Abstract
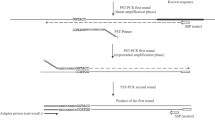
An enhanced universal fast walking (UFW) method adapted for the mapping of transposons is described. This protocol combines the original UFW method with the use of agarase to unravel composite nucleotide sequence, thereby forgoing molecular cloning steps and the use of restriction enzymes and ligases necessary in other available genome walking methods such as the prominent inverse PCR. The minuscule automatable chemistry of UFW is completed within one reaction vessel using a constant enzyme buffer, and the intrinsic DNA fingerprints, from which amplicons may be quantitatively recovered, offer quality assurance. The core steps of the protocol, spanning half a day or less, comprise first-strand synthesis, primer destruction, random-ended-primer annealing, distal branched-end repair, second-primer destruction, lariat formation and final amplification. Distinctively, no starting or intermediate templates are wasted during the reaction series, thus achieving yields comparable to direct PCR. Ultimate per-reaction walk-lengths are schematically illimitable and sequence-ready amplicons can be produced immediately from prevalent single-copy genomic walk origins. The core UFW protocol may be applied, as described here, to expedited transposon boundary retrieval, but is also applicable to general genome walking and cDNA walking, as well as viral and other insertional element mapping.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Myrick, K.V. & Gelbart, W.M. Universal fast walking for direct and versatile determination of flanking sequence. Gene 284, 125–131 (2002).
Huet, F. et al. A deletion-generator compound element allows deletion saturation analysis for genomewide phenotypic annotation. Proc. Natl. Acad. Sci. USA 99, 9948–9953 (2002).
Park, D.J., Renfree, M.B. & Marshall Graves, J.A. Universal fast walking applied to cDNA. Prep. Biochem. Biotechnol. 34, 123–133 (2004).
Arkhipova, I.R. & Meselson, M. Diverse DNA transposons in rotifers of the class Bdelloidea. Proc. Natl Acad. Sci. USA 102, 11781–11786 (2005).
Schon, I. & Arkhipova, I.R. Two families of non-LTR retrotransposons, Syrinx and Daphne, from the Darwinulid ostracod, Darwinula stevensoni. Gene 371, 296–307 (2006).
Kuo, S., Chang, W.J. & Landweber, L.F. Complex germline architecture: two genes intertwined on two loci. Mol. Biol. Evol. 23, 4–6 (2006).
Chang, W.J., Kuo, S. & Landweber, L.F. A new scrambled gene in the ciliate Uroleptus. Gene 368, 72–77 (2006).
Walser, J.C., Chen, B. & Feder, M.E. Heat-shock promoters: targets for evolution by P transposable elements in Drosophila . PLoS Genet. 2, e165 (2006).
Ochman, H., Gerber, A.S. & Hartl, D.L. Genetic applications of an inverse polymerase chain reaction. Genetics 120, 621–623 (1988).
Triglia, T., Peterson, M.G. & Kemp, D.J. A procedure for in vitro amplification of DNA segments that lie outside the boundaries of known sequences. Nucleic Acids Res. 16, 8186 (1988).
Mueller, P.R. & Wold, B. In vivo footprinting of a muscle specific enhancer by ligation mediated PCR. Science 246, 780–786 (1989).
Riley, J. et al. A novel, rapid method for the isolation of terminal sequences from yeast artificial chromosome (YAC) clones. Nucleic Acids Res. 18, 2887–2890 (1990).
Jones, D.H. & Winistorfer, S.C. Sequence specific generation of a DNA panhandle permits PCR amplification of unknown flanking DNA. Nucleic Acids Res. 20, 595–600 (1992).
Megonigal, M.D. et al. Panhandle PCR for cDNA: a rapid method for isolation of MLL fusion transcripts involving unknown partner genes. Proc. Natl Acad. Sci. USA 97, 9597–9602 (2000).
Puskas, L.G., Fartmann, B. & Bottka, S. Restricted PCR: amplification of an individual sequence flanked by a highly repetitive element from total human DNA. Nucleic Acids Res. 22, 3251–3252 (1994).
Liu, Y.G. & Whittier, R.F. Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 25, 674–681 (1995).
Schmidt, M. et al. Detection and direct genomic sequencing of multiple rare unknown flanking DNA in highly complex samples. Hum. Gene Ther. 12, 743 (2001).
Walser, J.C., Evgen'ev, M.B. & Feder, M.E. A genomic walking method for screening sequence length polymorphism. Mol. Ecol. Notes 6, 563–567 (2006).
Huang, A.M., Rehm, E.J. & Rubin, G.M. in Drosophila Protocols (eds. Sullivan, W., Ashburner, M. & Hawley, R.S.) 429–437 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2000).
Maniatis, T., Fritsch, E.F. & Sambrook, J. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, 1982).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Acknowledgements
The work in this study was supported by the NIGMS and the NHGRI.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
A patent, no. 6,929,914, "Method for accelerated genome walking and DNA fingerprinting", awarded by the United States Patent Office to owner Harvard University, with Kyl V. Myrick as inventor.
Rights and permissions
About this article
Cite this article
Myrick, K., Gelbart, W. A modified universal fast walking method for single-tube transposon mapping. Nat Protoc 2, 1556–1563 (2007). https://doi.org/10.1038/nprot.2007.223
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.223
This article is cited by
-
Cyclic Digestion and Ligation-Mediated PCR Used for Flanking Sequence Walking
Scientific Reports (2020)
-
Applications and challenges of next-generation sequencing in Brassica species
Planta (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.