Abstract
Gain-of-function IDH mutations are initiating events that define major clinical and prognostic classes of gliomas1,2. Mutant IDH protein produces a new onco-metabolite, 2-hydroxyglutarate, which interferes with iron-dependent hydroxylases, including the TET family of 5′-methylcytosine hydroxylases3,4,5,6,7. TET enzymes catalyse a key step in the removal of DNA methylation8,9. IDH mutant gliomas thus manifest a CpG island methylator phenotype (G-CIMP)10,11, although the functional importance of this altered epigenetic state remains unclear. Here we show that human IDH mutant gliomas exhibit hypermethylation at cohesin and CCCTC-binding factor (CTCF)-binding sites, compromising binding of this methylation-sensitive insulator protein. Reduced CTCF binding is associated with loss of insulation between topological domains and aberrant gene activation. We specifically demonstrate that loss of CTCF at a domain boundary permits a constitutive enhancer to interact aberrantly with the receptor tyrosine kinase gene PDGFRA, a prominent glioma oncogene. Treatment of IDH mutant gliomaspheres with a demethylating agent partially restores insulator function and downregulates PDGFRA. Conversely, CRISPR-mediated disruption of the CTCF motif in IDH wild-type gliomaspheres upregulates PDGFRA and increases proliferation. Our study suggests that IDH mutations promote gliomagenesis by disrupting chromosomal topology and allowing aberrant regulatory interactions that induce oncogene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008)
The Cancer Genome Atlas Research Network. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015)
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009)
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010)
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011)
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012)
Cairns, R. A. & Mak, T. W. Oncogenic isocitrate dehydrogenase mutations: mechanisms, models, and clinical opportunities. Cancer Dis. 3, 730–741 (2013)
Pastor, W. A., Aravind, L. & Rao, A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nature Rev. Mol. Cell Biol. 14, 341–356 (2013)
Kohli, R. M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013)
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17, 510–522 (2010)
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012)
Bickmore, W. A. & van Steensel, B. Genome architecture: domain organization of interphase chromosomes. Cell 152, 1270–1284 (2013)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014)
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012)
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015)
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 (2000)
Hark, A. T. et al. CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus. Nature 405, 486–489 (2000)
The GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015)
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl Acad. Sci. USA 111, 996–1001 (2014)
Sturm, D. et al. Paediatric and adult glioblastoma: multiform (epi)genomic culprits emerge. Nature Rev. Cancer 14, 92–107 (2014)
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013)
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1 . Cancer Cell 17, 98–110 (2010)
Wang, H. et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 22, 1680–1688 (2012)
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014)
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnol. 32, 347–355 (2014)
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome – biological and translational implications. Nature Rev. Cancer 11, 726–734 (2011)
Costello, J. F., Berger, M. S., Huang, H. S. & Cavenee, W. K. Silencing of p16/CDKN2 expression in human gliomas by methylation and chromatin condensation. Cancer Res. 56, 2405–2410 (1996)
Koivunen, P. et al. Transformation by the (R)-enantiomer of 2-hydroxyglutarate linked to EGLN activation. Nature 483, 484–488 (2012)
Chi, A. S. et al. Prospective, high-throughput molecular profiling of human gliomas. J. Neurooncol. 110, 89–98 (2012)
Rheinbay, E. et al. An aberrant transcription factor network essential for Wnt signaling and stem cell maintenance in glioblastoma. Cell Rep. 3, 1567–1579 (2013)
Suvà, M. L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014)
Wakimoto, H. et al. Maintenance of primary tumor phenotype and genotype in glioblastoma stem cells. Neuro Oncol. 14, 132–144 (2012)
Luchman, H. A. et al. An in vivo patient-derived model of endogenous IDH1-mutant glioma. Neuro Oncol. 14, 184–191 (2012)
Lai, R. K. et al. Genome-wide methylation analyses in glioblastoma multiforme. PLoS ONE 9, e89376 (2014)
Abyzov, A., Urban, A. E., Snyder, M. & Gerstein, M. CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res. 21, 974–984 (2011)
Chudnovsky, Y. et al. ZFHX4 interacts with the NuRD core member CHD4 and regulates the glioblastoma tumor-initiating cell state. Cell Rep. 6, 313–324 (2014)
Bao, S. et al. Targeting cancer stem cells through L1CAM suppresses glioma growth. Cancer Res. 68, 6043–6048 (2008)
Sandelin, A., Alkema, W., Engstrom, P., Wasserman, W. W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004)
de Laat, W. & Dekker, J. 3C-based technologies to study the shape of the genome. Methods 58, 189–191 (2012)
Hagège, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nature Protocols 2, 1722–1733 (2007)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008)
Acknowledgements
We thank J. Kim, the MGH Neuro Oncology Tissue Repository, and the MGH Pathology Flow Cytometry Core for assistance with clinical samples and analysis, and E. Lander and W. Kaelin for discussions. W.A.F. is supported by a basic research fellowship from the American Brain Tumor Association. B.B.L. is supported by a Jane Coffin Childs fellowship. B.E.B. is an American Cancer Society Research Professor. This research was supported by funds from Howard Hughes Medical Institute, the National Brain Tumor Society and the National Human Genome Research Institute.
Author information
Authors and Affiliations
Contributions
Conception and experimental design: W.A.F., Y.D., B.B.L., S.M.G., M.L.S. and B.E.B. Methodology and data acquisition: W.A.F., Y.D., B.B.L., S.M.G., A.S.V., A.O.S.-R., M.L.S. and B.E.B. Analysis and interpretation of data: W.A.F., Y.D. and B.E.B. Manuscript writing: W.A.F., Y.D. and B.E.B. W.A.F. and Y.D. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DNA methylation and CTCF binding at deregulated boundaries.
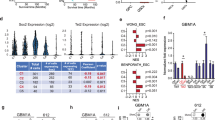
a, Box plots show DNA methylation levels over CTCF sites (200-bp window centred on the peak) within boundaries predicted by gene pair correlation analysis to be disrupted. All CTCF sites located within a 1-kb window centred on a disrupted boundary were considered. Methylation levels were determined from whole-genome bisulfite data for three IDH mutant (red labels) and three IDH wild-type (black labels) tumours. b, Bars show average normalized ChIP-seq signal over all CTCF sites located inside a 1-kb window centred on a disrupted boundary.
Extended Data Figure 2 Expression of Fip1l1 in mouse brain cells and survival effects of PDGFRA and FIP1L1.
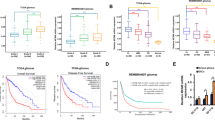
a, Expression of Fip1l1 in a published data set for isolated mouse brain cell types44. b, Kaplan–Meier plot based on TCGA data23 indicates that combined FIP1L1 and PDGFRA expression is a negative prognostic factor in IDH1 mutant lower-grade gliomas. Multivariate analysis including the known prognostic factor 1p/19q deletion diminished this effect into non-significance, suggesting that other predictors of survival may also have a role in this model.
Extended Data Figure 3 CTCF-anchored loop in the PDGFRA region.
a, Schematic depiction of a HiC interaction signature of a CTCF-anchored loop domain, compared to an ordinary domain, as described previously15. CTCF-anchored loop domains are characterized by an increased interaction score at the apex of the domain, representing a CTCF–CTCF dimeric interaction. b, IMR90 HiC contact matrix for the PDGFRA/FIP1L1 locus, as presented in Fig. 3a. Solid circle indicates CTCF dimer interaction point; dashed circles indicate lack of CTCF dimeric anchor signature. c, IMR90 HiC contact matrix as in b, but with an expanded heatmap scale, more clearly conveys the CTCF-anchored loop that insulates PDGFRA. d, e, HiC contact matrix for GM12878 cells for the same region confirms a single CTCF-anchored loop (solid circle) between PDGFRA and FIP1L1. These data support the significance of this specific boundary in locus topology and PDGFRA insulation.
Extended Data Figure 4 Characterization of the FIP1L1 enhancer.
a, H3K27ac ChIP-seq track for GSC6 gliomaspheres reveals strong enrichment over the FIP1L1 enhancer. CTCF ChIP-seq track reveals location of the boundary element insulator (as in Fig. 3a). FIP1L1 enhancer (i) and promoter (ii) are indicated. b, H3K27ac ChIP-seq tracks for IDH mutant and wild-type gliomaspheres and glioma specimens reveal enrichment over the FIP1L1 enhancer. c, ChIP-seq tracks for glioma master transcription factors and other histone modifications support the enhancer identity of the element (H3K27ac, H3K4me1, SOX2, OLIG2; lacks H3K4me3, lacks H3K27me3). By contrast, the FIP1L1 promoter has a distinct ‘promoter-like’ chromatin state.
Extended Data Figure 5 Interaction of the FIP1L1 enhancer with nearby promoters and PDGFRA quantified by reciprocal 3C.
Top, the H3K27ac, CTCF and genetic architecture of the FIP1L1/PDGFRA locus is indicated, highlighting the 3C strategy. Bottom, plots indicate the interaction signal of the indicated sites (black lines) with the common enhancer primer. The FIP1L1 enhancer interacts with local promoters in wild-type and mutant tumours and models. In IDH wild-type gliomas, it shows essentially no interaction with the PDGFRA promoter. In IDH mutant gliomas, it interacts with the PDGFRA promoter with comparable strength to the local interactions, despite the much larger intervening distance (900 kb). Error bars reflect s.d.
Extended Data Figure 6 Crenolanib reverses the increased growth of PDGFRA insulator disrupted cells.
Insulator CRISPR-infected gliomaspheres exhibit a roughly twofold increase in proliferation rate, compared to control sgRNA-infected gliomaspheres. This proliferative advantage is eliminated by treatment with the PDGFRα inhibitor crenolanib. Crenolanib and dasatinib both inhibit PDGFRα, but their other targets are non-overlapping. Hence, this sensitivity provides further support that PDGFRA induction drives the increased proliferation of the insulator CRISPR gliomaspheres. Error bars reflect s.d.
Extended Data Figure 7 Signature of boundary deregulation in IDH mutant gliomas is robust.
Volcano plot depicts the significance (y axis) of gene pairs that are either more or less correlated in IDH mutant than IDH wild-type gliomas. This plot was generated by repeating the analysis in the main text and shown in Fig. 1f, except that here the statistics were performed using only the 14,055 genes expressed at >1 TPM in at least half of the samples. This indicates that the boundary deregulation signature in IDH mutant gliomas is not sensitive to noise from lowly expressed genes.
Supplementary information
Supplementary Table
This file contains a list of deregulated boundaries and genes. The table lists domains whose boundaries are predicted to be lost based on cross-boundary gene pairs that gain correlation and intra-domain gene pairs that lose correlation. All supporting gene pairs (FDR < 1%) are listed along with their significance. The overall significance of each domain is also shown. Genes that are up-regulated by at least 2-fold in IDH mutant gliomas, relative to the median of the wild-type, are indicated. For each domain, the coordinates of boundaries predicted to be lost are also listed. (XLSX 23 kb)
Rights and permissions
About this article
Cite this article
Flavahan, W., Drier, Y., Liau, B. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016). https://doi.org/10.1038/nature16490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16490
This article is cited by
-
Advances in the multimodal analysis of the 3D chromatin structure and gene regulation
Experimental & Molecular Medicine (2024)
-
The potential of epigenetic therapy to target the 3D epigenome in endocrine-resistant breast cancer
Nature Structural & Molecular Biology (2024)
-
Defining super-enhancers by highly ranked histone H4 multi-acetylation levels identifies transcription factors associated with glioblastoma stem-like properties
BMC Genomics (2023)
-
Stalled oligodendrocyte differentiation in IDH-mutant gliomas
Genome Medicine (2023)
-
Co-localization of clusters of TCR-regulated genes with TAD rearrangements
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.