Abstract
T cells reorganize their metabolic profiles after being activated, but the systemic metabolic effect of sustained activation of the immune system has remained unexplored. Here we report that augmented T cell responses in Pdcd1−/− mice, which lack the inhibitory receptor PD-1, induced a metabolic serum signature characterized by depletion of amino acids. We found that the depletion of amino acids in serum was due to the accumulation of amino acids in activated Pdcd1−/− T cells in the lymph nodes. A systemic decrease in tryptophan and tyrosine led to substantial deficiency in the neurotransmitters serotonin and dopamine in the brain, which resulted in behavioral changes dominated by anxiety-like behavior and exacerbated fear responses. Together these data indicate that excessive activation of T cells causes a systemic metabolomic shift with consequences that extend beyond the immune system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
O'Neill, L.A., Kishton, R.J. & Rathmell, J. A guide to immunometabolism for immunologists. Nat. Rev. Immunol. 16, 553–565 (2016).
Loftus, R.M. & Finlay, D.K. Immunometabolism: cellular metabolism turns immune regulator. J. Biol. Chem. 291, 1–10 (2016).
Okazaki, T., Chikuma, S., Iwai, Y., Fagarasan, S. & Honjo, T. A rheostat for immune responses: the unique properties of PD-1 and their advantages for clinical application. Nat. Immunol. 14, 1212–1218 (2013).
Good-Jacobson, K.L. et al. PD-1 regulates germinal center B cell survival and the formation and affinity of long-lived plasma cells. Nat. Immunol. 11, 535–542 (2010).
Kawamoto, S. et al. The inhibitory receptor PD-1 regulates IgA selection and bacterial composition in the gut. Science 336, 485–489 (2012).
Agata, Y. et al. Expression of the PD-1 antigen on the surface of stimulated mouse T and B lymphocytes. Int. Immunol. 8, 765–772 (1996).
Taylor, S. et al. PD-1 regulates KLRG1+ group 2 innate lymphoid cells. J. Exp. Med. 214, 1663–1678 (2017).
Hyde, R., Taylor, P.M. & Hundal, H.S. Amino acid transporters: roles in amino acid sensing and signalling in animal cells. Biochem. J. 373, 1–18 (2003).
Sinclair, L.V. et al. Control of amino-acid transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat. Immunol. 14, 500–508 (2013).
Iwai, Y., Terawaki, S. & Honjo, T. PD-1 blockade inhibits hematogenous spread of poorly immunogenic tumor cells by enhanced recruitment of effector T cells. Int. Immunol. 17, 133–144 (2005).
Hayaishi, O., Rothberg, S., Mehler, A.H. & Saito, Y. Studies on oxygenases; enzymatic formation of kynurenine from tryptophan. J. Biol. Chem. 229, 889–896 (1957).
Takikawa, O., Habara-Ohkubo, A. & Yoshida, R. Induction of indoleamine 2,3-dioxygenase in tumor cells transplanted into allogeneic mouse: interferon-γ is the inducer. Adv. Exp. Med. Biol. 294, 437–444 (1991).
Wikoff, W.R. et al. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl. Acad. Sci. USA 106, 3698–3703 (2009).
Yano, J.M. et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 161, 264–276 (2015).
Palmiter, R.D. Dopamine signaling in the dorsal striatum is essential for motivated behaviors: lessons from dopamine-deficient mice. Ann. NY Acad. Sci. 1129, 35–46 (2008).
Darvas, M., Wunsch, A.M., Gibbs, J.T. & Palmiter, R.D. Dopamine dependency for acquisition and performance of Pavlovian conditioned response. Proc. Natl. Acad. Sci. USA 111, 2764–2769 (2014).
Mosienko, V. et al. Exaggerated aggression and decreased anxiety in mice deficient in brain serotonin. Transl. Psychiatry 2, e122 (2012).
Bauer, E.P. Serotonin in fear conditioning processes. Behav. Brain Res. 277, 68–77 (2015).
Santos, J.M., Martinez, R.C. & Brandão, M.L. Effects of acute and subchronic treatments with fluoxetine and desipramine on the memory of fear in moderate and high-intensity contextual conditioning. Eur. J. Pharmacol. 542, 121–128 (2006).
Maki, Y. et al. Monoamine oxidase inhibitors reduce conditioned fear stress-induced freezing behavior in rats. Eur. J. Pharmacol. 406, 411–418 (2000).
Anthony, J. & Trevor, B.G.K. Marieke Kruidering-Hall. Antidepressants. in Katzung & Trevor′s Pharmacology: Examination & Board Review, 11e. Ch 30 (McGraw-Hill Education, 2015).
Carr, E.L. et al. Glutamine uptake and metabolism are coordinately regulated by ERK/MAPK during T lymphocyte activation. J. Immunol. 185, 1037–1044 (2010).
Chang, C.H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Patsoukis, N. et al. PD-1 alters T-cell metabolic reprogramming by inhibiting glycolysis and promoting lipolysis and fatty acid oxidation. Nat. Commun. 6, 6692 (2015).
Nishimura, H., Minato, N., Nakano, T. & Honjo, T. Immunological studies on PD-1 deficient mice: implication of PD-1 as a negative regulator for B cell responses. Int. Immunol. 10, 1563–1572 (1998).
Malissen, M. et al. Altered T cell development in mice with a targeted mutation of the CD3-epsilon gene. EMBO J. 14, 4641–4653 (1995).
Tagawa, Y., Sekikawa, K. & Iwakura, Y. Suppression of concanavalin A-induced hepatitis in IFN-γ−/− mice, but not in TNF-α−/− mice: role for IFN-γ in activating apoptosis of hepatocytes. J. Immunol. 159, 1418–1428 (1997).
Xia, J., Sinelnikov, I.V., Han, B. & Wishart, D.S. MetaboAnalyst 3.0--making metabolomics more meaningful. Nucleic Acids Res. 43, W1, W251–W257 (2015).
Moroji, T., Takahashi, K., Ogura, K., Toishi, T. & Arai, S. Rapid microwave fixation of rat brain. J Microwave Power. 12, 273–286 (1977).
Sugiura, Y., Honda, K., Kajimura, M. & Suematsu, M. Visualization and quantification of cerebral metabolic fluxes of glucose in awake mice. Proteomics 14, 829–838 (2014).
Oka, M. et al. Arl8b is required for lysosomal degradation of maternal proteins in the visceral yolk sac endoderm of mouse embryos. J. Cell. Sci. http://dx.doi:10.1242/jcs.200519 (2017).
Hu, S. et al. Targeted metabolomic analysis of head and neck cancer cells using high performance ion chromatography coupled with a Q exactive HF mass spectrometer. Anal. Chem. 87, 6371–6379 (2015).
Zhou, Z. Non-target impurity profiling of marketplace Cetirizine using high-resolution mass spectrometry and multivariate data analysis. Rapid Commun. Mass Spectrom. 30, 1941–1950 (2016).
Morikawa, T. et al. Hypoxic regulation of the cerebral microcirculation is mediated by a carbon monoxide-sensitive hydrogen sulfide pathway. Proc. Natl. Acad. Sci. USA 109, 1293–1298 (2012).
Shariatgorji, M. et al. Direct targeted quantitative molecular imaging of neurotransmitters in brain tissue sections. Neuron 84, 697–707 (2014).
Sugiyama, E., Masaki, N., Matsushita, S. & Setou, M. Ammonium Sulfate Improves Detection of Hydrophilic Quaternary Ammonium Compounds through Decreased Ion Suppression in Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. Anal. Chem. 87, 11176–11181 (2015).
Marshall, A.G., Hendrickson, C.L. & Jackson, G.S. Fourier transform ion cyclotron resonance mass spectrometry: a primer. Mass Spectrom. Rev. 17, 1–35 (1998).
Benveniste, H. & Hansen, A.J. Practical aspects of using microdialysis for determination of brain interstitial concentrations. In: Robinson, T.E., Jr., J.B.J. & Huston, J.P. (eds). Microdialysis in the neurosciences. Elsevier: New York, USA, 1991, pp 81–97.
Komada, M., Takao, K. & Miyakawa, T. Elevated plus maze for mice. J. Vis. Exp. 22, e1088 (2008).
Takao, K. & Miyakawa, T. Light/dark transition test for mice. J. Vis. Exp. 1, e104 (2006).
Miyakawa, T. et al. Conditional calcineurin knockout mice exhibit multiple abnormal behaviors related to schizophrenia. Proc. Natl. Acad. Sci. USA 100, 8987–8992 (2003).
Koshimizu, H. et al. Adenomatous polyposis coli heterozygous knockout mice display hypoactivity and age-dependent working memory deficits. Front. Behav. Neurosci. 5, 85 (2011).
Pattwell, S.S. et al. Altered fear learning across development in both mouse and human. Proc. Natl. Acad. Sci. USA 109, 16318–16323 (2012).
Hogquist, K.A. et al. T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27 (1994).
Barnden, M.J., Allison, J., Heath, W.R. & Carbone, F.R. Defective TCR expression in transgenic mice constructed using cDNA-based alpha- and beta-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Acknowledgements
We thank S. Yamamoto, S. Oonawa and Y. Doi for technical help; S. Kawamoto, T. Chaya, and T. Kozuka for the help with metabolome and brain dissection and preparations; S. Narumiya, T. Sato and K. Tanaka for discussions and suggestion; Y. Iwakura (Tokyo University of Science) and M. Kubo (IMS RIKEN) for Ifng−/− mice; B. Malissen (Centre d'Immunologie de Marseille-Luminy) for Cd3e−/− mice; and N. Lonberg (Bristol-Myers Squibb) for anti-PD-1. Supported by Japan Agency for Medical Research and Development–Core Research for Evolutional Science and Technology (14532135 to S.F.), Japan Agency for Medical Research and Development (145208 and 16770835 to T.H.) and the Cell Science Foundation (K.C).
Author information
Authors and Affiliations
Contributions
M.Mi. performed most of the in vivo experiments; B.Z. performed all in vitro experiments and behavioral studies at Kyoto University facility; Y.S., K.So. and K.H. performed all metabolome analyses; M.M.G. performed quantitative PCR and 5-HT staining and collaborated in writing the manuscript; Y.T. and S.I. performed behavioral studies at the RIKEN IMS facility; M.Mi and M.Ma. performed all germ-free and gnotobiotic experiments; A.V. collaborated in the writing and revision of the manuscript; K.C. contributed to tumour and anti-PD-1 blockade experiments; T.Hi. contributed expertise in behavioral studies; H.Q. contributed initial observations of mouse behavior; R.S. performed brain-region dissection; K.Su. contributed to OT-I and OT-II in vivo experiments; T.F. Y.I., F.M., M.S. and T.Ho. contributed expertise in metabolome and behavioral studies and conceptual design; and S.F. conceived of the conceptual design, analyzed the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Serum metabolome profiles of Pdcd1−/− mice and Pdcd1−/−Ifng−/− mice.
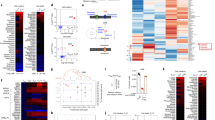
(a) Schematics of IFN-γ induced tryptophan degradation by the kynurenine pathway. (b) Principal component (PC) analysis of the serum metabolome of Pdcd1−/−Ifng−/− mice and Pdcd1−/− mice (n=12-14 mice per group). Each symbol represents an individual mouse. (c) GC-MS of proteinogenic amino acids (grouped as in Fig. 1c,d) in the serum of Pdcd1−/−Ifng−/− mice, presented relative to the mean values for Pdcd1−/− mice (n=12-14 mice per group). ND, not detected. NS, not significant (two-tailed unpaired Student’s t-test). Experiment performed once (b,c) (mean + s.e.m. in c).
Supplementary Figure 2 Non-targeted metabolome analysis of liver, lymph nodes, small intestine and colon.
(a) Non-targeted analyses by Orbitrap Fourier transform mass spectrometry (FT-MS) detected 8575 signals. Among them, 354 compounds were identified by both (i) formula prediction based on accurate m/z value and isotope patterns, as well as (ii) MS/MS structural validation (light blue dots in c). (b) Principal component (PC) analysis of the metabolome profile of the colon, liver, LNs and upper and lower segments of the small intestine (SI UP and SI Low, respectively) of Pdcd1−/− mice and wild-type controls using the above compounds (n = 5 per group). Each symbol represents the result for an individual mouse. (c) Volcano plots displaying probability values (log10) and fold-change (log2 values) show that LNs contained the largest number of altered metabolites between wild-type and Pdcd1−/− mice. Circles inside the green square represent significantly increased (P < 0.05, more than 2 times) metabolites in Pdcd1−/− mice. Circles inside the pink square were metabolites significantly decreased (less than half). Each symbol represents an individual compound. (d) Levels of indicated aromatic amino acids in the LNs of Pdcd1−/− mice compared to wild-type (WT) controls. ***P < 0.0005, ****P < 0.0001 (two-tailed unpaired Student’s t-test). Data are pooled from two experiments (b-d) (mean ± s.e.m. in d).
Supplementary Figure 3 Amino-acid profiles of tissues from wild-type and Pdcd1−/− mice.
Levels of proteinogenic amino acids (grouped as in Fig. 1c,d) in the spleen, small intestine, colon and liver isolated from wild-type and Pdcd1−/− mice relative to the mean of wild-type mice (n=5 mice per group). All measurements were by LC-MS. ND, not detected. NS, not significant; *P < 0.05, **P < 0.005 (two-tailed unpaired Student’s t-test). Data are pooled from two experiments (mean + s.e.m.).
Supplementary Figure 4 Tryptophan, tyrosine and related neurotransmitters in Pdcd1−/− mice.
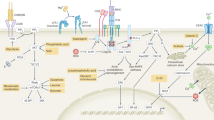
(a,b) Schematics of metabolic pathways for the synthesis of dopamine from tyrosine (a) and 5-HT from tryptophan (b). (c) Levels of metabolites measured by LC-MS or HPLC-ECD in the brain of wild-type and Pdcd1−/− mice. (d) Imaging mass spectrometry showing the distribution and relative levels of GABA in sagittal sections of brain from wild-type and Pdcd1−/− germ-free (GF) mice (left): GABA signal, 169215089 pixels (wild-type) and 126534111 pixels (Pdcd1−/−). Relative levels of indicated amino acids in the brain of wild-type and Pdcd1−/− GF mice were measured by LC-MS (right) (n=4-5 mice per group). (e) Microdialysis measurements of tryptophan and tyrosine in the prefrontal cortex levels in steady state (upper and lower left) and the levels of 5-HT upon high potassium challenge in wild-type and Pdcd1−/− mice (right). NS, not significant; *P < 0.05, **P < 0.005, ****P < 0.0001 (two-tailed unpaired Student’s t-test). Data are representative of two experiments (c), one experiment (d,e) (mean + s.e.m. in c,d).
Supplementary Figure 5 Metabolic and behavioral characteristics of Pdcd1−/− mice.
(a) Metabolic profiling of wild-type and Pdcd1−/− mice of indicated parameters plotted at 1 h intervals (n=4-5 mice per group). (b) Representative traces of wild-type and Pdcd1−/− mice in the dark-light transition test (left) and the indicated parameters results (right) (n=15 mice per group). (c) Forced-swim test. Distance travelled and percentage immobility (time spent without swimming) measured for the min 3 to min 6 of a 10 min session (n=12-13 mice per group). (d) Social-interaction test. The number and total duration of contacts and the mean duration per contact were shown (n=24 mice per group). (e) Hot-plate test showing the latency of hind paw withdrawal as a measure of pain sensitivity (n=24 mice per group). (f) Fear-extinction test. Percentage of freezing time on day 1 fear-conditioning in response to conditioned stimulus (CS) was shown in left panel. For fear-extinction training, mice received 5 presentations of the CS per day from day 2 to day 5. Percentages of freezing time in response to CS on day 2 and day 5 (middle two panels) were shown (n=8-9 mice per group). Fear-extinction indices were displayed (right panel) as differential freezing: freezing percentage = (CS1 + CS2 at day 2)-(CS4 + CS5 at day 5). (g) Representative traces of wild-type and Pdcd1−/− mice in the Y-maze test (left) and alternation rate between the arms of the Y maze (right) in wild-type and Pdcd1−/− mice (n=9-12 mice per group). NS, not significant; *P < 0.05, **P < 0.005 (two-tailed unpaired t-test (a-e and right panel in f) or ANOVA test (left and middle two panels in f)). Data are representative of at least two experiments with similar results (a-c,f,g) or one experiment (d,e) (means in a, mean + s.e.m. in b-e, right panel of f and g, and mean ± s.e.m. in left and middle two panels of f).
Supplementary Figure 6 Behavioral phenotyping of wild-type and Pdcd1-deficient littermates.
(a) Open field test of littermate wild-type and Pdcd1−/− mice. Locomotion parameters (measured as total distance travelled and total movement duration), total movement episodes and percentage of time spent in the center field were shown (n=12-14 per group). The littermate Pdcd1−/− mice which manifested freezing behaviour (n=4) in the center of platform were excluded from calculation. (b) Elevated plus-maze test of littermate wild-type and Pdcd1−/− mice. Total distance travelled, total entries and total time spent in the open arms, in the center of the maze and in the closed arms were shown (n=8-14 mice per group). (c) Percentage of freezing time on day 1 fear conditioning was shown in left panel (dark grey area: duration for presenting pairing of conditioned stimulus (CS) and unconditioned stimulus). One day later, percentage of freezing time of littermate wild-type and Pdcd1−/− mice were measured by contextual and cued test (right two panels; light grey area: duration for CS) (n=5-6 mice per group). (d) Open field test of wild-type and Pdcd1−/− mice after phenelzine treatment (n=6-7 mice per group). NS, not significant; *P < 0.05, **P < 0.005 (two-tailed unpaired t-test (a,b,d) or ANOVA test (c)). Data are representative of two experiments (a,b,c) or pooled from two experiments (d) (mean + s.e.m. in a,b,d and mean ± s.e.m. in c).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1832 kb)
Supplementary Table 1
Supplementary Excel Table of serum metabolites (XLS 1716 kb)
Rights and permissions
About this article
Cite this article
Miyajima, M., Zhang, B., Sugiura, Y. et al. Metabolic shift induced by systemic activation of T cells in PD-1-deficient mice perturbs brain monoamines and emotional behavior. Nat Immunol 18, 1342–1352 (2017). https://doi.org/10.1038/ni.3867
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3867
This article is cited by
-
The cancer-immune dialogue in the context of stress
Nature Reviews Immunology (2024)
-
Non-coding RNAs expression in SARS-CoV-2 infection: pathogenesis, clinical significance, and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Insights from a 30-year journey: function, regulation and therapeutic modulation of PD1
Nature Reviews Immunology (2023)
-
Amino acid catabolite markers for early prognostication of pneumonia in patients with COVID-19
Nature Communications (2023)
-
The effect of SSRIs on fear learning: a systematic review and meta-analysis
Psychopharmacology (2023)