Abstract
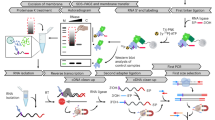
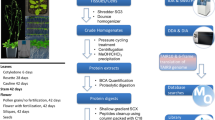
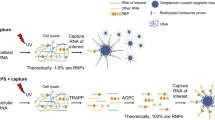
Tandem affinity purification coupled to mass spectrometry (TAP-MS) is one of the most advanced methods to characterize protein complexes in plants, giving a comprehensive view on the protein-protein interactions (PPIs) of a certain protein of interest (bait). The bait protein is fused to a double affinity tag, which consists of a protein G tag and a streptavidin-binding peptide separated by a very specific protease cleavage site, allowing highly specific protein complex isolation under near-physiological conditions. Implementation of this optimized TAP tag, combined with ultrasensitive MS, means that these experiments can be performed on small amounts (25 mg of total protein) of protein extracts from Arabidopsis cell suspension cultures. It is also possible to use this approach to isolate low abundant protein complexes from Arabidopsis seedlings, thus opening perspectives for the exploration of protein complexes in a plant developmental context. Next to protocols for efficient biomass generation of seedlings (∼7.5 months), we provide detailed protocols for TAP (1 d), and for sample preparation and liquid chromatography-tandem MS (LC-MS/MS; ∼5 d), either from Arabidopsis seedlings or from cell cultures. For the identification of specific co-purifying proteins, we use an extended protein database and filter against a list of nonspecific proteins on the basis of the occurrence of a co-purified protein among 543 TAP experiments. The value of the provided protocols is illustrated through numerous applications described in recent literature.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Braun, P., Aubourg, S., Van Leene, J., De Jaeger, G. & Lurin, C. Plant protein interactomes. Annu. Rev. Plant Biol. 64, 161–187 (2013).
Van Leene, J., Boruc, J., De Jaeger, G., Russinova, E. & De Veylder, L. A kaleidoscopic view of the Arabidopsiscore cell cycle interactome. Trends Plant Sci. 16, 141–150 (2011).
Arabidopsis Interactome Mapping Consortium. Evidence for network evolution in an Arabidopsis interactome map. Science 333, 601–607 (2011).
Backstrom, S., Elfving, N., Nilsson, R., Wingsle, G. & Bjorklund, S. Purification of a plant mediator from Arabidopsis thaliana identifies PFT1 as the Med25 subunit. Mol. Cell 26, 717–729 (2007).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Witte, C.P., Noel, L.D., Gielbert, J., Parker, J.E. & Romeis, T. Rapid one-step protein purification from plant material using the eight-amino acid StrepII epitope. Plant Mol. Biol. 55, 135–147 (2004).
Smaczniak, C. et al. Proteomics-based identification of low-abundance signaling and regulatory protein complexes in native plant tissues. Nat. Protoc. 7, 2144–2158 (2012).
Fröhlich, A. et al. Looking deep inside: detection of low-abundance proteins in leaf extracts of Arabidopsis and phloem exudates of pumpkin. Plant Physiol. 159, 902–914 (2012).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032 (1999).
Van Leene, J., Witters, E., Inzé, D. & De Jaeger, G. Boosting tandem affinity purification of plant protein complexes. Trends Plant Sci. 13, 517–520 (2008).
Puig, O. et al. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24, 218–229 (2001).
Kühner, S. et al. Proteome organization in a genome-reduced bacterium. Science 326, 1235–1240 (2009).
Gavin, A.-C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Krogan, N.J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Hutchins, J.R.A. et al. Systematic analysis of human protein complexes identifies chromosome segregation proteins. Science 328, 593–599 (2010).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005).
Van Leene, J. et al. Isolation of transcription factor complexes from Arabidopsis cell suspension cultures by tandem affinity purification. Methods Mol. Biol. 754, 195–218 (2011).
Van Leene, J. et al. A tandem affinity purification-based technology platform to study the cell cycle interactome in Arabidopsis thaliana. Mol. Cell. Proteomics 6, 1226–1238 (2007).
Takahashi, N. et al. The DNA replication checkpoint aids survival of plants deficient in the novel replisome factor ETG1. EMBO J. 27, 1840–1851 (2008).
Takahashi, N. et al. The MCM-binding protein ETG1 aids sister chromatid cohesion required for postreplicative homologous recombination repair. PLoS Genet. 6, e1000817 (2010).
Pauwels, L. et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464, 788–791 (2010).
Gadeyne, A. et al. The TPLATE adaptor complex drives clathrin-mediated endocytosis in plants. Cell 156, 691–704 (2014).
Spinner, L. et al. A protein phosphatase 2A complex spatially controls plant cell division. Nat. Commun. 4, 1863 (2013).
Van Leene, J. et al. Targeted interactomics reveals a complex core cell cycle machinery in Arabidopsis thaliana. Mol. Syst. Biol. 6, 397 (2010).
Eloy, N.B. et al. SAMBA, a plant-specific anaphase-promoting complex/cyclosome regulator is involved in early development and A-type cyclin stabilization. Proc. Natl. Acad. Sci. USA 109, 13853–13858 (2012).
Heyman, J. et al. Arabidopsis ULTRAVIOLET-B-INSENSITIVE4 maintains cell division activity by temporal inhibition of the anaphase-promoting complex/cyclosome. Plant Cell 23, 4394–4410 (2011).
Heyman, J. et al. ERF115 controls root quiescent center cell division and stem cell replenishment. Science 342, 860–863 (2013).
Bürckstümmer, T. et al. An efficient tandem affinity purification procedure for interaction proteomics in mammalian cells. Nat. Methods 3, 1013–1019 (2006).
Vercruyssen, L. et al. ANGUSTIFOLIA3 binds to SWI/SNF chromatin remodeling complexes to regulate transcription during Arabidopsis leaf development. Plant Cell 26, 210–229 (2014).
Lamesch, P. et al. The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 40, D1202–D1210 (2012).
Xu, X. et al. The tandem affinity purification method: an efficient system for protein complex purification and protein interaction identification. Protein Expr. Purif. 72, 149–156 (2010).
Li, Y. The tandem affinity purification technology: an overview. Biotechnol. Lett. 33, 1487–1499 (2011).
Kyriakakis, P., Tipping, M., Abed, L. & Veraksa, A. Tandem affinity purification in Drosophila—the advantages of the GS-TAP system. Fly 2, 229–235 (2008).
Ma, H. et al. A highly efficient multifunctional tandem affinity purification approach applicable to diverse organisms. Mol. Cell. Proteomics 11, 501–511 (2012).
Li, F. et al. Structure of the core editing complex (L-complex) involved in uridine insertion/deletion RNA editing in trypanosomatid mitochondria. Proc. Natl. Acad. Sci. USA 106, 12306–12310 (2009).
Sundberg, M. et al. The heteromultimeric debranching enzyme involved in starch synthesis in Arabidopsis requires both isoamylase1 and isoamylase2 subunits for complex stability and activity. PLoS ONE 8, e75223 (2013).
Heijde, M. et al. Constitutively active UVR8 photoreceptor variant in Arabidopsis. Proc. Natl. Acad. Sci. USA 110, 20326–20331 (2013).
Benhamed, M. et al. Genome-scale Arabidopsis promoter array identifies targets of the histone acetyltransferase GCN5. Plant J. 56, 493–504 (2008).
Domenichini, S. et al. Evidence for a role of Arabidopsis CDT1 proteins in gametophyte development and maintenance of genome integrity. Plant Cell 24, 2779–2791 (2012).
Van Damme, D. et al. Adaptin-like protein TPLATE and clathrin recruitment during plant somatic cytokinesis occurs via two distinct pathways. Proc. Natl. Acad. Sci. USA 108, 615–620 (2011).
Boudolf, V. et al. CDKB1;1 forms a functional complex with CYCA2;3 to suppress endocycle onset. Plant Physiol. 150, 1482–1493 (2009).
Van Aken, O. et al. Mitochondrial type-I prohibitins of Arabidopsis thaliana are required for supporting proficient meristem development. Plant J. 52, 850–864 (2007).
Cromer, L. et al. Centromeric cohesion is protected twice at meiosis, by SHUGOSHINs at anaphase I and by PATRONUS at interkinesis. Curr. Biol. 23, 2090–2099 (2013).
Berckmans, B. et al. Auxin-dependent cell cycle reactivation through transcriptional regulation of Arabidopsis E2Fa by lateral organ boundary proteins. Plant Cell 23, 3671–3683 (2011).
Nelissen, H. et al. Plant Elongator regulates auxin-related genes during RNA polymerase II transcription elongation. Proc. Natl. Acad. Sci. USA 107, 1678–1683 (2010).
Fernández-Calvo, P. et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 23, 701–715 (2011).
Geerinck, J., Pauwels, L., De Jaeger, G. & Goossens, A. Dissection of the one-megaDalton JAZ1 protein complex. Plant Signal. Behav. 5, 1039–1041 (2010).
Antoni, R. et al. PYRABACTIN RESISTANCE1-LIKE8 plays an important role for the regulation of abscisic acid signaling in root. Plant Physiol. 161, 931–941 (2013).
Di Rubbo, S. et al. The clathrin adaptor complex AP-2 mediates endocytosis of BRASSINOSTEROID INSENSITIVE1 in Arabidopsis. Plant Cell 25, 2986–2997 (2013).
Sauer, M. et al. MTV1 and MTV4 encode plant-specific ENTH and ARF GAP proteins that mediate clathrin-dependent trafficking of vacuolar cargo from the trans-Golgi network. Plant Cell 25, 2217–2235 (2013).
Nodzyński, T. et al. Retromer subunits VPS35A and VPS29 mediate prevacuolar compartment (PVC) function in Arabidopsis. Mol. Plant 6, 1849–1862 (2013).
Bassard, J.-E. et al. Protein-protein and protein-membrane associations in the lignin pathway. Plant Cell 24, 4465–4482 (2012).
Menges, M. & Murray, J.A.H. Synchronous Arabidopsissuspension cultures for analysis of cell-cycle gene activity. Plant J. 30, 203–212 (2002).
Verkest, A. et al. A generic tool for transcription factor target gene discovery in Arabidopsis cell suspension cultures based on tandem chromatin affinity purification. Plant Physiol. 164, 1122–1133 (2014).
Gibson, T.J., Seiler, M. & Veitia, R.A. The transience of transient overexpression. Nat. Methods 10, 715–721 (2013).
Facette, M.R., Shen, Z., Björnsdóttir, F.R., Briggs, S.P. & Smith, L.G. Parallel proteomic and phosphoproteomic analyses of successive stages of maize leaf development. Plant Cell 25, 2798–2812 (2013).
Wang, G., Wu, W.W., Zhang, Z., Masilamani, S. & Shen, R.-F. Decoy methods for assessing false positives and false discovery rates in shotgun proteomics. Anal. Chem. 81, 146–159 (2009).
Van Bel, M. et al. Dissecting plant genomes with the PLAZA comparative genomics platform. Plant Physiol. 158, 590–600 (2012).
Carbon, S. et al. AmiGO: online access to ontology and annotation data. Bioinformatics 25, 288–289 (2009).
Pardo, M. & Choudhary, J.S. Assignment of protein interactions from affinity purification/mass spectrometry data. J. Proteome Res. 11, 1462–1474 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Zhang, X., Henriques, R., Lin, S.-S., Niu, Q.-W. & Chua, N.-H. Agrobacterium-mediated transformation of Arabidopsis thalianausing the floral dip method. Nat. Protoc. 1, 641–646 (2006).
Clough, S.J. & Bent, A.F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Hubner, N.C. et al. Quantitative proteomics combined with BAC TransgeneOmics reveals in vivo protein interactions. J. Cell Biol. 189, 739–754 (2010).
Horiguchi, G., Kim, G.-T. & Tsukaya, H. The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J. 43, 68–78 (2005).
De Bodt, S., Hollunder, J., Nelissen, H., Meulemeester, N. & Inzé, D. CORNET 2.0: integrating plant coexpression, protein-protein interactions, regulatory interactions, gene associations and functional annotations. New Phytol. 195, 707–720 (2012).
Acknowledgements
J.V.L. is a postdoctoral Fellow of the Research Foundation-Flanders. We thank S. Ghorbani for practical support; A. Staes and J. Vandenbussche for support with mass spectrometry; G. Gonnelli for help with data analysis; K. Verleye, J.V. Driessche and N. Helderwert for general support; and A. Bleys for help in preparing the manuscript.
Author information
Authors and Affiliations
Contributions
J.V.L., D.E. and G.D.J. designed the research. B.C., N.D.W., G.P., E.V.D.S., L.V. and J.V.L. performed experiments. D.E., J.V.L., K.V., L.M. and G.D.J. analyzed the data. K.G. and E.W. provided protocols for LC-MS/MS analysis. M.D., A.V., K.V., K.G., L.M. and E.W. commented on the manuscript. J.V.L., D.E. and G.D.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Distribution of the average Normalized Spectral Abundance Factors (NSAF) of non-specific proteins.
Non-specific proteins were grouped in subsets by the number of different baitgroups the non-specific proteins were identified in.
Supplementary Figure 2 Identification of two non-specific proteins as genuine interactions.
Identification of two Actin-related proteins as genuine interactions using AN3 as bait, either with TAP on cell culture or on seedlings. In all TAP experiments where ARP4 or ARP7 were identified, the difference was calculated between the NSAF in that sample and the minimal NSAF value of 0.4% for low abundant non-specific proteins (See Supplementary Fig. 1).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 1 (PDF 703 kb)
Supplementary Table 2
Excel list of 760 nonspecific proteins, based on the occurrence of a co-purified protein among 543 TAP experiments using 115 different bait proteins, classified into 62 unrelated bait groups. In column three, the number of baitgroups in which a protein was identified, is shown. (XLS 110 kb)
Rights and permissions
About this article
Cite this article
Van Leene, J., Eeckhout, D., Cannoot, B. et al. An improved toolbox to unravel the plant cellular machinery by tandem affinity purification of Arabidopsis protein complexes. Nat Protoc 10, 169–187 (2015). https://doi.org/10.1038/nprot.2014.199
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.199
This article is cited by
-
Adaptor protein complex interaction map in Arabidopsis identifies P34 as a common stability regulator
Nature Plants (2023)
-
TraB family proteins are components of ER-mitochondrial contact sites and regulate ER-mitochondrial interactions and mitophagy
Nature Communications (2022)
-
NuA4 and H2A.Z control environmental responses and autotrophic growth in Arabidopsis
Nature Communications (2022)
-
Mapping of the plant SnRK1 kinase signalling network reveals a key regulatory role for the class II T6P synthase-like proteins
Nature Plants (2022)
-
The membrane-localized protein kinase MAP4K4/TOT3 regulates thermomorphogenesis
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.