Abstract
Zebrafish have several advantages compared to other vertebrate models used in modeling human diseases, particularly for large-scale genetic mutant and therapeutic compound screenings, and other biomedical research applications. With the impactful developments of CRISPR and next-generation sequencing technology, disease modeling in zebrafish is accelerating the understanding of the molecular mechanisms of human genetic diseases. These efforts are fundamental for the future of precision medicine because they provide new diagnostic and therapeutic solutions. This review focuses on zebrafish disease models for biomedical research, mainly in developmental disorders, mental disorders, and metabolic diseases.
Similar content being viewed by others
Introduction
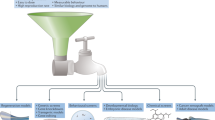
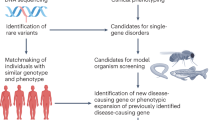
For use in genetic studies as an animal model, zebrafish was initially introduced by Streisinger and colleagues1 in the early 1980s. Large-scale N-ethyl-N-nitrosourea (ENU) mutagenesis was conducted in combination with extensive phenotypic screening2,3. Further phenotypic characterization of these ENU mutations in most of the major organ systems was performed4. However, later positional cloning of each ENU mutation after a forward genetic screen was time-consuming and laborious5. Since the increase in resolution of the zebrafish genome map, advanced gene-targeting technologies involving ZFNs, TALENs, and CRISPR/Cas96,7,8,9,10,11,12, have overcome challenges in generating specific gene-knockout mutations. CRISPR/Cas9 utilizes an efficient reverse genetic approach to provide knockout animals for zebrafish researchers13. Furthermore, the high level of genome structure shared between zebrafish and humans (~70% of human genes have at least one obvious zebrafish ortholog, compared to 80% of human genes with mouse orthologs)14,15 has facilitated the use of zebrafish for understanding human genetic diseases. Recent advancements in next-generation sequencing (NGS) coupled with the demand for personalized medicine has further driven zebrafish uses in identifying causal relationships between the genotype and phenotype of various human diseases.
Additionally, zebrafish possess several advantages over rodent models in the study of vertebrate development and disease. These include hundreds of embryos in a single clutch and optical clarity of the developing embryo, which allows live imaging at the organism level16,17. In addition, the use of tissue-specific transgenic animals can be easily generated under the control of various selected gene promoters. Recent improvement of the Tol2-based transgenic system in zebrafish18 has allowed the control of gene expression in a spatiotemporal manner by coupling with regulatory elements such as GAL4/UAS or Cre/LoxP19,20. These advantages allow live imaging of cells and tracking of cellular dynamics in vivo to study the underlying molecular mechanisms of various developing organs.
The necessity of a model organism to recapitulate metabolic symptoms and associated disease development in humans has led to the exploitation of several animal species, among which rodents have been widely employed. For the past several decades, mice have been the leading experimental animal model in the field of biomedical research due to powerful genetic tools, amenable diagnostic parameters that are comparable to those in humans, and standardized protocols for developing, diagnosing, and treating metabolic syndromes. However, factors inherently different from those in humans, such as dietary requirements, lifestyle, and microbiomes, have called for alternative animal model systems to be utilized in parallel21. Zebrafish is a fascinating animal model for understanding the human pathogenesis of metabolic diseases and identifying potential therapeutic options21. However, all animal models have unique shortcomings, are the zebrafish model is no exception: first, zebrafish are poikilothermal animals living under water. Nonetheless, zebrafish possess metabolic characteristics similar to humans to complement data obtained from other model organisms, including rodents. This possibility has been clearly shown in recent studies22,23,24,25 in which drugs that had been approved for alleviating metabolic syndromes in humans were also effective in a zebrafish model.
This review addresses the use of zebrafish as an animal model for biomedical research, mainly in developmental disorders, mental disorders, and communication between the brain and organs. In addition to biomedical research, we also discuss the utility of zebrafish in metabolic control, focusing on cellular metabolic organelles.
Biomedical research I: developmental disorders
During early animal development, an organizer can induce a complete body axis when transplanted to the ventral side of a host embryo. Studies have suggested that head inducers can inhibit Wnt signaling during the early development of anterior brain structures. In zebrafish, for example, it was demonstrated that head defects in the headless mutant were caused by a mutation in T-cell factor 35. Loss of gene function in the headless mutant revealed that headless can repress Wnt target genes. These data provide the first genetic evidence that a component of the Wnt signaling pathway is essential in head/brain formation and patterning in vertebrate animals.
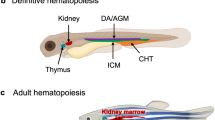
Zebrafish have been a tractable animal model for identifying developing neurons and the in vivo architecture of the brain, from neurogenesis at the early neural plate stage to the adult brain (Fig. 1). The zebrafish HuC homolog, which is 89% identical to the human HuC protein, is one of the earliest discovered markers of neuronal precursor cells in zebrafish, which are apparent during neurogenesis as early as the neural plate stage (Fig. 1a)26,27. Zebrafish are useful for studying the functional role of novel genes in neuronal development through directed expression studies of the zebrafish nervous system (Fig. 1b)28. In addition, recently, tissue-clearing technology has allowed visualization of neural networks in the whole brain of adult zebrafish (Fig. 1c). These molecular tools and technologies are useful for investigating phenotypic changes in zebrafish disease models of human developmental disorders.
a Detection of early neuronal precursor cells by whole-mount in situ hybridization with a pan-neuronal marker, huC, at the neural plate stage (10.5 h after fertilization). Unpublished data. b Immunostaining of axonal growth in the spinal cord of one-day-old zebrafish. Double-staining with anti-gicerin antibody and anti-HNK-1 antibody28. c Confocal image of myelin structure in an isolated adult zebrafish brain visualized by mbp promoter-driven membrane-tagged GFP, Tg(mbp:mEGFP). Arrows indicate the olfactory, optic, and otic nerves. Unpublished data.
The Genome-wide Association Study (GWAS) investigates a genomewide set of variants in human genetic diseases to identify the causative gene variant associated with a particular disease. GWAS data can be used to identify single-nucleotide polymorphisms (SNPs) and other variants in the genome associated with genetic disease29. In contrast to the identification of SNPs and variants, phenotypic abnormalities and haploinsufficiency of the various genes are derived from microdeletions or chromosomal translocation of different genomes30,31. For instance, Potocki-Shaffer syndrome (PSS) is a disorder that affects the development of bones, nerve cells in the brain, and other tissues due to the interstitial deletion of band p11.2 in chromosome 1132. Developmental disorders in PSS were investigated using phf21a-knockdown zebrafish, producing developmental abnormalities in the head, face, and jaw, in addition to increased neuronal apoptosis33. Another example of a disease studied by zebrafish models is Miles–Carpenter syndrome (MCS), in which syndromic X-linked intellectual disability is characterized by severe intellectual deficit, microcephaly, exotropia, distal muscle wasting, and low digital arches. By whole-exome sequencing of MCS families, ZC4H2 was identified as an MCS gene candidate. ZC4H2, a zinc-finger protein, is located in Xq11.2, and point mutations in ZC4H2 were found in MCS patients. Homozygous zc4h2-knockout zebrafish larvae showed motor hyperactivity, abnormal swimming, and continuous jaw movement. Motor hyperactivity was caused by a reduction in V2 GABAergic interneurons, arising from misspecification of neural progenitors in the brain and spinal cord of the zc4h2-knockout zebrafish34. The knockout animals also exhibited contractures of the pectoral fins and abnormal eye positioning, suggestive of exotropia, indicating that zebrafish disease models can be used to study the underlying cellular and molecular mechanisms of human developmental disorders.
Biomedical research II: mental disorders
Mental disorders, also called psychiatric disorders, are characterized by defective behavioral or mental patterns that cause significant distress to the subject. The International Classification of Diseases (ICD) published by the World Health Organization (WHO) is the international standard for classifying medical disease conditions. Over 450 different definitions of mental disorders are represented in the Diagnostic and Statistical Manual of Mental Disorders (DSM), the standard reference for psychiatry published by the American Psychiatric Association. Zebrafish are highly social animals that exhibit shoaling and schooling behaviors and are suitable for social behavioral tests in relation to mental disorders. Using mutagenesis screening, Kim and colleagues recently identified a novel chemokine-like gene family, samdori (sam), involved in mental disorders in zebrafish. Among the five sam family members, sam2 is specifically expressed in the habenular nuclei of the brain and is associated with intellectual disability and autism spectrum disorder35,36. Sam2-knockout animals (both zebrafish and mouse) showed defects in emotional responses, such as fear and anxiety, that are involved in anxiety-related disorders and/or autism35,36.
Additionally, by whole-exome sequencing, FAM50A was identified as the causative gene for Armfield X-linked intellectual disability (XLID) syndrome. XLID refers to forms of mental disorders with intellectual disability that are explicitly associated with X-linked recessive inheritance. Approximately 100 genes have been found to be involved in XLID syndrome37. XLID accounts for ~16% of all cases of intellectual disability in males, who are more likely to be affected than females. The biological activity of human FAM50A missense variants was functionally validated by rescue experiments in a zebrafish fam50a-knockout model38. Using the zebrafish disease model, it was recently found that Armfield XLID syndrome is a spliceosomopathy associated with aberrant mRNA processing during development38.
Biomedical research III: communication between the brain and other organs
Human puberty is a dynamic process that initiates the complex interactions of the hypothalamic-pituitary-gonadal axis (HPG axis), which refers to single endocrine glands as individual entities. The HPG axis plays a critical role in developing and regulating many of the body’s systems, particularly reproduction39. Gonadotropin-releasing hormone (GnRH), secreted by the hypothalamus in the brain, circulates through the anterior portion of the pituitary hypophyseal portal system and binds to receptors on the secretory cells of the adenohypophysis40. In response to GnRH stimulation, these cells produce luteinizing hormone and follicle-stimulating hormone, which circulate in the bloodstream41. Therefore, an adolescent develops into a mature adult with a body capable of sexual reproduction42. Kallmann syndrome (KS) is a genetic disorder known to prevent a person from starting or fully completing puberty. In a study showing that the WDR11 gene mutation is involved in KS pathogenicity, the zebrafish wdr11 gene was demonstrated to be expressed in the brain region, indicating a potential role for WDR11-EMX1 protein interaction43.
Additionally, acute inflammation is known to initiate regenerative response after traumatic injury in the adult zebrafish brain. The cysteinyl leukotriene receptor 1 (cysltr1)–leukotriene C4 (LTC4) pathway is required and sufficient for enhanced proliferation and neurogenesis44. LTC4, one of the ligands for CysLT1, binds to its receptor Cysltr1 expressed on radial glial cells in the zebrafish brain44. In a study by Kyritsis et al., cysltr1 was increasingly expressed on radial glial cells after traumatic brain injury, suggesting cross talk between components of the inflammatory response and the central nervous system during traumatic brain injury44.
The nicotinamide adenine dinucleotide phosphate (NADPH) oxidase (NOX) family is involved in the production of reactive oxygen species in response to various extracellular signals. The NOX family member dual oxidase (DUOX) was identified as thyroid NADPH oxidase. In humans, DUOX2 mutations were identified among children diagnosed with congenital hypothyroidism. Recently, it was demonstrated that, in addition to goitrous thyroid glands and growth retardation, defects in anxiety response and social interaction were found in duox-knockout zebrafish45. These results suggest that duox-knockout zebrafish could serve as an effective animal model for studies in thyroid development and related neurological diseases, including intellectual disability and autism.
A large percentage of children with ASD are known to have gastrointestinal problems, such as constipation, diarrhea, and abdominal pain. Recent studies on the brain-gut axis have also shown that interactions with host-associated microbial communities, either directly by microbial metabolites or indirectly via immune, metabolic or endocrine systems, can act as sources of environmental cues. Molecular signals from the gut provide environmental cues for communication between the gut and the brain during episodes related to anxiety, depression, cognition or autism spectrum disorder (ASD)46. Moreover, modulation of intrinsic signaling pathways and extrinsic cues in resident intestinal bacteria enhances the stability of β-catenin in intestinal epithelial cells, promoting cell proliferation47.
Biomedical research IV: metabolic disorders
Zebrafish an animal models for metabolic research
A high-calorie diet, a sedentary lifestyle, and a family history of metabolic disorders increase the prevalence of risk factors such as low HDL levels, high triglyceride levels, high blood glucose, high blood pressure, and abdominal obesity48. Such metabolic disorders may arise from an imbalance between nutritional intake and energy expenditure, leading to the development of serious illnesses, including diabetes, stroke, and fatty liver disease49.
In addition to general similarities with human metabolism, zebrafish metabolism also exhibits unique characteristics. Zebrafish embryos consume yolk for the first five days of development, after which they are fed for further growth to prevent them from undergoing fasting. The feeding-to-fasting transition at 5–6 days post fertilization (dpf) has been utilized to develop mechanistic insights into metabolic homeostasis upon energy deprivation50,51. Another unique feature of zebrafish is the composition and development of adipose tissue. As a poikilothermal animal, zebrafish do not seem to require brown adipose tissue, on which mammals do depend. Adipose development occurs late in development, with the first adipocyte being detected 8 dpf50.
Interestingly, late adipogenesis may also provide an experimental setting by which the role of adipose tissue in the pathogenesis of metabolic disorders can be investigated. Modeling metabolism to recapitulate human disorders can be appropriately established during the larval period. Similarly, metabolic disorders can be modeled in adults to explore phenotype references in the presence of all major metabolic organs. Many metabolic similarities and discrepancies between humans and zebrafish and the modeling of different types of metabolic diseases have been reviewed elsewhere52,53,54.
Zebrafish models for organelle biology research
Body metabolism is regulated by metabolic organelles, such as the endoplasmic reticulum (ER), mitochondria, peroxisomes, lipid droplets, and lysosomes. Whole-body metabolism is the sum of all metabolic activity of individual organs that originates from the metabolic function of individual cells. The function of subcellular organelles is critical for responding to environmental changes and regulating metabolic outputs to maintain metabolic homeostasis.
Zebrafish have served as an excellent model system to assess in vivo toxicity in response to treatment of a chemical of interest, and numerous studies have illustrated metabolic changes related to mitochondrial function upon chemical treatment55,56,57. After an initial study of mitochondrial activity and distribution in zebrafish oocytes reported in 1980, many reports regarding the mitochondrial genome and functional homologs of mitochondrial proteins in zebrafish were published in the late 1990s and early 2000s. More recently, zebrafish models have drawn extensive interest for use in testing a range of bioactive chemicals, including those that induce or disrupt development, improve disease conditions, or induce unfavorable side effects in daily human health or anticancer treatments58,59,60,61,62,63,64,65,66,67,68. In addition, the use of zebrafish as in vivo models for studying gene functions involved in metabolic activities has recently increased. Among the new molecular tools in developmental genetics, CRISPR/Cas9 is the most recent example of a reverse genetics technique, and mechanistic studies of the regulation of biogenesis, degradation and the quality maintenance of an organelle of interest have been conducted using zebrafish models69.
CRISPR/Cas9 is the most advanced gene editing system
Recent findings and the development of CRISPR/Cas9, evolutionary gene-editing machinery that originated from the defense system of bacteria that earned its developers the Nobel Prize in Chemistry in 2020. Highly efficient gene targeting made it possible to edit a gene of interest in any genome. Accordingly, studies utilizing CRISPR/Cas9 in zebrafish have rapidly increased. In particular, studies to elucidate the role of mitochondria in neutrophil motility70, tRNA biogenesis and the physiology of cardiomyocytes71,72, neuronal regeneration73, neurodegeneration in Parkinson’s disease74,75 and cellular metabolism regulation of mitochondrial abundance76,77 have been reported. Furthermore, studies illustrating the role of the endoplasmic (sarcoplasmic) reticulum included REEP5-gene knockout, which was used to elucidate the previously unknown regulation of ER/SR membrane protein organization and stress response in cardiac myocytes78. In addition, the demonstration of MCTP (multiple C2 domain proteins with two transmembrane regions) gene function acting as a novel ER calcium sensor was also reported76.
Moreover, molecular pathogenesis studies based on the analysis of genes, such as ATP13a79, NPC180,81, and GBA182 to understand Niewmann-Pick disease type C1 (NPC1) and other lysosomal storage diseases resulting from defective intracellular trafficking or lysosome function have been reported. Efforts have also been made to elucidate molecules and regulatory mechanisms leading to autophagosome formation, autolysosome formation, and autophagy83,84,85,86. Recently, a possible knock-in strategy to edit mitochondrial DNA and genomic DNA has been reported87, facilitating research on organelle function in metabolic diseases.
Transgenic approach to track organelle dynamics, abundance, and interaction
Mitochondria have long been foci due to their roles in bioenergetics and apoptosis, leading to a plethora of transgenic zebrafish. Several transgenic zebrafish, such as Mnx1:MITO-Kaede88, hspa8:MITO-YC289, and MLS-EGFP90, have been generated to mark mitochondria with fluorescent proteins GFP, YFP, Kaede, and yellow cameleon (YC), which are induced explicitly by a pan-expression promoter, an inducible heat shock promoter, a cell-type-specific promoter, or a combination of the GAL4-UAS system and are localized to the mitochondria using a mitochondria-targeting sequence91. One of the best examples of live mitochondrial imaging was illustrated in sensory axons of Rohon-Beard neurons, in which mitochondrial shape, dynamics, and transport were analyzed quantitatively92. A similar in vivo technique using zebrafish has since become popular to demonstrate the connection between mitochondrial behavior and neuronal health93,94. In addition to mitochondria, other organelles have been studied to reveal their roles during zebrafish development. For instance, a peroxisomal solute carrier, slc25a17, is involved in the maintenance of functional peroxisomes by showing substrate specificity towards coenzyme A95. To visualize peroxisomes in zebrafish embryos in vivo, the transgenic line Tg(Xla.Eef1a:RFP-SKL) was established and used under different metabolic conditions96. The use of double transgenic zebrafish allows simultaneous tracking of the dynamics of mitochondria and peroxisomes in vivo, as shown in Fig. 2a.
a Using transgenic zebrafish lines 5 dpf, mitochondria, Tg(Xla.Eef1a:MLS-EGFP), and peroxisomes, Tg(Xla. Eef1a:RFP-SKL), in the skin of the developing larva are visualized. b Motile cilia (green) in the hindbrain 4th ventricle are visualized with anti-acetylated tubulin antibody, and nuclei are shown in red. Unpublished data.
Another example is a transgenic line that marks the Golgi apparatus using the Golgi-Venus together with a cis-Golgi marker, GM-13097, to elucidate its role in dendrite specification of Purkinje cells. A trans-Golgi marker, GalT-GFP, was also established to reveal the dynamic localization of a connexin variant that influences cellular behavior98. A more systematic approach was applied to the study of secretory pathways, where a series of transgenic lines were generated based on different Rab proteins marking different types of endosomal vesicles99. A handful of transgenic lines were added to improve the identification of the cellular secretory pathways, and Lamp2-EGFP was used to mark lysosome-related vacuoles in the zebrafish notochord, GFP-CaaX (mem-GFP) was used to visualize the plasma membrane100, and NLS-mCherry or NLS-EGFP was used to the identify nucleus101,102. Another transgenic zebrafish used to mark the apoptotic cell membrane specifically, Annexin-Cy5, was also generated103. Moreover, transgenic zebrafish can be used to visualize transient and dynamic structures, with EGFP-LC3 used to monitor phagophore formation during autophagy104, Kif17-GFP105 used to analyze vesicles trafficking towards microtubule plus-ends, and EB1-GFP106 or EB2-GFP used to view microtubules growing in the plus-end. In combination with vital dyes, these transgenic zebrafish have been utilized extensively to advance our understanding of the dynamics of subcellular structures under physiological conditions and during pathological progression107.
Bioimaging tools that enable in vivo analysis
Advanced imaging tools that allow the examination of subcellular structures may facilitate the identification of previously unknown processes. These processes include communication between organelles upon membrane contact108, organelle biogenesis (peroxisome biogenesis109), organelle dynamics responding to an environmental cue110 and organelle trafficking along microtubules111. Notably, recent advances in microscopy have greatly enhanced the ability to observe cells in their native state and even monitor in vivo dynamics of organelles as well as ductal structure in the liver in zebrafish110,112. Motile cilia in the 4th ventricle of the hindbrain and bile duct of the developing liver can be visualized under confocal microscopy after specimens are immunostained with anti-acetylated tubulin (Fig. 2b) and with anti-cytokeratin 18 antibody (Fig. 3), respectively. High-speed, high-resolution, 3-dimensional in vivo imaging has enabled the dissection of dynamic intracellular processes and cellular behavior in response to different environments, which can enable the prediction of physiological conditions at the organism level. In this regard, a drug discovery platform based on organelle biology in zebrafish may play an essential role in the development of precision medicine and next-generation disease therapy.
Conclusion
In summary, the zebrafish is a very useful vertebrate animal model in biomedical research and drug discovery. In particular, with the aid of CRISPR-based-knockout technology and big data from next-generation DNA sequencing, functional validation of GWAS candidates in zebrafish is greatly enhancing the ability and accuracy of identifying causative genes and molecular mechanisms underlying the pathogenesis of human genetic diseases. These efforts are fundamental to the establishment of a platform for the future of precision medicine, providing new molecular targets for diagnostic and therapeutic strategies, especially those involving rare diseases.
References
Streisinger, G., Walker, C., Dower, N., Knauber, D. & Singer, F. Production of clones of homozygous diploid zebra fish (Brachydanio rerio). Nature 291, 293–296 (1981).
Grunwald, D. J. & Streisinger, G. Induction of recessive lethal and specific locus mutations in the zebrafish with ethyl nitrosourea. Genet. Res. 59, 103–116 (1992).
Kimmel, C. B. Genetics and early development of zebrafish. Trends Genet. 5, 283–288 (1989).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996).
Kim, C. H. et al. Repressor activity of Headless/Tcf3 is essential for vertebrate head formation. Nature 407, 913–916 (2000).
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat. Biotechnol. 26, 702–708 (2008).
Meng, X., Noyes, M. B., Zhu, L. J., Lawson, N. D. & Wolfe, S. A. Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat. Biotechnol. 26, 695–701 (2008).
Bedell, V. M. et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491, 114–118 (2012).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Varshney, G. K., Sood, R. & Burgess, S. M. Understanding and Editing the Zebrafish Genome. Adv. Genet. 92, 1–52 (2015).
Sung, Y. H. et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Res. 24, 125–131 (2014).
Postlethwait, J. H. et al. Vertebrate genome evolution and the zebrafish gene map. Nat. Genet. 18, 345–349 (1998).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Shin, J. T. & Fishman, M. C. From Zebrafish to human: modular medical models. Annu Rev. Genomics Hum. Genet. 3, 311–340 (2002).
Lieschke, G. J. & Currie, P. D. Animal models of human disease: zebrafish swim into view. Nat. Rev. Genet 8, 353–367 (2007).
Kwan, K. M. et al. The Tol2kit: a multisite gateway-based construction kit for Tol2 transposon transgenesis constructs. Dev. Dyn. 236, 3088–3099 (2007).
Halpern, M. E. et al. Gal4/UAS transgenic tools and their application to zebrafish. Zebrafish 5, 97–110 (2008).
Langenau, D. M. et al. Cre/lox-regulated transgenic zebrafish model with conditional myc-induced T cell acute lymphoblastic leukemia. Proc. Natl Acad. Sci. USA 102, 6068–6073 (2005).
Santoro, M. M. Zebrafish as a model to explore cell metabolism. Trends Endocrinol. Metab. 25, 546–554 (2014).
Nakayama, H. et al. Anti-obesity natural products tested in juvenile zebrafish obesogenic tests and mouse 3T3-L1 adipogenesis assays. Molecules 25, https://doi.org/10.3390/molecules25245840 (2020).
Misselbeck, K. et al. A network-based approach to identify deregulated pathways and drug effects in metabolic syndrome. Nat. Commun. 10, 5215 (2019).
Asaoka, Y., Terai, S., Sakaida, I. & Nishina, H. The expanding role of fish models in understanding non-alcoholic fatty liver disease. Dis. Model Mech. 6, 905–914 (2013).
Nakayama, H. et al. Novel anti-obesity properties of palmaria mollis in zebrafish and mouse models. Nutrients 10, https://doi.org/10.3390/nu10101401 (2018).
Kim, C. H. et al. Zebrafish elav/HuC homologue as a very early neuronal marker. Neurosci. Lett. 216, 109–112 (1996).
Itoh, M. et al. Mind bomb is a ubiquitin ligase that is essential for efficient activation of Notch signaling by Delta. Dev. Cell 4, 67–82 (2003).
Kim, C. H. et al. Neuron-specific expression of a chicken gicerin cDNA in transient transgenic zebrafish. Neurochem. Res. 21, 231–237 (1996).
Manolio, T. A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Koolen, D. A. et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat. Genet. 38, 999–1001 (2006).
Kalscheuer, V. M. et al. Disruption of the serine/threonine kinase 9 gene causes severe X-linked infantile spasms and mental retardation. Am. J. Hum. Genet. 72, 1401–1411 (2003).
Potocki, L. & Shaffer, L. G. Interstitial deletion of 11(p11.2p12): a newly described contiguous gene deletion syndrome involving the gene for hereditary multiple exostoses (EXT2). Am. J. Med. Genet. 62, 319–325 (1996).
Kim, H. G. et al. Translocations disrupting PHF21A in the Potocki-Shaffer-syndrome region are associated with intellectual disability and craniofacial anomalies. Am. J. Hum. Genet. 91, 56–72 (2012).
May, M. et al. ZC4H2, an XLID gene, is required for the generation of a specific subset of CNS interneurons. Hum. Mol. Genet. 24, 4848–4861 (2015).
Choi, J. H. et al. Targeted knockout of a chemokine-like gene increases anxiety and fear responses. Proc. Natl Acad. Sci. USA 115, E1041–E1050 (2018).
Kim, S. et al. Deficiency of a brain-specific chemokine-like molecule, SAM3, induces cardinal phenotypes of autism spectrum disorders in mice. Sci. Rep. 7, 16503 (2017).
Stevenson, R. E. & Schwartz, C. E. X-linked intellectual disability: unique vulnerability of the male genome. Dev. Disabil. Res Rev. 15, 361–368 (2009).
Lee, Y. R. et al. Mutations in FAM50A suggest that Armfield XLID syndrome is a spliceosomopathy. Nat. Commun. 11, 3698 (2020).
Millar, R. P. et al. Gonadotropin-releasing hormone receptors. Endocr. Rev. 25, 235–275 (2004).
Charlton, H. Hypothalamic control of anterior pituitary function: a history. J. Neuroendocrinol. 20, 641–646 (2008).
Vadakkadath Meethal, S. & Atwood, C. S. The role of hypothalamic-pituitary-gonadal hormones in the normal structure and functioning of the brain. Cell Mol. Life Sci. 62, 257–270 (2005).
Boehm, U. et al. Expert consensus document: European Consensus Statement on congenital hypogonadotropic hypogonadism-pathogenesis, diagnosis and treatment. Nat. Rev. Endocrinol. 11, 547–564 (2015).
Kim, H. G. et al. WDR11, a WD protein that interacts with transcription factor EMX1, is mutated in idiopathic hypogonadotropic hypogonadism and Kallmann syndrome. Am. J. Hum. Genet. 87, 465–479 (2010).
Kyritsis, N. et al. Acute inflammation initiates the regenerative response in the adult zebrafish brain. Science 338, 1353–1356 (2012).
Park, J. S. et al. Targeted knockout of duox causes defects in zebrafish growth, thyroid development, and social interaction. J. Genet Genomics 46, 101–104 (2019).
Sharon, G., Sampson, T. R., Geschwind, D. H. & Mazmanian, S. K. The central nervous system and the gut microbiome. Cell 167, 915–932 (2016).
Cheesman, S. E., Neal, J. T., Mittge, E., Seredick, B. M. & Guillemin, K. Epithelial cell proliferation in the developing zebrafish intestine is regulated by the Wnt pathway and microbial signaling via Myd88. Proc. Natl Acad. Sci. USA 108(Suppl 1), 4570–4577 (2011).
Park, W. Y., Choe, S. K., Park, J. & Um, J. Y. Black raspberry (Rubus coreanus Miquel) promotes browning of preadipocytes and inguinal white adipose tissue in cold-induced mice. Nutrients 11, https://doi.org/10.3390/nu11092164 (2019).
Jung, U. J. & Choi, M. S. Obesity and its metabolic complications: the role of adipokines and the relationship between obesity, inflammation, insulin resistance, dyslipidemia and nonalcoholic fatty liver disease. Int J. Mol. Sci. 15, 6184–6223 (2014).
Flynn, E. J. 3rd, Trent, C. M. & Rawls, J. F. Ontogeny and nutritional control of adipogenesis in zebrafish (Danio rerio). J. Lipid Res. 50, 1641–1652 (2009).
Minchin, J. E. N. & Rawls, J. F. A classification system for zebrafish adipose tissues. Dis. Model Mech. 10, 797–809 (2017).
Seth, A., Stemple, D. L. & Barroso, I. The emerging use of zebrafish to model metabolic disease. Dis. Model Mech. 6, 1080–1088 (2013).
Schlegel, A. & Gut, P. Metabolic insights from zebrafish genetics, physiology, and chemical biology. Cell Mol. Life Sci. 72, 2249–2260 (2015).
Salmi, T. M., Tan, V. W. T. & Cox, A. G. Dissecting metabolism using zebrafish models of disease. Biochem Soc. Trans. 47, 305–315 (2019).
Oulmi, Y. & Braunbeck, T. Toxicity of 4-chloroaniline in early life-stages of zebrafish (Brachydanio rerio): I. cytopathology of liver and kidney after microinjection. Arch. Environ. Contam Toxicol. 30, 390–402 (1996).
Braunbeck, T., Gorge, G., Storch, V. & Nagel, R. Hepatic steatosis in zebra fish (Brachydanio rerio) induced by long-term exposure to gamma-hexachlorocyclohexane. Ecotoxicol. Environ. Saf. 19, 355–374 (1990).
Peterman, E. M. et al. Neutralization of mitochondrial superoxide by superoxide dismutase 2 promotes bacterial clearance and regulates phagocyte numbers in zebrafish. Infect. Immun. 83, 430–440 (2015).
Hill, A. J., Teraoka, H., Heideman, W. & Peterson, R. E. Zebrafish as a model vertebrate for investigating chemical toxicity. Toxicol. Sci. 86, 6–19 (2005).
Basu, S. & Sachidanandan, C. Zebrafish: a multifaceted tool for chemical biologists. Chem. Rev. 113, 7952–7980 (2013).
Kanungo, J., Cuevas, E., Ali, S. F. & Paule, M. G. Zebrafish model in drug safety assessment. Curr. Pharm. Des. 20, 5416–5429 (2014).
McGrath, P. & Li, C. Q. Zebrafish: a predictive model for assessing drug-induced toxicity. Drug Disco. Today 13, 394–401 (2008).
Parng, C. In vivo zebrafish assays for toxicity testing. Curr. Opin. Drug Disco. Devel 8, 100–106 (2005).
McGrath, P. & Seng, W. L. Use of zebrafish apoptosis assays for preclinical drug discovery. Expert Opin. Drug Disco. 8, 1191–1202 (2013).
Baxendale, S., van Eeden, F. & Wilkinson, R. The Power of Zebrafish in Personalised Medicine. Adv. Exp. Med. Biol. 1007, 179–197 (2017).
Fernandez, I., Gavaia, P. J., Laize, V. & Cancela, M. L. Fish as a model to assess chemical toxicity in bone. Aquat. Toxicol. 194, 208–226 (2018).
Green, A. J. & Planchart, A. The neurological toxicity of heavy metals: a fish perspective. Comp. Biochem Physiol. C. Toxicol. Pharm. 208, 12–19 (2018).
Horzmann, K. A. & Freeman, J. L. Making waves: new developments in toxicology with the zebrafish. Toxicol. Sci. 163, 5–12 (2018).
Yoganantharjah, P. & Gibert, Y. The use of the zebrafish model to aid in drug discovery and target validation. Curr. Top. Med Chem. 17, 2041–2055 (2017).
Chen, Z., Berquez, M. & Luciani, A. Mitochondria, mitophagy, and metabolic disease: towards assembling the puzzle. Cell Stress 4, 147–150 (2020).
Zhou, W. et al. Neutrophil-specific knockout demonstrates a role for mitochondria in regulating neutrophil motility in zebrafish. Dis. Model Mech. 11, https://doi.org/10.1242/dmm.033027 (2018).
Chen, D. et al. Deletion of Gtpbp3 in zebrafish revealed the hypertrophic cardiomyopathy manifested by aberrant mitochondrial tRNA metabolism. Nucleic Acids Res. 47, 5341–5355 (2019).
Sasagawa, S. et al. Downregulation of GSTK1 Is a common mechanism underlying hypertrophic cardiomyopathy. Front Pharm. 7, 162 (2016).
Yang, L. Q., Chen, M., Ren, D. L. & Hu, B. Dual oxidase mutant retards mauthner-cell axon regeneration at an early stage via modulating mitochondrial dynamics in zebrafish. Neurosci. Bull. 36, 1500–1512 (2020).
Soman, S. K., Bazala, M., Keatinge, M., Bandmann, O. & Kuznicki, J. Restriction of mitochondrial calcium overload by mcu inactivation renders a neuroprotective effect in zebrafish models of Parkinson’s disease. Biol. Open 8, https://doi.org/10.1242/bio.044347 (2019).
Edson, A. J. et al. Dysregulation in the brain protein profile of zebrafish lacking the Parkinson’s disease-related protein DJ-1. Mol. Neurobiol. 56, 8306–8322 (2019).
Zhang, Q. et al. Deletion of Mtu1 (Trmu) in zebrafish revealed the essential role of tRNA modification in mitochondrial biogenesis and hearing function. Nucleic Acids Res 46, 10930–10945 (2018).
Munro, B., Horvath, R. & Muller, J. S. Nucleoside supplementation modulates mitochondrial DNA copy number in the dguok -/- zebrafish. Hum. Mol. Genet 28, 796–803 (2019).
Lee, S. H. et al. REEP5 depletion causes sarco-endoplasmic reticulum vacuolization and cardiac functional defects. Nat. Commun. 11, 965 (2020).
Nyuzuki, H. et al. Degeneration of dopaminergic neurons and impaired intracellular trafficking in Atp13a2 deficient zebrafish. IBRO Rep. 9, 1–8 (2020).
Zhang, T. & Peterson, R. T. Modeling lysosomal storage diseases in the zebrafish. Front Mol. Biosci. 7, 82 (2020).
Tseng, W. C. et al. Modeling Niemann-Pick disease type C1 in zebrafish: a robust platform for in vivo screening of candidate therapeutic compounds. Dis. Model Mech. 11, https://doi.org/10.1242/dmm.034165 (2018).
Lelieveld, L. T. et al. Role of beta-glucosidase 2 in aberrant glycosphingolipid metabolism: model of glucocerebrosidase deficiency in zebrafish. J. Lipid Res 60, 1851–1867 (2019).
Jewett, C. E., Appel, B. H. & Prekeris, R. The Rab11 effectors Fip5 and Fip1 regulate zebrafish intestinal development. Biol. Open 9, https://doi.org/10.1242/bio.055822 (2020).
Meneghetti, G. et al. The epg5 knockout zebrafish line: a model to study Vici syndrome. Autophagy 15, 1438–1454 (2019).
Fodor, E. et al. Methods to study autophagy in zebrafish. Methods Enzymol. 588, 467–496 (2017).
Sasaki, T. et al. Autolysosome biogenesis and developmental senescence are regulated by both Spns1 and v-ATPase. Autophagy 13, 386–403 (2017).
Bian, W. P. et al. Knock-in strategy for editing human and zebrafish mitochondrial DNA using mito-CRISPR/Cas9 system. ACS Synth. Biol. 8, 621–632 (2019).
Bergamin, G., Cieri, D., Vazza, G., Argenton, F. & Mostacciuolo, M. L. Zebrafish Tg(hb9:MTS-Kaede): a new in vivo tool for studying the axonal movement of mitochondria. Biochim Biophys. Acta 1860, 1247–1255 (2016).
Mizuno, H., Sassa, T., Higashijima, S., Okamoto, H. & Miyawaki, A. Transgenic zebrafish for ratiometric imaging of cytosolic and mitochondrial Ca2+ response in teleost embryo. Cell Calcium 54, 236–245 (2013).
Kim, M. J., Kang, K. H., Kim, C. H. & Choi, S. Y. Real-time imaging of mitochondria in transgenic zebrafish expressing mitochondrially targeted GFP. Biotechniques 45, 331–334 (2008).
Engerer, P. et al. Imaging subcellular structures in the living zebrafish embryo. J Vis Exp, e53456, https://doi.org/10.3791/53456 (2016).
Paquet, D., Plucinska, G. & Misgeld, T. In vivo imaging of mitochondria in intact zebrafish larvae. Methods Enzymol. 547, 151–164 (2014).
Mandal, A., Pinter, K. & Drerup, C. M. Analyzing neuronal mitochondria in vivo using fluorescent reporters in zebrafish. Front. Cell Dev. Biol. 6, 144 (2018).
Xu, Y., Chen, M., Hu, B., Huang, R. & Hu, B. In vivo Imaging of mitochondrial transport in single-axon regeneration of zebrafish mauthner cells. Front. Cell Neurosci. 11, 4 (2017).
Kim, Y. I. et al. Slc25a17 acts as a peroxisomal coenzyme A transporter and regulates multiorgan development in zebrafish. J. Cell Physiol. 235, 151–165 (2020).
Kim, D. et al. Fis1 depletion in osteoarthritis impairs chondrocyte survival and peroxisomal and lysosomal function. J. Mol. Med (Berl.) 94, 1373–1384 (2016).
Tanabe, K. et al. Atypical protein kinase C regulates primary dendrite specification of cerebellar Purkinje cells by localizing Golgi apparatus. J. Neurosci. 30, 16983–16992 (2010).
Gerhart, S. V. et al. The Cx43-like connexin protein Cx40.8 is differentially localized during fin ontogeny and fin regeneration. PLoS ONE 7, e31364 (2012).
Clark, B. S., Winter, M., Cohen, A. R. & Link, B. A. Generation of Rab-based transgenic lines for in vivo studies of endosome biology in zebrafish. Dev. Dyn. 240, 2452–2465 (2011).
Ellis, K., Bagwell, J. & Bagnat, M. Notochord vacuoles are lysosome-related organelles that function in axis and spine morphogenesis. J. Cell Biol. 200, 667–679 (2013).
Yokota, Y. et al. Endothelial Ca 2+ oscillations reflect VEGFR signaling-regulated angiogenic capacity in vivo. Elife 4, https://doi.org/10.7554/eLife.08817 (2015).
Yeo, S. Y., Kim, M., Kim, H. S., Huh, T. L. & Chitnis, A. B. Fluorescent protein expression driven by her4 regulatory elements reveals the spatiotemporal pattern of Notch signaling in the nervous system of zebrafish embryos. Dev. Biol. 301, 555–567 (2007).
Peri, F. & Nusslein-Volhard, C. Live imaging of neuronal degradation by microglia reveals a role for v0-ATPase a1 in phagosomal fusion in vivo. Cell 133, 916–927 (2008).
Cui, J., Sim, T. H., Gong, Z. & Shen, H. M. Generation of transgenic zebrafish with liver-specific expression of EGFP-Lc3: a new in vivo model for investigation of liver autophagy. Biochem Biophys. Res. Commun. 422, 268–273 (2012).
Bader, J. R., Kusik, B. W. & Besharse, J. C. Analysis of KIF17 distal tip trafficking in zebrafish cone photoreceptors. Vis. Res 75, 37–43 (2012).
Tran, L. D. et al. Dynamic microtubules at the vegetal cortex predict the embryonic axis in zebrafish. Development 139, 3644–3652 (2012).
Vacaru, A. M. et al. In vivo cell biology in zebrafish—providing insights into vertebrate development and disease. J. Cell Sci. 127, 485–495 (2014).
Ng, A. Q. E., Ng, A. Y. E. & Zhang, D. Plasma membrane furrows control plasticity of ER-PM Contacts. Cell Rep. 30, 1434–1446 e1437 (2020).
Sugiura, A., Mattie, S., Prudent, J. & McBride, H. M. Newly born peroxisomes are a hybrid of mitochondrial and ER-derived pre-peroxisomes. Nature 542, 251–254 (2017).
Liu, T. L. et al. Observing the cell in its native state: Imaging subcellular dynamics in multicellular organisms. Science 360, https://doi.org/10.1126/science.aaq1392 (2018).
Yildiz, A. Sorting out microtubule-based transport. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-020-00320-y (2020).
Choi, T. Y., Ninov, N., Stainier, D. Y. & Shin, D. Extensive conversion of hepatic biliary epithelial cells to hepatocytes after near total loss of hepatocytes in zebrafish. Gastroenterology 146, 776–788 (2014).
Acknowledgements
This work was supported by grants NRF-2020R1I1A3070817 (TYC), NRF-2018M3A9B8021980 (CHK), and MOF-20180430 (SKC). Zebrafish were obtained from the Zebrafish Center for Disease Modeling. Tissue-clearing reagents were kindly provided by Binaree, Inc.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choi, TY., Choi, TI., Lee, YR. et al. Zebrafish as an animal model for biomedical research. Exp Mol Med 53, 310–317 (2021). https://doi.org/10.1038/s12276-021-00571-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-021-00571-5
This article is cited by
-
Purification and Characterization of a Novel Anticoagulant Peptide from Protein Hydrolysate of Eupolyphaga sinensis Walker
International Journal of Peptide Research and Therapeutics (2024)
-
Cannabinoids and healthy ageing: the potential for extending healthspan and lifespan in preclinical models with an emphasis on Caenorhabditis elegans
GeroScience (2024)
-
Proteomic analysis unveils Gb3-independent alterations and mitochondrial dysfunction in a gla−/− zebrafish model of Fabry disease
Journal of Translational Medicine (2023)
-
Comparison of the effects of empagliflozin and sotagliflozin on a zebrafish model of diabetic heart failure with reduced ejection fraction
Experimental & Molecular Medicine (2023)
-
Evaluation of the efficacy of cystinosin supplementation through CTNS mRNA delivery in experimental models for cystinosis
Scientific Reports (2023)