best of our knowledge, the expression patterns of Ascl1 and Wnt11 have not yet been examined in lung cancer. Therefore, in order to investigate Ascl1 and Wnt11 expression patterns, we stained surgically resected lung cancer tissues (17 SCLCs, 19 ADCs, and 17 squamous cell carcinomas (SCCs)) for Ascl1 and Wnt11 using IHC (Table 3 and Fig. 1a), and performed a WB analysis on 12 lung cancer cell lines (seven SCLCs, three ADCs, and two SCCs) (Fig. 1b). IHC revealed that ASCL1 and WNT11 were more strongly expressed in SCLC specimens than in NSCLC specimens. ASCL1 staining was detected in the nuclei of SCLC cells, and WNT11 staining was observed in the cytoplasmic and perinuclear areas of SCLC cells. While the majority of ASCL1-positive cases showed positive staining for WNT11 (seven out of eight cases), one ASCL1-positive, but WNT11-negative case was also observed (one out of eight cases) in SCLC. Furthermore, the WB analysis revealed that WNT11 was overexpressed in four out of the seven SCLC cell lines examined. The expression of ASCL1 and WNT7B in these lung cancer cell lines was also confirmed by the WB analysis. Three out of the seven SCLC cell lines simultaneously expressed ASCL1 and WNT11. SBC1 is an exceptional SCLC cell line that expressed ASCL1, but only a low level of WNT11. In contrast to WNT11, WNT7B was frequently overexpressed in the NSCLC cell lines examined. The expression patterns of the NE markers chromogranin A (CHGA), synaptophysin (SYP), and neural cell adhesion molecule (NCAM) were also shown. The similar expression patterns of ASCL1, WNT11, and WNT7B mRNAs were confirmed by RT-PCR in these human lung cancer cell lines (Fig. 1c). Furthermore, based on the fact that Ascl1 and Wnt11 were more likely to be co-expressed in SCLC, we surveyed public data sets of gene expression profiles in human SCLC samples and evaluated their relationships. The RNA-seq data set using the tumor samples of 79 SCLC patients confirmed the coordinated expression of ASCL1 and WNT11 in human SCLC tissue samples (GSE60052: ρ = 0.5283) (Fig. 1d) [33]. This result indicates that Wnt11 expression may be positively regulated by Ascl1.
a ASCL1 was positively immunostained in SCLC cell nuclei. Immunohistochemical staining showed that WNT11 was more strongly expressed in SCLC specimens than in non-SCLC specimens. Representative images are shown. Scale bar = 50 μm. b A WB analysis was conducted using lung cancer cell lines, including SCLC, ADC, and SCC. The expression of ASCL1, WNT11, WNT7B, CHGA, SYP, and NCAM in lung cancer cell lines. Three out of seven SCLC cell lines simultaneously expressed WNT11 and ASCL1. β-actin served as an internal control. c Expression patterns of ASCL1, WNT11, and WNT7B mRNA were shown by RT-PCR in 12 human lung cancer cell lines and were similar to protein expression patterns in the WB analysis. GAPDH served as an internal control. d Expression levels of ASCL1 and WNT11 in the RNA-seq data set of SCLC tissues. The GSE60052 (n = 79) data set [33] was analyzed
Ascl1 is a driver molecule of Wnt11 in SCLC
To investigate the biological effects of Ascl1 on Wnt11 in lung cancer cell lines, we performed ASCL1 transfection experiments. An A549 cell line stably transfected with ASCL1 was established; A549 is a representative ADC cell line that does not express Ascl1 or other NE molecules. The expression changes of the NE molecules, E-CADHERIN, SLUG, and SNAIL was induced by the forced expression of ASCL1, as described previously [14]. The results obtained showed that WNT11 was also induced in ASCL1-transfected ADC cells (Fig. 2a). IHC staining revealed that WNT11 was more strongly expressed after the transfection of ASCL1 in the xeno-transplanted tumors of immunocompromised mice (Fig. 2b). We used these cells in the RNA-seq analysis and compared mRNA expression by RT-PCR or heatmaps. We detected ten Wnt signaling-related molecules, including WNT11, that were significantly changed after the transfection with ASCL1. Wnt1-inducible signaling pathway protein 2 (WISP2), dickkopf-related protein (DKK) 1, secreted frizzled related protein 1 (SFRP1), and WNT7B expression levels decreased, whereas WNT11, WNT10A, DKK2, lymphoid enhancer binding factor 1 (LEF1), disheveled binding antagonist of beta catenin 1 (DACT1), and receptor tyrosine kinase like orphan receptor 2 (ROR2) expression levels increased (Fig. 2c, d). These results suggest that Ascl1 is strongly involved in the Wnt signaling pathway.
a) The WB analysis showed that the transfection of ASCL1 decreased the expression of E-CADHERIN and increased that of EMT-related transcription factors. In addition, we demonstrated that WNT11 was also induced in these cells. In contrast, WNT7B and DKK1 decreased. TF, transfected. b Xeno-transplantation of ASCL1-transfected A549 cells. Using IHC, ASCL1 transfection was shown to induce WNT11 protein expression. Representative images are shown. Scale bar = 50 μm. c RNA-seq data are shown using a heatmap. We extracted ten molecules related to the Wnt signaling pathway. d RT-PCR data are shown. We confirmed that these results corresponded with RNA-seq data. GAPDH served as an internal control. e, f Ascl1 affects Wnt11 expression in the SCLC cell lines H69, H889, and SBC1. The suppression of Wnt11 by siRNA for ASCL1 was confirmed in H69 and H889 cells by the WB analysis (e), and SBC1 cells by the qRT-PCR analysis (f). Data are shown as means ± SD. Asterisks indicate a significant difference. *p < 0.05. NT nontreated, si small interfering
To confirm whether Ascl1 affects Wnt11 expression in SCLC cell lines, we conducted an ASCL1 knockdown experiment using RNAi interference in H69, H889, and SBC1 cells. The knockdown of ASCL1 expression in H69 and H889 cells resulted in a significant reduction in WNT11 (Fig. 2e). qRT-PCR revealed that the knockdown of ASCL1 mRNA expression in SBC1 also led to a significant reduction in WNT11 (Fig. 2f). These results support Ascl1 functioning as a driver molecule for Wnt11 expression in SCLC.
Ascl1 regulates Wnt11 expression by altering lysine H3K27 acetylation (H3K27Ac)
To clarify the mechanisms by which Ascl1 regulates Wnt11 expression in SCLC, we conducted ChIP assays for H3K27Ac in H889, SBC1, ASCL1-transfected A549 cells (clones 1 and 2), and A549 mock cell lines. H3K27Ac is associated with the stronger activation of transcription, and thus, is defined as an active enhancer marker. H3K27Ac-marked chromatin, called an enhancer, is associated with key oncogenic genes in cancer cells [35]. Borromeo et al. identified WNT11 as one of the target genes of ASCL1 based on their ChIP sequencing (ChIP-seq) findings from SCLC cell lines, which included H889 cells [13]. An integrated genome viewer snapshot of H889, H128, and H2107 cells for ASCL1 was shown (Fig. 3a). We characterized H3K27Ac levels over the 10-kb upstream region from the WNT11 gene (hg19, chr11: 75,928,700–75,928,930 bp), which was detected as the highest peak examined. In H889 cells, H3K27Ac levels significantly increased in the targeted chromosome site by ChIP-qPCR assay. In contrast, we did not observe a significant change in H3K27Ac levels at the same location in SBC1 cells (Fig. 3b). Furthermore, we detected different H3K27Ac levels between A549 mock and ASCL1-transfected A549 clones (Fig. 3c). As described above, H889 cells strongly expressed Ascl1 and Wnt11, whereas SBC1 cells did not (Fig. 1b, c). It is also important to note that WNT11 and some NE markers showed different expression levels between ASCL1-transfected A549 clones, in spite of the similar expression levels of ASCL1 (Fig. 2a). These results suggest that H3K27Ac levels at the enhancer region of the WNT11 gene are related to the balance between ASCL1 and WNT11 protein expression in these cell lines.
a ChIP-seq shows ASCL1 binding near the WNT11 gene in human SCLC cell lines (common to NCI-H889, NCI-H128, and NCI-H2107). An integrated genome viewer snapshot was shown from the study by Borromeo et al. [13]. b Manual ChIP assays for H3K27Ac at the 10-kb upstream lesions of WNT11. The relative amounts of immunoprecipitated DNA are depicted as a percentage of input DNA. H3K27Ac levels significantly increased at the targeted chromosome site in H889 cells, but not in SBC1 cells. c Relative comparison of H3K27Ac levels between ASCL1-TF A549 clones and mock cells. H3K27Ac levels increased in accordance with WNT11 expression in ASCL1-TF A549 cells. Data are shown as means ± SD. Asterisks indicate a significant difference. *p < 0.05. TF transfected
The role of Wnt11 in SCLC cell lines
To investigate the biological significance of Wnt11 in SCLC cell lines, we conducted WNT11 knockdown experiments using RNAi interference in H69 and H889 cells. The knockdown of WNT11 in these cells resulted in significant reductions in E-CADHERIN and SNAIL. Moreover, the knockdown of WNT11 decreased NE molecules; however, not all NE molecules were downregulated in the cell lines examined. This result suggests that Wnt11 promotes NE differentiation in the presence of Ascl1. The knockdown of WNT11 did not affect ASCL1 expression in the cell lines examined. Glycogen synthase kinase-3a/β (GSK3α/β), one of the essential components in the canonical Wnt pathway, decreased in the WNT11 mRNA knockdown–H69 cell line, but not in the WNT11 knockdown–H889 cell line. This result suggests that Wnt11 affects the canonical Wnt pathway in SCLCs in a context-dependent manner in each cell line (Fig. 4a). The cell-counting assay revealed that the knockdown of WNT11 suppressed cell growth and the knockdown of ASCL1 in these cells (Fig. 4b). This result indicates that Wnt11 and Ascl1 positively affect the proliferation of SCLC cells.
a Wnt11 affects NE differentiation and E-cadherin expression in SCLC cell lines. The suppression of E-CADHERIN and SNAIL was observed in H69 and H889 cells with siRNA for WNT11. Wnt11 was also involved in the expression of NE molecules, but not Ascl1 in these cell lines. b Cell- counting assays with SCLC cell lines. The cell growth curve is shown. The suppression of cell proliferation was observed in H69 and H889 cells with siRNA against WNT11 and ASCL1. The analysis was performed in triplicate. Data are shown as means ± SD. Asterisks indicate a significant difference. *p < 0.05. c Wnt11 affected NE differentiation and the canonical Wnt pathway in ASCL1-transfected A549 cells. The suppression of CHGA was observed in these cells with siRNA. Wnt11 is also involved in the expression of the canonical Wnt pathway components, GSK-3α/β and MYC. The knockdown of WNT11 expression modulated the apoptotic pathway and proliferative activity. The knockdown of WNT11 expression increased cleaved CASPASE3, but decreased p-HH3. NT nontreated, si small interfering, p- phospho-, TF transfected
The role of Wnt11 in an ASCL1-transfected ADC cell line
To investigate the biological significance of Wnt11 in an ASCL1-transfected ADC cell line, we conducted WNT11 knockdown experiments using RNAi interference in ASCL1-transfected A549 cells. The knockdown of WNT11 expression in this cell line resulted in a significant reduction in the protein expression of GSK3α/β, MYC, and CHGA. The canonical Wnt pathway induces Myc through Gsk3α/β and other Wnt signal components. This result suggests that Wnt11 affects canonical pathways in this cell. CHGA, but not SYP, decreased after the knockdown of WNT11. This result shows that Wnt11 may also promote NE differentiation in an Ascl1-dependent manner in lung cancer cell lines. Furthermore, the knockdown of WNT11 also decreased the phosphorylation of histone H3 (p-HH3) and increased the caspase pathway (Fig. 4c). These results suggest that Wnt11 affects cell proliferation and apoptosis in lung cancer.
The role of Wnt11 in proliferation and apoptosis in SCLC cell lines
To investigate the biological significance of Wnt11 in SCLC cell lines, we performed a WNT11 transfection experiment. A SBC3 cell line stably transfected with WNT11 was established. SBC3 is a representative SCLC cell line that hardly expresses Ascl1 and Wnt11 molecules (Fig. 1b, c). WNT11 was clearly detected in the cytoplasmic and perinuclear areas of WNT11-transfected SBC3 cells. The same level of phase-contrast microscope images were shown in mock and WNT11-transfected SBC3 cells (Fig. 5a). This result showed that the phosphorylation of GSK3α/β, AKT, and p38 mitogen-activated protein kinase markedly increased after the overexpression of WNT11 (Fig. 5b). The cell growth curve showed that Wnt11 promoted cell proliferation in transfected cells (Fig. 5c). These results suggest that Wnt11 is positively involved in cell proliferation in SCLC cell lines. WNT11-transfected H1975 ADC cells, which were preliminarily established, showed similar results (Supplementary Fig. S1).
We also demonstrated that WNT11-transfected SBC3 cells did not show significant changes in NE markers or NE-promoting transcription factors, such as ASCL1 and INSM1, by qRT-PCR (Fig. S2). Notch pathway components, including NOTCH1 or NOTCH2, HES1, delta-like ligand (DLL) 1, DLL3, and jagged ligand 1 (JAG1), which may affect the NE to non-NE cell fate switch, did not show significant changes in the cell (Fig. S3). We also did not observe any significant changes in EMT markers or the promotion of the cell invasive capacity measured in these cells using invasion chambers (Fig. S4). Similarly, the expression levels of APC regulator of WNT signaling pathway (APC), which is reported to be mutated in several cancers and is an established negative regulator of β-catenin and Wnt signaling [36], did not change after the overexpression of Wnt11 in cells (Fig. S5).
Canonical Wnt pathway inhibition by IWR-1-endo is not sufficient to affect NE differentiation or E-CADHERIN and SNAIL expression in SCLC cell lines
We inhibited a canonical Wnt pathway using a treatment with IWR-1-endo in order to identify which Wnt pathways are more significant in the regulation of NE differentiation or E-CADHERIN and SNAIL expression in SCLC. IWR-1-endo is a potent inhibitor of the Wnt response, blocking the Wnt/β-catenin pathway. On the other hand, the Wnt11 ligand has been reported to activate canonical and noncanonical Wnt pathways [22, 23]. It may be beneficial to compare the effects of Wnt11 inhibition and IWR-1-endo in SCLC. Tankyrase, a poly-ADP-ribose polymerase enzyme, regulates canonical Wnt activity [37]. Busch et al. showed that IWR-1-endo dose-dependently repressed cell growth and increased tankyrase levels in NSCLC cell lines [38]. Taking these findings into consideration, we used tankyrase to monitor the effects of the IWR-1-endo treatment in this study. TANKYRASE was markedly upregulated after the IWR-1-endo treatment in the SCLC cell lines examined. However, there were no other changes in the expression of molecules that were associated with NE differentiation or E-CADHERIN and SNAIL (Fig. 6). This result suggests that the noncanonical Wnt pathway is needed to affect NE differentiation or E-CADHERIN and SNAIL expression.
a WNT11 transfection in SBC3 cells. Cell shapes did not appear to change after the transfection of WNT11. GFP showed the localization of WNT11 in the perinuclear region, cytoplasm, or cell membrane in WNT11-TF SBC3 cells. TF transfected. Scale bar = 100 μm. b WB analyses revealed that the overexpression of WNT11 in WNT11-TF SBC3 cells markedly increased the phosphorylation of GSK3α/β, AKT, and P38 mitogen-activated protein kinase. All analyses were performed in triplicate. β-actin was used as an internal control. NT nontreated. c A cell- counting assay with WNT11-transfected SBC3 and mock cells. The cell growth curve is shown. Cell proliferation was promoted by the forced expression of WNT11. The analysis was performed in triplicate. Data are shown as means ± SD. Asterisks indicate a significant difference. *p < 0.05
Canonical Wnt pathway inhibitory experiment using IWR-1-endo. The same amount of dimethyl sulfoxide (DMSO) was used as in the control. Apart from the upregulation of TANKYRASE, the expression of NE markers, E-CADHERIN and SNAIL, did not significantly change after the treatment. All analyses were performed in triplicate. β-actin was used as an internal control
Discussion
This study demonstrated that Wnt11 was more strongly expressed in SCLC than in NSCLC using IHC staining, a WB analysis, and RT-PCR. We elucidated the significance of Wnt11 in NE differentiation, cell proliferation, and E-cadherin expression, particularly in SCLC. The results obtained suggest the potential of Wnt11 as a therapeutic target in the treatment of SCLC.
We revealed that Ascl1 regulates Wnt11 and other Wnt pathway components in lung cancer cells. WNT11 is one of the target genes of ASCL1 [13]. ASCL1 associates with an enhancer in multiple human SCLC cell lines that strongly express ASCL1 [39]. H3K27Ac-marked chromatin, called an enhancer, may contribute to Wnt11 expression, particularly in the presence of Ascl1. The overexpression of Ascl1 in the ADC cell line induced a decrease in the expression level of Dkk1, a negative Wnt signaling pathway regulator [11, 40]. Meder et al. also reported that Ascl1 was sufficient to trigger canonical Wnt signaling via the phosphorylation of the co-receptor LDL receptor-related protein 6 [40]. Osada et al. showed that the knockdown of Ascl1 induced growth inhibition and apoptosis in SCLC cell lines [12]. The present results are consistent with these findings, and as a further novel insight, Wnt11 is one of the most important molecules involved in these molecular mechanisms and cellular biological behaviors. Ascl1-induced Wnt11 may regulate not only cell proliferation, but also NE differentiation and E-cadherin expression in human lung cancer.
A shift between canonical and noncanonical Wnt signaling pathways has been suggested to modulate the lung carcinogenic process. The upregulation of canonical Wnt7b has been detected in ADC cells, whereas the increased expression of Wnt5a was found in primary SCCs [30, 41]. In a similar manner, the upregulation of Wnt11 was detected, whereas the decreased expression of Wnt7b was observed in ASCL1-transfected A549 cells in this study. We also demonstrated that SCLC cells strongly expressed Wnt11, whereas Wnt7b was frequently expressed in NSCLC cell lines. Ascl1 may be a key regulator in coordinating the Wnt11–Wnt7b balance. Not only Wnt ligands, but also various Wnt signaling molecules changed their expression levels in an Ascl1-dependent manner. For example, SFRP1, which inhibits Wnt signaling by binding Wnt proteins, decreased after the transfection of Ascl1. Fukui et al. reported that SFRP1 is a candidate tumor suppressor gene for lung cancer [42]. Ascl1 may advantageously affect lung tumorigenesis by modulating the Wnt signaling pathway. ROR2, a Wnt binding receptor with an interaction via FZD2 [43], is induced under Ascl1-positive conditions. Bai et al. found that the ROR2 receptor responded to the Wnt11 ligand and regulated cell migration and cell morphology during gastrulation in zebrafish [44]. Minami indicated the importance of Wnt5a/ROR2 signaling in the progression of different types of cancers in humans [45]. Based on these findings, we hypothesize that ROR2 may function in the Wnt signaling pathway of SCLC in an Ascl1-dependent manner.
The knockdown of Wnt11 expression in the H69 and H889 SCLC cell lines resulted in significant reductions in E-cadherin and Snail. Bartis et al. investigated the molecular events in canonical and noncanonical Wnt signaling pathways in lung SCC, and reported that Wnt11 and Wnt5a potentiated the relocation of β-catenin to the nucleus as an important step in decreased cell adhesion [30]. We showed that Ascl1 and Wnt11 also regulated E-cadherin expression in SCLC cell lines. Our results strengthened their findings, and as a further novel insight, Wnt11 may affect E-cadherin expression and NE differentiation in a context-dependent manner, particularly in the presence of Ascl1. The SBC3 cell line stably transfected with WNT11 did not show changes in the expression of E-cadherin or NE markers in this study. On the other hand, Ascl1 was sufficiently potent to induce the modulation of E-cadherin expression and NE differentiation in lung cancer [14, 19]. Lim et al. reported that the endogenous activation of the Notch pathway may result in a NE to non-NE fate switch in 10–50% of tumor cells in a mouse model of SCLC and in human tumors [46]. Wnt11 overexpression did not affect the expression levels of Notch pathway components in the SBC3 cells examined. However, Wnt11 affected E-cadherin expression or the NE switch more strongly in an Ascl1-dependent manner. In other words, Wnt11 alone did not play a role in the non-NE to NE switch and E-cadherin expression in SCLC. Furthermore, Wagner et al. recently showed that relapsed SCLC exhibited recurrent mutations and the loss of heterozygosity in regulators of Wnt signaling, including APC [47]. Since they demonstrated that the activation of Wnt signaling through the knockdown of APC induced chemoresistance in human SCLC cell lines, we investigated whether Wnt11 overexpression affected APC expression. However, we did not detect significant APC changes in WNT11-transfected SBC3 cells.
Uysal-Onganer et al. previously showed that Wnt11 increased the expression of neuron-specific enolase (NSE) and Ascl1 in prostate cancer [29]. Their findings were somewhat contradictory with the present results. We revealed that Ascl1 drove Wnt11 and NE marker expression in lung cancer, whereas the opposite phenomenon in prostate cancer was not observed in our case. These differences suggest that the Ascl1–Wnt11 and Wnt11–Ascl1 axes both exist in cell-specific context-dependent manners. Unfortunately, they screened NE differentiation using NSE alone. CHGA, SYP, NCAM, and INSM1 are specific NE markers [19]; therefore, it will be interesting to examine Wnt11-expressing prostate cancer with any other NE marker in addition to NSE.
We performed a canonical Wnt inhibitory experiment using IWR-1-endo to compare the effects of Wnt11 inhibition in SCLC. Many types of Wnt ligands and receptors are known to collaborate with each other in order to regulate cell-specific Wnt signaling events. In this study, we aimed to compare the effects of Wnt11 inhibition by RNAi and the inhibition of canonical Wnt signaling by IWR-1-endo. The results obtained showed that IWR-1-endo was insufficient to affect NE differentiation or E-cadherin and Snail expression in SCLC. These results were in contrast to those from the Wnt11 inhibition experiment. Previous studies demonstrated that canonical and noncanonical Wnt pathway suppression may be induced by the inhibition of Wnt11 [22, 30]. These findings indicate that the noncanonical pathway is essential for regulating NE differentiation, and E-cadherin and Snail expression in SCLC. Previous findings showed that the PCP signaling pathway is activated by Wnt11, which activates RhoA and Ras-related C3 botulinum toxin substrate 1. These, in turn, activate the stress kinase Jun N-terminal kinase and Rho-associated coiled-coil-containing protein kinase 1, which initiates cytoskeleton remodeling, and ultimately, changes in cell adhesion and motility [48,49,50].
In summary, Wnt11 is frequently overexpressed in SCLC and is associated with the expression of Ascl1. Ascl1 plays an important role in regulating the Wnt signaling pathway. We demonstrated that Wnt11 regulates NE differentiation and E-cadherin expression in SCLC cell lines in vitro. We also revealed the significance of Wnt11 and Ascl1 in the cell growth of Ascl1-positive SCLC cell lines. Collectively, these results suggest the potential of the Ascl1–Wnt11 axis as a therapeutic target and support the clinical utility of Wnt11 as a biological marker in SCLC. In further studies on Wnt11, it will be necessary to focus on highly specific molecules that are involved in Wnt11 signaling in SCLC. This study promotes our understanding of the significance of Wnt11 in SCLC, which may lead to the development of novel targeted therapies and improvements in the prognosis of patients with SCLC.
References
Rodriguez E, Lilenbaum RC. Small cell lung cancer: past, present, and future. Curr Oncol Rep. 2010;12:327–34.
Mitsudomi T, Morita S, Yatabe Y, Negoro S, Okamoto I, Tsurutani J, et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): an open label, randomised phase 3 trial. Lancet Oncol. 2010;11:121–8.
Kwak EL, Bang YJ, Camidge DR, Shaw AT, Solomon B, Maki RG, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med. 2010;363:1693–703.
Wu YL, Yang JC, Kim DW, Lu S, Zhou J, Seto T, et al. Phase II study of crizotinib in east asian patients with ROS1-positive advanced non-small-cell lung cancer. J Clin Oncol. 2018;36:1405–11.
Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S, Hatano S, et al. RET, ROS1 and ALK fusions in lung cancer. Nat Med. 2012;18:378–81.
Gazdar AF, Bunn PA, Minna JD. Small-cell lung cancer: what we know, what we need to know and the path forward. Nat Rev Cancer. 2017;17:765.
May M, Siegsmund M, Hammermann F, Loy V, Gunia S. Prognostic significance of proliferation activity and neuroendocrine differentiation to predict treatment failure after radical prostatectomy. Scand J Urol Nephrol. 2007;41:375–81.
Linnoila RI, Zhao B, DeMayo JL, Nelkin BD, Baylin SB, DeMayo FJ, et al. Constitutive achaete-scute homologue-1 promotes airway dysplasia and lung neuroendocrine tumors in transgenic mice. Cancer Res. 2000;60:4005–9.
Travis WD. Pathology and diagnosis of neuroendocrine tumors: lung neuroendocrine. Thorac Surg Clin. 2014;24:257–66.
Ball DW, Azzoli CG, Baylin SB, Chi D, Dou S, Donis-Keller H, et al. Identification of a human achaete-scute homolog highly expressed in neuroendocrine tumors. Proc Natl Acad Sci USA. 1993;90:5648–52.
Osada H, Tomida S, Yatabe Y, Tatematsu Y, Takeuchi T, Murakami H, et al. Roles of achaete-scute homologue 1 in DKK1 and E-cadherin repression and neuroendocrine differentiation in lung cancer. Cancer Res. 2008;68:1647–55.
Osada H, Tatematsu Y, Yatabe Y, Horio Y, Takahashi T. ASH1 gene is a specific therapeutic target for lung cancers with neuroendocrine features. Cancer Res. 2005;65:10680–5.
Borromeo MD, Savage TK, Kollipara RK, He M, Augustyn A, Osborne JK, et al. ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep. 2016;16:1259–72.
Ito T, Kudoh S, Ichimura T, Fujino K, Hassan WA, Udaka N. Small cell lung cancer, an epithelial to mesenchymal transition (EMT)-like cancer: significance of inactive Notch signaling and expression of achaete-scute complex homologue 1. Hum Cell. 2017;30:1–10.
Rizzo P, Osipo C, Foreman K, Golde T, Osborne B, Miele L. Rational targeting of Notch signaling in cancer. Oncogene. 2008;27:5124–31.
Wael H, Yoshida R, Kudoh S, Hasegawa K, Niimori-Kita K, Ito T. Notch1 signaling controls cell proliferation, apoptosis and differentiation in lung carcinoma. Lung Cancer. 2014;85:131–40.
Ito T, Udaka N, Yazawa T, Okudela K, Hayashi H, Sudo T, et al. Basic helix-loop-helix factors regulate the neuroendocrine differentiation of fetal mouse pulmonary epithelium. Development. 2000;127:3913–21.
Morimoto M, Nishinakamura R, Saga Y, Kopan R. Different assemblies of Notch receptors coordinate the distribution of the major bronchial Clara, ciliated and neuroendocrine cells. Development. 2012;139:4365–73.
Fujino K, Motooka Y, Hassan WA, Ali Abdalla MO, Sato Y, Kudoh S, et al. INSM1 is a crucial regulator of neuroendocrine differentiation in lung cancer. Am J Pathol. 2015;185:3164–77.
Ball DW. Achaete-scute homolog-1 and Notch in lung neuroendocrine development and cancer. Cancer Lett. 2004;204:159–69.
Pongracz JE, Stockley RA. Wnt signaling in lung development and diseases. Respir Res. 2006;7:15.
Rapp J, Jaromi L, Kvell K, Miskei G, Pongracz JE. WNT signaling - lung cancer is no exception. Respir Res. 2017;18:167.
Stewart DJ. Wnt signaling pathway in non-small cell lung cancer. J Natl Cancer Inst. 2014;106:djt356.
Uysal-Onganer P, Kypta RM. Wnt11 in 2011-the regulation and function of a non-canonical Wnt. Acta Physiol(Oxf). 2012;204:52–64.
Tada M, Concha ML, Heisenberg CP. Non-canonical Wnt signalling and regulation of gastrulation movements. Semin Cell Dev Biol. 2002;13:251–60.
Kirikoshi H, Sekihara H, Katoh M. Molecular cloning and characterization of human WNT11. Int J Mol Med. 2001;8:651–6.
Ueno K, Hazama S, Mitomori S, Nishioka M, Suehiro Y, Hirata H, et al. Down-regulation of frizzled-7 expression decreases survival, invasion and metastatic capabilities of colon cancer cells. Br J Cancer. 2009;101:1374–81.
Lin Z, Reierstad S, Huang CC, Bulun SE. Novel estrogen receptor-alpha binding sites and estradiol target genes identified by chromatin immunoprecipitation cloning in breast cancer. Cancer Res. 2007;67:5017–24.
Uysal-Onganer P, Kawano Y, Caro M, Walker MM, Diez S, Darrington RS, et al. Wnt-11 promotes neuroendocrine-like differentiation, survival and migration of prostate cancer cells. Mol Cancer. 2010;9:55.
Bartis D, Csongei V, Weich A, Kiss E, Barko S, Kovacs T, et al. Down-regulation of canonical and up-regulation of non-canonical Wnt signalling in the carcinogenic process of squamous cell lung carcinoma. PLos ONE. 2013;8:e57393.
Travis WD, Brambilla E, Burke AP, Marx A, Nicholson AG. WHO classification of tumours of the lung, pleura, thymus and heart. 4th edn. Lyon: IARC; 2015.
Yoshida R, Nagata M, Nakayama H, Niimori-Kita K, Hassan W, Tanaka T, et al. The pathological significance of Notch1 in oral squamous cell carcinoma. Lab Invest. 2013;93:1068–81.
Jiang L, Huang J, Higgs BW, Hu Z, Xiao Z, Yao X, et al. Genomic landscape survey identifies SRSF1 as a key oncodriver in small cell lung cancer. PLoS Genet. 2016;12:e1005895.
Chen B, Dodge ME, Tang W, Lu J, Ma Z, Fan CW, et al. Small molecule-mediated disruption of Wnt-dependent signaling in tissue regeneration and cancer. Nat Chem Biol. 2009;5:100–7.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-André V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Fearnhead NS, Britton MP, Bodmer WF. The ABC of APC. Hum. Mol. Genet. 2001;10:721–33.
Huang SM, Mishina YM, Liu S, Cheung A, Stegmeier F, Michaud GA, et al. Tankyrase inhibition stabilizes axin and antagonizes Wnt signalling. Nature. 2009;461:614–20.
Busch AM, Johnson KC, Stan RV, Sanglikar A, Ahmed Y, Dmitrovsky E, et al. Evidence for tankyrases as antineoplastic targets in lung cancer. BMC Cancer. 2013;13:211.
Christensen CL, Kwiatkowski N, Abraham BJ, Carretero J, Al-Shahrour F, Zhang T, et al. Targeting transcriptional addictions in small cell lung cancer with a covalent CDK7 inhibitor. Cancer Cell. 2014;26:909–22.
Meder L, König K, Ozretić L, Schultheis AM, Ueckeroth F, Ade CP, et al. NOTCH, ASCL1, p53 and RB alterations define an alternative pathway driving neuroendocrine and small cell lung carcinomas. Int J Cancer. 2016;138:927–38.
Vesel M, Rapp J, Feller D, Kiss E, Jaromi L, Meggyes M, et al. ABCB1 and ABCG2 drug transporters are differentially expressed in non-small cell lung cancers (NSCLC) and expression is modified by cisplatin treatment via altered Wnt signaling. Respir Res. 2017;18:52.
Fukui T, Kondo M, Ito G, Maeda O, Sato N, Yoshioka H, et al. Transcriptional silencing of secreted frizzled related protein 1 (SFRP1) by promoter hypermethylation in non-small-cell lung cancer. Oncogene. 2005;24:6323–7.
Li C, Chen H, Hu L, Xing Y, Sasaki T, Villosis MF, et al. Ror2 modulates the canonical Wnt signaling in lung epithelial cells through cooperation with Fzd2. BMC Mol Biol. 2008;9:11.
Bai Y, Tan X, Zhang H, Liu C, Zhao B, Li Y, et al. Ror2 receptor mediates Wnt11 ligand signaling and affects convergence and extension movements in zebrafish. J Biol Chem. 2014;289:20664–76.
Minami Y, Oishi I, Endo M, Nishita M. Ror-family receptor tyrosine kinases in noncanonical Wnt signaling: their implications in developmental morphogenesis and human diseases. Dev Dyn. 2010;239:1–15.
Lim JS, Ibaseta A, Fischer NM, Cancilla B, O'Young G, Cristea S, et al. Intratumoral heterogeneity generated by Notch signaling promotes small-cell lung cancer. Nature. 2017;545:360–4.
Wagner AH, Devarakonda S, Skidmore ZL, Krysiak K, Ramu A, Trani L, et al. Recurrent WNT pathway alterations are frequent in relapsed small cell lung cancer. Nat Commun. 2018;9:3787.
Vladar EK, Antic D, Axelrod JD. Planar cell polarity signaling: the developing cell’s compass. Cold Spring Harb Perspect Biol. 2009;1:a002964.
Wang Y. Wnt/planar cell polarity signaling: a new paradigm for cancer therapy. Mol Cancer Ther. 2009;8:2103–9.
Li C, Bellusci S, Borok Z, Minoo P. Non-canonical Wnt signalling in the lung. J. Biochem. 2015;158:355–65.
Acknowledgements
We thank Ms. Sachiko Nakata, Ms. Yuko Fukuchi, and Ms. Takako Maeda for their technical assistance; Mr. Shingo Usuki and the staff of the LILA for their technical support; and the Institute of Molecular Embryology and Genetics, Kumamoto University, for their help with the RNA sequence analysis. This study was supported in part by the program of the Joint Usage/Research Center for Developmental Medicine, Institute of Molecular Embryology and Genetics, Kumamoto University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tenjin, Y., Kudoh, S., Kubota, S. et al. Ascl1-induced Wnt11 regulates neuroendocrine differentiation, cell proliferation, and E-cadherin expression in small-cell lung cancer and Wnt11 regulates small-cell lung cancer biology. Lab Invest 99, 1622–1635 (2019). https://doi.org/10.1038/s41374-019-0277-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-019-0277-y
This article is cited by
-
Beyond Hormones: Investigating the Impact of Progesterone Receptor Membrane Component 1 in Lung Adenocarcinoma
The Journal of Membrane Biology (2024)
-
WNT ligands in non-small cell lung cancer: from pathogenesis to clinical practice
Discover Oncology (2023)
-
Heterogeneous expression and role of receptor tyrosine kinase-like orphan receptor 2 (ROR2) in small cell lung cancer
Human Cell (2022)
-
Long non-coding RNAs in lung cancer: implications for lineage plasticity-mediated TKI resistance
Cellular and Molecular Life Sciences (2021)
-
Talin1 Induces Epithelial-Mesenchymal Transition to Facilitate Endometrial Cell Migration and Invasion in Adenomyosis Under the Regulation of microRNA-145-5p
Reproductive Sciences (2021)