Abstract
A variety of laboratory methods are available for the detection of deletions of tumor suppressor genes and losses of their proteins. The clinical utility of fluorescence in situ hybridization (FISH) for the identification of deletions of tumor suppressor genes has previously been limited by difficulties in the interpretation of FISH signal patterns. The first deletion FISH assays using formalin-fixed paraffin-embedded tissue sections had to deal with a significant background level of signal losses affecting nuclei that are truncated by the cutting process of slide preparation. Recently, more efficient probe designs, incorporating probes adjacent to the tumor suppressor gene of interest, have increased the accuracy of FISH deletion assays so that true chromosomal deletions can be readily distinguished from the false signal losses caused by sectioning artifacts. This mini-review discusses the importance of recurrent tumor suppressor gene deletions in human cancer and reviews the common FISH methods being used to detect the genomic losses encountered in clinical specimens. The use of new probe designs to recognize truncation artifacts is illustrated with a four-color PTEN FISH set optimized for prostate cancer tissue sections. Data are presented to show that when section thickness is reduced, the frequency of signal truncation losses is increased. We also provide some general guidelines that will help pathologists and cytogeneticists run routine deletion FISH assays and recognize sectioning artifacts. Finally, we summarize how recently developed sequence-based approaches are being used to identify recurrent deletions using small DNA samples from tumors.
Similar content being viewed by others
Introduction
Genomic deletions are one of the major types of somatic mutations that inactivate tumor suppressor genes in human cancers. Some of these inactivation events occur as point mutations or deletions of a few base pairs of DNA that are only detectable by sequence-based methods. However, many of the clinically most important deletions occur as interstitial losses of the entire genomic region containing the tumor suppressor gene. Although there are a number of ways of inactivating a tumor suppressor gene, the end result is nearly always a mutational disruption or loss of coding information, leading to the absence of a functional protein. If protein expression is lost it means that often antibody based methods, such as immunohistochemistry may be used to evaluate loss of tumor suppressor genes.
Historically, classical cytogenetic analysis of tumor karyotypes was used to identify interstitial deletions of a chromosomal region known to contain a suppressor gene. However, fluorescence in situ hybridization (FISH) has a greatly improved sensitivity, and the technique is particularly useful when examining pathology tumor tissue preparations in which a suppressor gene deletion is of clinical importance. One of the main advantages of FISH lies in its ability to detect deletions directly in formalin-fixed paraffin-embedded (FFPE) tumor sections, so that gene loss may be examined in the context tumor tissue morphology.
Deletion testing by FISH of solid tumors is usually performed using FFPE tissue sections of a biopsy, with an adjacent section hematoxylin–eosin stained section available for reference purposes during analysis. Enumerating the gene copy number by FISH may be compromised by the partial sectioning or ‘signal truncation’ of tumor nuclei during the cutting process of slide preparation [1, 2]. The loss of some signals due to sectioning truncations mean that a high background of ‘false deletions’ can make interpretation more complicated (discussed in detail below). One of the most important sample types to study by FISH or immunohistochemistry in many newly diagnosed cancers are taken by fine needle and core biopsies [3]. Often these initial analyses obtained from needle cores can be used for FISH of immunohistochemistry to provide additional prognostic or diagnostic information depending on the tumor type and the assay of interest. These tiny cylindrical cores of tissue are usually embedded in paraffin blocks. Sections are then cut from the blocks using a microtome typically set to produce sections ranging from 3 to 5 µm in thickness in most laboratories. Sections of 3 µm thickness are particularly challenging for deletion FISH analyses because of loss of parts of the nuclear material during slide sectioning. This issue may have limited the uptake of FISH deletion assays in clinical laboratories however, there are new probe designs that can be used to recognize when slide sectioning losses may be present. In this mini-review, we will summarize the various laboratory approaches taken to determine whether genomic deletion of a tumor suppressor gene has taken place. We will summarize the common FISH methods used to detect the genomic losses encountered in clinical specimens, and we will also present one of the multicolor FISH probe designs that have been used recently to increase the accuracy of FISH deletion assays, even when using thin 3 µm sections. Finally, we will provide some guidelines that may be helpful for laboratories that are considering running FISH deletion analysis of tumor suppressor genes using FFPE tissue sections.
Clinically significant gene losses in human cancer
Over the years, loss of heterozygosity analysis, copy-number array comparative genomic hybridization and more recently, whole genome sequencing of tumor DNA, have uncovered a large number of gene deletions, complex insertions and sequence deletions (indels). However, the number of tumor suppressor gene deletion assays being routinely performed by FISH methods using clinical samples is still surprisingly limited. Reduced uptake of these tests for clinical uses may be partly because the diagnostic or prognostic value of loss of a tumor suppressor gene is only just emerging for some tumor types. Also, for many tumors, the best laboratory approach to determining whether a suppressor gene of interest has been deleted has not been established at the rigorous level required for a clinical grade test. Some of the clinically most relevant tumor suppressor gene deletion assays are shown in Table 1. This list is not intended to be comprehensive; we have highlighted assays that illustrate their clinical importance and diversity of the laboratory approaches that are currently being used to detect gene loss.
Homozygous deletion of 9p21, the cytoband harboring the p16/CDKN2A gene, has been reported as the most common genetic alteration in malignant mesotheliomas [4]. FISH using probes from the p16/CDKN2A gene have been helpful for distinguishing between benign and malignant mesothelial proliferations [5]. More recently the use of immunohistochemistry in combination with FISH has also been helpful in mesothelioma diagnosis [6].
Determination of chromosome 1p36 and 19q13 status by FISH is a crucial step in the diagnosis and the management of oligodendroglial brain tumors (reviewed in ref. 7). Based on current WHO criteria, demonstrating 1p36/19q13 co-deletion is a requirement for confirming the diagnosis of oligodendroglioma [8]. While there are alternative molecular method methods that have been used to identify co-deletion of 1p and 19q, such as loss of heterozygosity or quantitative PCR analysis, the use of deletion FISH seems to be the favored means of testing at the present time [9]. In typical FISH assays probes of different colors are used to label DNA probes to detect 1p36 and 19q13 losses within the interphase nuclei of individual glioma cells from FFPE tissue sections. Changes in the 1p36 and 19q13 probe signal pattern are then compared with controls to determine if the 1p36 and 19q13 co-deletion is present. Co-deletion is strongly correlated with a better response to standard treatment with radiotherapy and chemotherapy as well as a better overall survival.
In chronic lymphocytic leukemia deletions of the 17p13 chromosomal region, which includes the TP53 gene encoding the p53 protein, is a powerful predictor of resistance to chemotherapy and an overall poor outcome [10]. Abnormalities of p53 are also a common finding with potential therapeutic targeting in multiple myelomas [11]. The presence of hemizygous TP53 loss and nuclear p53 protein expression by immunohistochemistry in multiple myeloma can be an indication of an adverse outcome [12].
There is a strong correlation between loss of the PTEN gene and its protein and adverse pathologic features in prostate cancer (reviewed in [13]). PTEN gene loss as determined by both FISH and immunohistochemistry has been associated with biochemical recurrence [14], development of castrate resistant disease [15, 16], and prostate cancer-specific death [17]. Based on recent studies using diagnostic needle biopsies, it has been proposed that an initial analysis of PTEN expression could be carried out using immunohistochemistry. Thereafter, regions of tumor or suspicious areas in the biopsy that have reduced expression of PTEN protein, or are otherwise indeterminate by immunohistochemistry, could then be analyzed by PTEN FISH as a reflex test [18]. This general approach would be in keeping with the emerging consensus for several types of oncologic pathology tests such as those for breast [19], lung [6, 20], and melanomas [21], in which the primary immunohistochemistry analysis can be followed by reflex FISH for cases that are inconclusive.
FISH deletion testing using solid tumor tissue sections
In typical FISH deletion assays of solid tumors, the tissue biopsy has been fixed in formalin and then embedded in wax in a mold to produce tissue blocks that are then cut on a microtome (Supplementary Fig. 1A). Some of the earliest probe designs used for tumor suppressor gene deletion assays incorporated two-color FISH probes from two regions on the same chromosome: one specific probe targeting the tumor suppressor gene of interest, and a second control probe, often a centromere probe, that is used to count the number of chromosomes present [22]. Unfortunately, the interpretation of the FISH signals using this basic assay design for FFPE sections can be complicated by signal truncation losses generated by the histological slide making process itself (Supplementary Fig. 1B). Signal truncations are more evident in sections when the genomic distance between the control centromere probe and the tumor suppressor gene probe is large (>3–4 Mb). The significant background percentage of nuclei with ‘false deletions’ caused by the truncated nuclei in each section requires careful assay design and interpretation since the background truncation losses increase the probability of incorrectly classifying a tumor sample as deleted [2]. It is therefore essential to have matched control tissue and rigorous ‘cutoff’ values that take these truncation losses into account [22–24].
FISH analysis using a four-color deletion assay
A newer approach to increase the specificity of FISH deletion assays is to utilize additional probes labeled with different colors that are positioned to allow sectioning artifacts to be recognized and excluded from the overall scoring. Three- and four-color FISH assays have been used in a number different diagnostic settings to increase the specificity for detecting the aberration of interest [25–28]. Multicolor deletion FISH assays usually incorporate a specific probe for detecting loss of the tumor suppressor gene, in combination with one or more control probes to determine whether signal truncation has taken place.
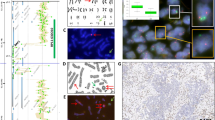
One of the best examples of four-color deletion FISH is the PTEN gene in prostate cancer [29]. We will use this tumor suppressor gene assay to illustrate the benefit of using additional control probes to identify nuclei that have been truncated by sectioning. The control probe combination was selected based on the idea that having adjacent probes that map either side of the PTEN tumor suppressor gene would allow for: (i) improved recognition of nuclear truncation losses; and (ii) would be helpful for mapping any larger deletions that may have extended outside the PTEN region. The best position for choosing flanking probes was found to be at the edge of the region of most frequent PTEN gene loss (Fig. 1a, b). When a nucleus loses both flanking probes together with the PTEN gene, there is a high chance that the nucleus was subject to truncation. In Fig. 1c, we show how the flanking probes are used to recognize a truncated nucleus as a schematic diagram. The concurrent loss of PTEN and both its flanking probes is a strong indication that the nucleus was sectioned during preparation. In Fig. 1d, the way that the flanking probes are used to recognize truncated nuclei in an analysis of part of a tissue section is depicted as a three-dimensional schematic representation.
Design of flanking probes to detect tumor suppressor gene deletions illustrated using an example of the PTEN gene. a Genomic deletion sizes of chromosome 10 derived from public domain datasets. Each horizontal thin red line represents an individual deletion and the best location for the selection of probes flanking PTEN (red vertical line) is just outside the minimal region of PTEN deletion (green and blue vertical lines). b Position of flanking probes WAPAL (green box) and FAS (aqua box) selected because they map outside the PTEN (red box) minimal region of deletion shown in b. The centromere probe (magenta box) is included to help determine the number of chromosome 10 s in the sample. c Schematic diagram to illustrate how a four-color probe design can be used to recognize truncation losses due to sectioning in a nucleus (gray spheres). In this illustration, the PTEN probe (red spots) is flanked either side of the gene by two probes (blue and green spots) on the two black wavy lines representing chromosome 10, which is enumerated by the CEP10 magenta probe. When there is loss of red PTEN with both blue and green flanking probes it indicates that sectioning (shown schematically as a dashed blue line in middle nucleus) has removed nuclear material that contained part of chromosome 10. However, when an actual interstitial deletion of PTEN is present, both flanking probes are retained (shown in right nucleus), indicating that the loss of a PTEN signal is due to an actual deletion. d Schematic three-dimensional representation to illustrate how four-color FISH is used to identify various truncated nuclei in a FFPE section. The nuclei are depicted as uniform pale blue spheres that are distributed randomly throughout the cuboid three-dimensional volume that represents part of a 5 µm section (left). In each nucleus the same spot colors as a are used to depict PTEN (red), the flanking probes (blue and green), and CEP10 (magenta). The thick blue dashed double arrowed vertical line points to the interface of two adjacent sections illustrating how truncated nuclei may be identified. It can be seen that there are six nuclei (red arrows) which will be recognizable with this probe set as truncated nuclei by the concurrent losses of a red PTEN probe with both green and blue flanking probes
Four-color FISH has been used to characterize the PTEN deletion status of prostatic carcinomas from both radical prostatectomy samples [29–31] and from needle core biopsies [32]. In these analyses, tumors in which there is no PTEN deletion (Fig. 2a) can readily be distinguished from truncated nuclei with PTEN losses caused by sectioning affects. For tumors with a homozygous deletion of PTEN (Fig. 2b) there is a complete absence of the PTEN specific probe so that tumors with this aberration are relatively easy to interpret. Nevertheless, some truncated nuclei can be readily detected in sections as signal losses affecting the flanking probes and a centromere. The benefit of including flanking probes is more apparent for tumors with a hemizygous deletion in which only one copy of the PTEN gene is lost (Fig. 2c). Careful examination of nuclei in these cases shows that the predominant pattern in scorable tumor nuclei is cells with only one PTEN gene copy. In a few cells, nuclear truncation effects can lead to loss of both PTEN genes, but the concurrent loss of flanking probes with PTEN is used to minimize the risk of misclassifying a hemizygous tumor as being homozygously deleted.
Four-color detection of PTEN deletions in FFPE sections. Representative four-color FISH images of 5 µm sections from radical prostatectomy samples are used to show the typical distribution of red PTEN signals and to illustrate how the blue FAS flanking probe and green WAPAL flanking probes aid in the detection of truncated nuclei that could otherwise be misleading. In this figure and all other the PTEN, FAS and WAPAL probe are labeled with these same colors, and the CEP10 probe is magenta (PTEN DEL-TECT Biocare, Pacheco, CA, USA). In all panels, the scorable nuclei without apparent loss of signal due to sectioning are indicated with blue arrows. Nuclei that are not scorable as a result sectioning affects are indicated with red arrows. One hundred nuclei were scored for each tumor to determine their respective PTEN deletion status. Scale bar, 2 µm. A schematic interpretation of the probe configuration on each chromosome 10 is shown beneath each panel. a Undeleted for PTEN. In the right panel, two red PTEN signals are apparent in all four scorable nuclei. However, two nuclei that have an apparent loss of PTEN can be recognized as being truncated (red arrows) as other probes are also lost with PTEN. b Homozygous PTEN deletion. In center panel, no PTEN signals are apparent in any nuclei. Some nuclei can be seen to be truncated (red arrows) because of loss of other probes. c Hemizygous PTEN deletion. In the right panel, the nuclei with blue arrows were scorable, and each has just one copy of the PTEN gene but retains flanking probes. The typical signal configuration shows seven signals with one red signal missing on all scorable nuclei. It is likely that this tumor has an interstitial deletion of chromosome 10 removing one copy of PTEN. This tumor is therefore classified as having a ‘hemizygous deletion’. Truncated nuclei with other signal losses in other probes are identified by red arrows
Another advantage of a four-color PTEN FISH assays is that is possible characterize the genomic heterogeneity of deletions within different regions of carcinoma [30]. The flanking probes can be used to identify any clonal variations of deletion size in more detail within the tumor sections. For some prostate cancers, the occurrence of two distinct regions bearing hemizygous and homozygous clonal deletions has been used to identify heterogeneity of PTEN losses (Fig. 3).
Detection of clonally distinct hemizygous and homozygous deletions in different regions of a tumor. The four-color PTEN FISH probe set can help characterize genomic heterogeneity and variation in deletion size. In this example, one side of the prostate gland (upper panel A) had tumor cells with a large hemizygous PTEN deletion (red probe) that extended into the WAPAL green flanking probe (see the schematic interpretation of deletion on the right). On the other side of the gland (lower panel B) tumor cells had this same large deletion on one chromosome 10, but the other copy of the PTEN gene was also deleted, retaining both flanking probes. Thus the tumor cells in this region of the gland was homozygously deleted for PTEN with no red signals evident (see the schematic interpretation of both deletions to the right of this panel)
Effect of section thickness on nuclear truncation losses
Nuclei in FFPE sections of cancers have a wide range of nuclear diameter. Prostate cancers typically have a mean diameter of about 6–10 µm [33, 34], and morphometric analysis has shown prostate cancer nuclei in FFPE sections often have an elliptical shape rather than being perfect spheres [35]. FISH signals are usually small discrete signals distributed within the nuclear space. To investigate the three-dimensional appearance and the spatial distribution of two-colored FISH signals within nuclei of tumor sections we performed high-resolution confocal imaging using FFPE prostate cancer sections. Three-dimensional views were created by building stacks of each optical slice collected at sequential Z-axis locations (Supplementary Movie 1). This analysis showed that the FISH signals were present randomly at different depths within the nuclear space and that some nuclei were partially present in the section and had a reduced number of FISH signals. We then captured images at different focal planes through a 5 µm prostate cancer section (which was not deleted for PTEN) to show how the PTEN and flanking probe FISH signals were distributed within nuclei and the effects of signal truncation (Supplementary Movie 2).
Early theoretical studies of the general effect of section thickness on the loss of FISH signals by truncation have also shown that when the thickness is <50% of the mean nuclear diameter the proportion of truncation losses increases [36]. We have estimated that the theoretical effect of reducing section thickness from 5 to 3 µm would increase the number of signal losses due to truncation by ~20% (Supplementary Material 1).
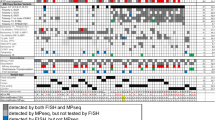
To directly investigate the influence of section thickness and truncation of nuclei on scoring results, we compared the FISH results of the four-color PTEN probe set on 3 µm FFPE tissue sections (Fig. 4a) to signal counts obtained from 5 µm FFPE normal tissue sections (Fig. 4b). Slides were evaluated by conventional fluorescence microscopy in the same regions previously defined and marked on each slide. These results showed that, while the percentage of signal truncations leading to potential misclassification of PTEN as a hemizygous loss was 11% in 5 µm sections, this frequency was much higher in 3 µm sections, with 37% having signal truncations leading to the false classification of PTEN deletion (Supplementary Material 2). Similarly when a PTEN hemizygously deleted tumor was analyzed using a 5 µm section, only 5% of cells could have been misclassified as being homozygously deleted without the use of flanking probes. This frequency of misclassification would have increased to 13% in the thinner 3 µm sections if the flanking probes had not been used to recognize signal truncation artifacts.
Effect of section thickness on nuclear truncation. The four-color PTEN FISH probe set was used to compare the relative amount of nuclear truncation when 3 µm (upper panel A) sections were compared to 5 µm (lower panel B) sections. In this comparison, we used one prostate carcinoma without an apparent PTEN deletion or other genomic alteration affecting the copy number of chromosome 10. In both panels, the scorable nuclei without apparent loss of signal due to sectioning are indicated with blue arrows. Nuclei that are not scorable as a result sectioning effects are indicated with red arrows. One hundred nuclei were scored for both 3 and 5 µm sections to determine their respective frequencies of nuclear truncation. Scale bar, 2 µm
Guiding principles for interpretation of FISH deletion assays
Tissue sections derived from solid tumors present a complex and challenging target for FISH assays. The distribution of nuclei within the section space is irregular, determined by multiple factors including the architecture of the tissue, as well as normal histologic variables such as differing levels of the intermingled tumor and adjacent stroma. In addition to these general histologic features, the nuclei within the tumor tissue vary in size as a function of their overall cell shape, their ploidy level, and the degree of local nuclear compaction present in the preparation. In Table 2, we present some of the general principles to consider when examining FFPE tissue sections using any deletion FISH assay.
Future of clinical deletion assays of solid tumors
This review has focused primarily on recent improvements for FISH testing in clinical laboratories to detect deletions of tumor suppressor genes in a solid tumor using FFPE tissue sections. However, newer ‘next-generation sequencing (NGS)’ and sequence-based technologies are providing alternative approaches to detecting both changes in gene copy number, and somatic point mutations [37]. NGS approaches can be optimized for analyzing DNA derived from tumor specimens that have been fixed in formalin [38]. These assays typically use methods such as multiplex PCR to isolate clinically relevant DNA segments of the genome, such as mutation hotspots or coding exons of entire oncogenes or tumor suppressor genes [39]. Counting the number of sequencing reads that align to a given genomic location is analogous to enumerating interphase FISH signals for a specific gene probe, but at extremely high resolution. One of the main general limitations of sequence-based methods for deletion detection is that often a solid tumor specimen contains significant infiltrations of normal stromal and inflammatory cells that may reduce the sensitivity of DNA copy-number measurements or make copy-number counting inconclusive. Future assessment of deletions for clinical needs will likely be based on combining findings from FISH, immunohistochemistry with customized NGS sequence-based detection methods.
Change history
08 March 2018
Figure 2 is incorrect in the original version of this article. The correct figure 2 is provided below
References
Hopman AH, van Hooren E, van de Kaa CA, et al. Detection of numerical chromosome aberrations using in situ hybridization in paraffin sections of routinely processed bladder cancers. Mod Pathol. 1991;4:503–13.
Piqueras M, Subramaniam MM, Navarrom SG, et al. Fluorescence in situ hybridridization (FISH) on formalin-fixed paraffin-embedded (FFPE) tissue sections. In: Stanta G editor. Guidelines for molecular analysis in archive tissues. Berlin Heidelberg: Springer-Verlag; 2011. p. 225–30.
Schneider F, Smith MA, Lane MC, et al. Adequacy of core needle biopsy specimens and fine-needle aspirates for molecular testing of lung adenocarcinomas. Am J Clin Pathol. 2015;143:193–200.
Chiosea S, Krasinskas A, Cagle PT, et al. Diagnostic importance of 9p21 homozygous deletion in malignant mesotheliomas. Mod Pathol. 2008;6:742–7.
Chung C, Santos GC, Hwang DM, et al. FISH assay development for the detection of p16/CDKN2A deletion in malignant pleural mesothelioma. J Clin Pathol. 2010;7:630–4.
Hida T, Hamasaki M, Matsumoto S, et al. Immunohistochemical detection of MTAP and BAP1 protein loss for mesothelioma diagnosis: comparison with 9p21 FISH and BAP1 immunohistochemistry. Lung Cancer. 2017;104:98–105.
Aldape K, Burger PC, Perry A. Clinicopathologic aspects of 1p/19q loss and the diagnosis of oligodendroglioma. Arch Pathol Lab Med. 2007;1:242–51.
Louis DN, Perry A, Reifenberger G, von Deimling A, et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131:803–20.
Woehrer A, Hainfellner JA, Molecular diagnostics: techniques and recommendations for 1p/19q assessment. CNS Oncol. 2015;4:295–306.
Zenz T, Krober A, Scherer K, et al. Monoallelic TP53 inactivation is associated with poor prognosis in chronic lymphocytic leukemia: results from a detailed genetic characterization with long-term follow-up. Blood. 2008;112:3322–9.
Teoh PJ, Chng WJ p53 abnormalities and potential therapeutic targeting in multiple myeloma, [Internet], Biomed Res Int. 2014;article ID:717919. Availble at http://dx.doi.org/10.1155/2014/717919.
Chang H, Yeung J, Qi C, Xu W. Aberrant nuclear p53 protein expression detected by immunohistochemistry is associated with hemizygous P53 deletion and poor survival for multiple myeloma. Br J Haematol. 2007;138:324–9.
Wise HM, Hermida MA, Leslie NR. Prostate cancer, PI3K, PTEN and prognosis. Clin Sci. 2017;131:197–210.
Yoshimoto M, Cunha IW, Coudry RA, et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br J Cancer. 2007;97:678–85.
Sircar K, Yoshimoto M, Monzon FA, et al. PTEN genomic deletion is associated with p-Akt and AR signalling in poorer outcome, hormone refractory prostate cancer. J Pathol. 2009;218:505–13.
Choucair K, Ejdelman J, Brimo F, et al. PTEN genomic deletion predicts prostate cancer recurrence and is associated with low AR expression and transcriptional activity. BMC Cancer. 2012;12:543
Reid AHM, Attard G, Ambroisine L, et al. Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at low and high risk of death from prostate cancer. Br J Cancer. 2010;102:678–84.
Lotan TL, Wei W, Ludkovski O, et al. Analytic validation of a clinical-grade PTEN immunohistochemistry assay in prostate cancer by comparison with PTEN FISH. Mod Pathol. 2016;29:904–14.
Rakha EA, Pinder SE, Bartlett JMS, et al. Updated UK Recommendations for HER2 assessment in breast cancer. J Clin Pathol. 2015;68:93–99.
Lindeman NI, Cagle PT, Beasley MB, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: Guideline from the College of American Pathologists, International Association for the study of lung cancer, and Association for Molecular Pathology. Arch Pathol Lab Med. 2013;137:828–60.
Tetzlaff MT, Wang W-L, Milless TL, et al. Ambiguous melanocytic tumors in a tertiary referral center: the contribution of fluorescence in situ hybridization (FISH) to conventional histopathologic and immunophenotypic analyses. Am J Surg Pathol. 2013;3712:1783–96.
Ventura RA, Martin-Subero JI, Jones M, et al. FISH analysis for the detection of lymphoma-associated chromosomal abnormalities in routine paraffin-embedded tissue. J Mol Diagn. 2006;8:141–51.
Wiktor AE, Van Dyke DL, Stupca PJ, et al. Preclinical validation of fluorescence in situ hybridization assays for clinical practice. Genet Med. 2006;8:16–23.
Wolff DJ, Bagg A, Cooley LD, et al. Guidance for fluorescence in situ hybridization testing in hematologic disorders. J Mol Diagn. 2007;9:134–43.
Chen S, Deniz K, Sung Y-S, et al. Ewing sarcoma with ERG gene rearrangements: A molecular study focusing on the prevalence of FUS-ERG and common pitfalls in detecting EWSR1-ERG fusions by FISH. Genes Chromosomes Cancer. 2016;55:340–9.
Lee HJ, Shin DH, Noh GY, et al. Combination of immunohistochemistry, FISH and RT-PCR shows high incidence of Xp11 translocation RCC: comparison of three different diagnostic methods. Oncotarget. 2017;8:30756–65.
Trejo Bittar HE, Luvison A, Miller C, et al. A comparison of ALK gene rearrangement and ALK protein expression in primary lung carcinoma and matched metastasis. Histopathology. 2017;71:269–77.
Yoshimoto M, Joshua AM, Chilton-MacNeill S, et al. Three-color FISH analysis of TMPRSS2/ERG fusions in prostate cancer indicates that genomic microdeletion of chromosome 21 is associated with rearrangement. Neoplasia. 2006;8:465–9.
Yoshimoto M, Ludkovski O, Degrace D, et al. PTEN genomic deletions that characterize aggressive prostate cancer originate close to segmental duplications. Genes Chromosomes Cancer. 2012;51:149–60.
Yoshimoto M, Ding K, Sweet JM, et al. PTEN losses exhibit heterogeneity in multifocal prostatic adenocarcinoma and are associated with higher Gleason grade. Mod Pathol. 2013;26:435–47.
Troyer DA, Jamaspishvili T, Wei W, et al. A multicenter study shows PTEN deletion is strongly associated with seminal vesicle involvement and extracapsular extension in localized prostate cancer. Prostate. 2015;75:1206–15.
Picanço-Albuquerque CG, Morais CL, Carvalho FLF, et al. In prostate cancer needle biopsies, detections of PTEN loss by fluorescence in situ hybridization (FISH) and by immunohistochemistry (IHC) are concordant and show consistent association with upgrading. Virchows Arch. 2016;468:606–17.
Bektaş S, Bahadir B, Doǧan Gün B, et al. The relation between Gleason score, and nuclear size and shape factors in prostatic adenocarcinoma. Turk J Med Sci. 2009;39:381–7.
Montironi R, Filho AL, Santinelli AM, et al. Nuclear changes in the normal-looking columnar epithelium adjacent to and distant from prostatic intraepithelial neoplasia and prostate cancer. Morphometric analysis in whole-mount sections. Virchows Arch. 2000;43:625–34.
Mohler JL, Figlesthaler WM, Zhang XZ, et al. Nuclear shape analysis for the assessment of local invasion and metastases in clinically localized prostate carcinoma. Cancer. 1994;74:2996–3001.
Pahlplatz MM, de Wilde PC, Poddighe P, et al. A model for evaluation of in situ hybridization spot-count distributions in tissue sections. Cytometry. 1995;20:193–202.
Khotskaya YB, Mills GB, Shaw KR. Next-generation sequencing and result interpretation in clinical oncology: challenges of personalized cancer therapy. Annu Rev Med. 2017;68:113–25.
Manson-Bahr D, Ball R, Gundem G, et al. Mutation detection in formalin-fixed prostate cancer biopsies taken at the time of diagnosis using next-generation DNA sequencing. J Clin Pathol. 2015;68:212–7.
Serrati S, De Summa S, Pilato B, et al. Next-generation sequencing: advances and applications in cancer diagnosis. Oncol Targets Ther. 2016;9:7355–65.
European Cytogenetics Association. FISH on histological sections of solid tumors: E.C.A. RECOMMENDATIONS. E.C.A. European Cytogeneticists Asscoiation Newsletter; 2012 p. 28–30.
Mascarello JT, Hirsch B, Kearney H, et al. Section E9 of the American College of Medical Genetics technical standards and guidelines: fluorescence in situ hybridization. Genet Med. 2011;13:667–75.
Deutsch EW, Ball C, Berman G, et al. Minimum information specification for in situ hybridization and immunohistochemistry experiments (MISFISHIE). Nat Biotechnol. 2008;26:305–12.
Squire JA, Marrano P, Kolomietz E. FISH in clinical cytogenetics. In: Beatty B, Mai S, Squire J, editors. FISH a practicsl approach. New York, NY: Oxford Univ. Press; 2002 p. 183–202.
Liehr T In Fluorecence in situ hybridization (FISH) application guide; Springer-Verlag, Berlin, 2016.
Acknowledgements
This research was initially supported by the Canadian Cancer Society grant no. 018124 and then by FAPESP grant no. 2015/09111-5. JAS has the CNPq Bolsa Produtividade em Pesquisa - Nível: PQ-1B grant no. 306864/2014-2.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
JAS has consulted for CymoGenDx LLC, and MY and JAS have a provisional patent application on probe sets to detect deletions
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Yoshimoto, M., Ludkovski, O., Good, J. et al. Use of multicolor fluorescence in situ hybridization to detect deletions in clinical tissue sections. Lab Invest 98, 403–413 (2018). https://doi.org/10.1038/s41374-017-0007-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-017-0007-2
This article is cited by
-
Intratumoral heterogeneity of CDKN2A deletions in IDH-mutant astrocytoma
Brain Tumor Pathology (2024)
-
CCND1 Amplification in Breast Cancer -associations With Proliferation, Histopathological Grade, Molecular Subtype and Prognosis
Journal of Mammary Gland Biology and Neoplasia (2022)
-
PTEN loss in Gleason grade 7 prostate tumors exhibits intratumoral heterogeneity and is associated with unfavorable pathological features
Applied Cancer Research (2019)