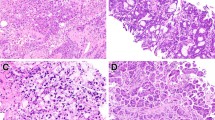
Abstract
In colorectal cancer, KRAS (exons 2, 3, and 4) and NRAS (exons 2, 3, and 4) mutations are associated with resistance to antiepidermal growth factor receptor monoclonal antibodies, and BRAF mutation is a molecular marker of poor prognosis. KRAS exon 2 and BRAF-mutated colorectal cancers have well-known distinct clinicopathological characteristics. Comparison of tumors with different RAS status (exons 2, 3, and 4 of KRAS and NRAS) based on their clinicopathological characteristics has never been established. All colorectal cancer patients with RAS and BRAF testing from 2011 to 2015 were included in this observational retrospective study. Patient and tumor characteristics were collected and correlation with RAS and BRAF status was evaluated. A total of 1735 patients with colorectal cancer were included. RAS-mutated colorectal cancers (n=1002), compared with RAS wild-type colorectal cancers (n=733), were significantly associated with male gender, classical adenocarcinoma subtype, well/moderately differentiated tumors, and microsatellite stable phenotype. KRAS codon 13-mutated colorectal cancers (n=171), compared with RAS wild-type colorectal cancers, more frequently presented classical adenocarcinoma subtype and microsatellite stable phenotype. In comparison with other RAS mutations, KRAS exon 3-mutated colorectal cancers (n=23) were associated with mucinous/rare histological subtypes and, most likely to located in the rectum. KRAS exon 4-mutated colorectal cancers (n=33) were more frequently associated with mucinous/rare histological subtypes. There was no significant association between NRAS mutation (n=37) and clinicopathological features. Colorectal cancers are associated with different clinicopathological features according to the type of RAS mutation. Consequently, these particular characteristics must be considered when assessing the prognostic value of RAS status in colorectal cancer.
Similar content being viewed by others
Main
Colorectal cancer is the third most common cancer in the world,1 with nearly 1.4 million new cases diagnosed each year. An estimated, 700 000 people die from colorectal cancer worldwide each year. The 5-year overall survival rate is ~59%.2 About 45% of colorectal cancers are associated with activating mutations of the KRAS gene. Mutations are located mostly in codons 12 (~30%) and 13 (~8%) of exon 2.3, 4 These somatic mutations result in a constitutive activation of the epidermal growth factor receptor (EGFR) pathway and, therefore, confer resistance to anti-EGFR therapy. Previous studies have shown that KRAS mutations in exon 2 are associated with no clinical benefit from anti-EGFR monoclonal antibodies in metastatic colorectal cancers.5, 6 KRAS mutation has been also associated with worse disease-free survival in adjuvant setting.7, 8 Recently, other mutations in KRAS (exons 3 and 4) and NRAS genes (exons 2, 3, and 4) have been reported to be associated with resistance to anti-EGFR monoclonal antibodies too.9 Anti-EGFR monoclonal antibodies, panitumumab and cetuximab, were consequently restricted to patients with metastatic colorectal cancers harboring a complete wild-type RAS genotype (KRAS and NRAS exons 2, 3, and 4 testing).9, 10, 11, 12
Approximately 10% of colorectal cancer are BRAF-mutated.13 BRAF mutation in colorectal cancer is associated with poor prognosis, especially in metastatic colorectal cancer.14 In addition, mutations in RAS and BRAF genes are mutually exclusive.15, 16, 17, 18, 19 Colorectal cancers with deficient mismatch repair system account for ~15% of all colorectal cancers. Deficient mismatch repair system is due to germline mutation in a mismatch repair gene (Lynch syndrome) or more commonly due to epigenetic inactivation of the MLH1 gene (sporadic cases). As compared with proficient mismatch repair colorectal cancers, deficient mismatch repair colorectal cancers are associated with good prognosis in non-metastatic setting20 and high disease control with drugs that target the immune checkpoints in metastatic setting.21
Recent studies have shown that KRAS exon 2 and BRAF-mutated colorectal cancers have distinct clinical and pathological characteristics. KRAS exon 2-mutated colorectal cancers occur more frequently in older patients, with a predominance of male gender, and are frequently located in the proximal colon compared with KRAS wild-type colorectal cancers.22, 23 High frequency of KRAS-mutation has been reported especially in the cecum.22, 24 Nevertheless, limited number of patients is a major weakness of those studies, leading to inconsistent conclusions. For instance, some studies have mentioned an association between KRAS mutations and mucinous differentiation, whereas others have not.22, 23 Moreover, correlation between KRAS mutation subtypes and clinicopathological characteristics was never clearly appraised. Gonsalves et al. showed that KRAS codon 13 mutations (p.Gly13Asp) are associated with deficient mismatch repair status and poor histologic grade.15 BRAF mutations are significantly associated with advanced age, female gender, poor histologic grade, mucinous differentiation, proximal colon tumor site, and deficient mismatch repair status.15, 25
Up until now, few studies have evaluated the possible correlation between clinicopathological features and complete RAS status (exons 2, 3, and 4 of KRAS and NRAS). A small series of 264 metastatic colorectal cancers in Japanese patients reported some association between KRAS exon 2 mutations and others RAS mutations with the rectal tumor site.25 The aim of this study was to identify new distinct subsets of colorectal cancers based on clinicopathological features and complete RAS and BRAF genotype of a large cohort of 1735 colorectal cancer patients.
Materials and methods
Population
All patients harboring colorectal cancer with complete RAS testing, performed at the Molecular Cancer Genetics Platform of Poitiers (French National Cancer Institute (INCa)) from January 2011 to April 2015, were included in the study (Figure 1). Since 2006, INCa has been supporting a national network of 28 hospital molecular genetics platforms throughout France, offering to patients an access to all essential molecular genetics testing for cancers. Overall, 2813 consecutive colorectal cancer cases with a molecular testing were reviewed to implement the cohort. Tumors with multiple testing were included only once (n=36). Tumors with incomplete RAS/BRAF status (n=834) were excluded. Indeed, before December 2013 there was no complete RAS testing (only KRAS exon 2 mutations were analyzed). It became systematic for every colorectal cancer after December 2013. Other rare histological types than adenocarcinoma were also excluded (n=6). Tumors with both RAS and BRAF mutations, or with two different RAS mutations, were also excluded (n=7). Finally, 1735 tumors with complete RAS and BRAF testing were included in the study. Results of microsatellite instability analysis were also recorded (n=700).
Molecular Analyses
Tumoral DNA was extracted from paraffin-embedded tumor sections using KAPA Express Extract Kits (KAPA Biosystems, CliniSciences, Nanterre, France) and GeneAmp PCR System 9700 Thermal Cycler (Applied Biosystems, Waltham, MA, USA).
According to the manufacturer’s instructions, RAS and BRAF molecular testing was performed using polymerase chain reaction (PCR) amplification and pyrosequencing technology Q24 PyroMark system (Qiagen, Hilden, Germany) with CE-IVD-certified allele-specific (Qiagen) or homemade primers.26 Mutational status of KRAS (exon 2: codons 12 and 13, exon 3: codons 59 and 61, and exon 4: codons 117 and 146), NRAS (exon 2: codons 12 and 13, exon 3: codons 59 and 61, and exon 4: codons 117 and 146), and BRAF (exon 15: codon 600) was analyzed.
Mismatch repair system status was determined with Microsatellite Instability Analysis System (Promega, Charbonnières-les-Bains, France) and multiplex PCR fluorescence assay using a panel of five mononucleotide repeat markers (BAT-25, BAT-26, NR-21, NR-24, and MONO-27) known to be monomorphic in Caucasian population. Samples were analyzed by capillary electrophoresis and data were deciphered using fragment analysis software (GeneMapper Software, Applied Biosystems). Colorectal cancers with two or more unstable loci were classified microsatellite-unstable and, thus, with deficient mismatch repair system.
Pathological Features
The following pathological features were analyzed using a pathological report database: histologic subtype (classical adenocarcinoma, mucinous adenocarcinoma, signet-ring cell adenocarcinoma, medullary adenocarcinoma, adenosquamous carcinoma, and undifferentiated carcinoma), histologic grade (well, moderately, or poorly differentiated), tumor site (proximal colon, transverse colon, distal colon, and rectum), vascular invasion (lymphatic and/or venous invasion/embols), and perineural invasion. Primary tumors located in the cecum and ascending colon were defined as proximal tumors, whereas tumors located in the splenic flexure, descending colon, and sigmoid colon were defined as distal tumors.27 Mucinous colorectal adenocarcinoma was defined by at least 50% of the tumor volume composed of extracellular mucin.28, 29
Definition of RAS Status and Different Subgroups
RAS and BRAF mutations are considered to be mutually exclusive (at 99%).15, 16, 17, 18, 19 Consequently, tumors with a RAS mutation were considered BRAF wild type, if no BRAF testing was performed. In the same way, tumors with BRAF mutation were considered RAS wild type, if RAS testing was incomplete.
Then, different groups of colorectal cancers were set, whether they presented RAS mutation or BRAF mutation, or none (super wild-type colorectal cancers), and they were compared with each other. Subtypes of KRAS mutants (KRAS exon 2: KRAS codon 12 and codon 13, KRAS exon 3 and KRAS exon 4 mutants) were also compared with each other.
Statistical Analyses
Clinical, pathological, and molecular variables collected at baseline were described as means and s.d.’s for quantitative variables and percentages for qualitative variables. Associations between mutational status and patients or tumor characteristics were assessed using the χ2-test (or Fisher’s exact test if appropriate) for qualitative variables and using Student's t-test for continuous variables. For all analyses an adjustment for multiple testing has been performed using Bonferroni correction. Statistical analyses were performed using the Statview software (Statview for Windows, SAS Institut, version 5.0).
Results
Patient and Tumor Characteristics
RAS and BRAF genotype was available for 1735 patients. The median age was 70.5 years and 57.9% were of male gender (Table 1). Histologic subtypes were 88.7% of classical adenocarcinoma, 10.1% of mucinous adenocarcinoma, and 1.2% of other adenocarcinoma subtypes. The majority of colorectal cancers were moderately differentiated (55.2%). Tumor sites were proximal, transverse, distal colon, and rectum at 37.0%, 4.3%, 39.2%, and 19.5%, respectively.
There was 55.6% of KRAS mutation (n=965/1735), 2.1% of NRAS mutation (n=37/1735), 15.5% of BRAF mutation (n=269/1735), 26.7% of super wild-type colorectal cancers (n=464/1735), and 13.4% of deficient mismatch repair colorectal cancers (n=94/700).
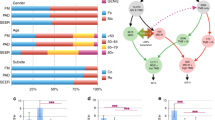
Comparison of subgroups of super wild-type, KRAS, NRAS, and BRAF mutants showed significant association with several features, namely age, gender, histologic subtype, histologic grade, tumor site, and microsatellite instability (all P<0.0001; Table 2). To highlight differences between mutational status, a 2 by 2 comparison was then conducted (mutated versus wild type).
Association Between RAS/Frequent KRAS Mutations and Clinicopathological Features
Among the 1735 colorectal cancers, 733 were RAS wild-type (42.2%) and 1002 were RAS mutants (57.8%). There was no age difference among patients according to RAS status, 70.4 years for RAS-mutated colorectal cancers, and 70.6 years for RAS wild-type colorectal cancers (Table 1). Compared with RAS wild-type colorectal cancers, RAS-mutated colo-rectal cancers were statistically associated with male gender (60.7% versus 54.0%; P=0.006), classical adenocarcinoma subtype (90.6% versus 86.1%; P=0.005), well/moderately differentiated tumors (93.4% versus 83.5%; P<0.0001), and microsatellite-stable phenotype (95.3% versus 75.8%; P<0.0001).
Secondly, we looked at the association between RAS mutation subtypes and clinicopathological features. Among RAS-mutated colorectal cancers, 965 were KRAS mutants (exons 2, 3, or 4; 96.3%) and 37 were NRAS mutants (exons 2, 3, or 4; 3.7%). Like RAS-mutated colorectal cancers, well/moderately differentiated tumors (P<0.0001) and microsatellite-stable phenotype (P<0.0001) were found to be associated with KRAS-mutated colorectal cancers as compared with KRAS wild-type colorectal cancers (Table 2). Moreover, results tended to show a possible association with male gender (P=0.016) and classical adenocarcinoma subtype (P=0.014). Among the colorectal cancer cohort, there were 909 KRAS exon 2 mutations (52.4%) distributed between 738 KRAS codon 12 mutants (42.5%) and 171 KRAS codon 13 mutants (9.9%; Table 3). KRAS exon 2 and KRAS codon 12 mutants, in comparison with RAS wild-type colorectal cancers, were also significantly associated with the same clinicopathological features as RAS-mutated colorectal cancers (classical adenocarcinoma subtype, well/moderately differentiated tumors and microsatellite stable phenotype; data not shown). Tumors with KRAS codon 13 mutants, in comparison with RAS wild-type colorectal cancers, were more frequently classical adenocarcinoma (95.3% versus 86.1%; P=0.004), with microsatellite-stable phenotype (90.9% versus 75.8%; P<0.0001), with perineural invasion (43.8% versus 29.9%; P=0.018) and located in the proximal colon (50.3% versus 37.4%; P=0.011).
KRAS codon 13-mutated colorectal cancers, compared with KRAS codon 12-mutated colorectal cancers, were more frequently poorly differentiated (11.5% versus 5.5%; P=0.005) and were located in the proximal colon (50.3% versus 33.8%; P=0.002; Table 3).
Association Between Rare KRAS/NRAS Mutations and Clinicopathological Features
Rare KRAS mutations were either KRAS exon 3 mutants (n=23, 1.3%) or KRAS exon 4 mutants (n=33, 1.9%; Table 3). Compared with RAS wild-type colorectal cancers, results tended to show that rare KRAS-mutated colorectal cancers (n=56) were associated with rectal site (31.2% versus 16.9%; P=0.043) and microsatellite-stable phenotype (93.5% versus 75.8%; P=0.024).
When comparing the different KRAS mutation groups to each other, regarding distribution of histologic subtypes and tumor site, we found significant discrepancies (Table 3). KRAS exon 3-mutated colorectal cancers were more frequently associated with mucinous/rare histological subtypes (17.3% versus 9.5%; P=0.002) and tended to be associated with rectal tumor site (33.3% versus 21.1%; P=0.009). KRAS exon 4-mutated colorectal cancers were associated with mucinous/rare histological subtypes (21.2% versus 9.5%; P=0.002). NRAS mutation was present in 37 cases (2.1%). Clinicopathological features in NRAS-mutated colorectal cancers were not significantly different compared with NRAS wild-type colorectal cancers (Table 2). When comparing NRAS-mutated and KRAS-mutated groups, no significant disparity was observed (data not shown).
Association Between BRAF Status and Clinicopathological Features
Patients with BRAF-mutated colorectal cancers were significantly older than patients with BRAF wild-type colorectal cancers (respectively, 74.7 and 68.2 years) and were associated with female gender (respectively, 59.5% and 38.9%; all P<0.0001; Table 2). Moreover, BRAF-mutated colorectal cancers were statistically associated with proximal tumor site (68.2% versus 31.0%), mucinous differentiation (24.5% versus 7.4%), poorly differentiated tumors (31.6% versus 7.0%), and microsatellite instability (57.0% versus 6.2%; all P<0.0001).
Clinicopathological Features of RAS and BRAF Wild-Type Colorectal Cancers
Patients with super wild-type colorectal cancers were significantly younger than patients with RAS or BRAF mutations (68.3 versus 71.3 years; P<0.0001). Moreover, super wild-type tumors were significantly associated with classical adenocarcinoma subtype (94.0% versus 86.8%; P<0.0001) and distal tumor site (54.5% versus 34.0%; P<0.0001) compared with RAS- or BRAF-mutated tumors (Table 2).
Discussion
To our knowledge, this retrospective study is one of the first to analyze associations between complete RAS and BRAF mutational status and clinicopathological features in a large cohort of colorectal cancers. As compared with RAS wild-type tumors, RAS mutants, KRAS mutants, KRAS exon 2 mutants, and KRAS codon 12 mutants tended to be associated with the same clinicopathological features, ie, male gender, classical adenocarcinoma subtype, well/moderately differentiated tumors, and microsatellite stable phenotype. KRAS codon 13 mutant colorectal cancers were more frequently classical adenocarcinoma subtype and were with microsatellite stable phenotype. For the first time, we highlighted some associations between rare RAS mutations (KRAS exons 3 and 4; NRAS mutations) and clinicopathological features. As compared with other KRAS mutations, KRAS exon 3-mutated colorectal cancers were more frequently associated with mucinous/rare histological subtypes and, most likely to the rectal tumor site, whereas KRAS exon 4-mutated colorectal cancers were associated only with mucinous/rare histological subtypes. In contrast, there was no significant association between NRAS mutation and any clinicopathological feature.
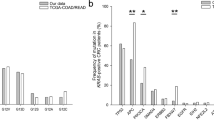
Mutation rates of RAS, KRAS, NRAS, and BRAF genes were 57.7%, 55.6%, 2.1%, and 15.5%, respectively. Among all of them, as expected, KRAS exon 2 (codons 12/13) mutation was the most frequent with 52.4%. These mutation rates were slightly above data in the literature. In recently published studies, around 50% of colorectal cancers presented a RAS mutation with an average of 45% of KRAS mutants and 40% of KRAS exon 2 mutants.3, 7, 8, 30 BRAF mutation rate in the literature is around 10%, which is slightly lower than the BRAF mutation rate found in our cohort of patients (15.5%).14, 31, 32 Sensitivity of the different molecular techniques used could explain the disparity. Since 2010, our genomic platform has been using pyrosequencing, a robust technique known to be more sensitive than Sanger sequencing used in some other platforms. In addition, our population may be enriched in RAS/BRAF mutants as colorectal cancers with incomplete RAS testing or colorectal cancers with RAS wild-type status but no BRAF testing or colorectal cancers with BRAF wild-type status but no RAS testing were automatically excluded (colorectal cancers analyzed between 2011 and 2013). Concerning rare RAS mutations (KRAS exons 3 and 4 and NRAS exons 2, 3, and 4), 5.4% colorectal cancers were concerned in our study. Rare RAS mutations were scattered between KRAS exon 3 mutants (1.3%), KRAS exon 4 mutants (1.9%), and NRAS mutants (2.1%). These rates are lower than the 10% rate previously reported by various studies.3, 9, 25, 33 Apart from the exclusion of colorectal cancers with incomplete RAS testing and from the use of different techniques with different specificity and sensitivity, no clear explanation can be drawn. The 13.4% of colorectal cancers with deficient mismatch repair system was in accordance with data found in the literature (≈12%).34, 35 In our study, patients and tumor characteristics were consistent with previously published data.30, 36, 37, 38, 39, 40
Complete RAS status with analyses of exons 2, 3, and 4 underlined some distinct clinicopathological and molecular characteristics. RAS-mutated colorectal cancers, and more precisely KRAS exon 2 and KRAS codon 12 mutants, were significantly associated with classical adenocarcinoma subtype, well/moderately differentiated tumors, and microsatellite stable phenotype. Most of our results were consistent with the literature concerning KRAS exon 2, notably the association with well/moderately histologic grade.25 Nevertheless, the data found in literature present some disparities, and some dissimilarity can be observed concerning KRAS mutations and tumor sites or histologic subtypes. Some studies have shown an association between KRAS mutation and either right colon8 or rectal tumor sites.25 Other studies precisely determined that cecal cancers exhibited significantly higher frequency of KRAS mutations than other tumor sites.22, 24 Rosty et al. has also observed gender-related distribution of KRAS-mutated carcinoma between different colonic segments.22 Association between KRAS mutation and mucinous differentiation was affirmed in some studies but denied in others.19, 20 In the same way, two studies observed that KRAS exon 2-mutated colorectal cancers seemed to occur more frequently in elderly patients, but other studies did not.22, 23 These inconsistent results could arise from a low patient number and lack of robustness of some studies. We analyzed a large cohort and found no association between KRAS exon 2 mutations with tumor site, neither with age nor with mucinous differentiation. No significant association between KRAS mutations and tumor site in our study could be explained by the fact that no data on cecum tumor site were available. Yamauchi et al. introduced a new concept of continuum colorectal cancer characterized by linear gradual changes of key tumor molecular frequencies from the rectum to the ascending colon.41 However, cecal cancers did not follow the same continuum trend and stand for a unique colorectal cancer subtype characterized by high frequency of KRAS mutation.24 Indeed, there are some differences among proximal colorectal cancers, which is a heterogeneous subgroup.
A recent publication showed that, when compared with other KRAS mutations, KRAS codon 13 mutation was associated with deficient mismatch repair phenotype and poor histologic grade.15 In our population, KRAS codon 13 mutation was correlated with poor tumor differentiation, but also with the proximal colon site. To our knowledge, there exist no published data concerning the association of rare RAS mutations (KRAS exons 3 and 4 and NRAS mutations) and clinicopathological features. We demonstrated that, in comparison with colorectal cancers with other KRAS mutations, KRAS exon 3-mutated colorectal cancers and KRAS exon 4-mutated colorectal cancers were both associated with mucinous/rare histological subtypes. Moreover, KRAS exon 3-mutated colorectal cancers tended to be associated with rectal tumor site. Finally, there was no significant association between NRAS mutations and clinical or pathological features. Therefore, correct and complete determination of the RAS genotype is required as clinicopathological features are associated with colorectal cancer prognosis value and varied according to RAS mutation.
According to previous studies, patients with BRAF-mutated colorectal cancers are aged, mostly women, with right colon tumor site, either poorly differentiated or mucinous tumors, and had tumor with deficient mismatch repair status.15, 25, 42, 43 Our results were in complete accordance with the literature.
Concerning super wild-type colorectal cancers, they were significantly associated with younger age, classical adenocarcinoma subtype, and distal colon tumor site. It is worth noting that no other study has ever evaluated the correlation between clinicopathological features and super wild-type colorectal cancers. As expected, the correlations found for clinicopathological features were opposite to those discovered for KRAS and BRAF mutated-colorectal cancers.
Part of our study limitations is because of missing clinical, histological, molecular data, or incomplete information on tumor site, explained by the fact that it is an observational retrospective study. Nevertheless, aside from vascular and perineural invasions, the major data were efficiently collected with less than 20% of missing information. Having no data on the exact proximal location site (ie, cecum or not) is one of the limitations of our study. Because of unique molecular status of cecal tumors, detailed site information in future molecular studies in colorectal cancers is necessary. Our study can be criticized on the choice made to consider RAS and BRAF mutations as mutually exclusive. However, only 1–2% colorectal cancers may carry both mutations, which is a too small percentage to change the statistical results obtained, considering the size of our study.15, 16, 17, 18, 19 On the other hand, the strength of our study was to collect and statistically correlate data from one of the largest cohorts of colorectal cancers (n=1735). To take into account multiple testing, we used Bonferroni correction but some P-value are above the alpha level and we cannot exclude true association. Above all, this is the first report correlating complete RAS and BRAF analysis with colorectal cancers’ clinicopathological features.
In conclusion, this study provides a novel broader approach of clinicopathological features according to mutational status in colorectal cancers. In particular, KRAS exon 2 (codon 13), exon 3, and 4 mutations have distinct clinical, pathological, and molecular characteristics, and must be carefully considered when assessing the prognosis value of RAS status and in clinical trials in colorectal cancers.
References
Institut National Du Cancer. Available from www.e-cancer.fr/.
De Angelis R, Sant M, Coleman MP et al. Cancer survival in Europe 1999-2007 by country and age: results of EUROCARE-5 - a population-based study. Lancet Oncol 2014;15:23–34.
Piton N, Lonchamp E, Nowak F et al. Real-life distribution of KRAS and NRAS mutations in metastatic colorectal carcinoma from French routine genotyping. Cancer Epidemiol Biomarkers Prev 2015;24:1416–1418.
Colussi D, Brandi G, Bazzoli F et al. Molecular pathways involved in colorectal cancer: implications for disease behavior and prevention. Int J Mol Sci 2013;14:16365–16385.
Karapetis CS, Khambata-Ford S, Jonker DJ et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med 2008;359:1757–1765.
Lièvre A, Bachet J-B, Corre DL et al. KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res 2006;66:3992–3995.
Yoon HH, Tougeron D, Shi Q et al. KRAS codon 12 and 13 mutations in relation to disease-free survival in BRAF-wild-type stage III colon cancers from an adjuvant chemotherapy trial (N0147 alliance). Clin Cancer Res 2014;20:3033–3043.
Taïeb J, Emile J-F, Le Malicot K et al. Prognostic value of KRAS exon 2 gene mutations in stage III colon cancer: post hoc analyses of the PETACC8 trial. J Clin Oncol 2014;25:2378–2385.
Douillard J-Y, Oliner KS, Siena S et al. Panitumumab–FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 2013;369:1023–1034.
Vaughn CP, Zobell SD, Furtado LV et al. Frequency of KRAS, BRAF, and NRAS mutations in colorectal cancer. Genes Chromosomes Cancer 2011;50:307–312.
De Roock W, Claes B, Bernasconi D et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol 2010;11:753–762.
Zlobec I, Bihl MP, Schwarb H et al. Clinicopathological and protein characterization of BRAF- and K-RAS-mutated colorectal cancer and implications for prognosis. Int J Cancer 2010;127:367–380.
Kocarnik JM, Shiovitz S, Phipps AI . Molecular phenotypes of colorectal cancer and potential clinical applications. Gastroenterol Rep 2015;3:269–276.
Seligmann JF, Fisher D, Smith CG et al. Investigating the poor outcomes of BRAF-mutant advanced colorectal cancer: Analysis from 2530 patients in randomised clinical trials. Ann Oncol 2016;28:562–568.
Gonsalves WI, Mahoney MR, Sargent DJ et al. Patient and tumor characteristics and BRAF and KRAS mutations in colon cancer, NCCTG/alliance N0147. J Natl Cancer Inst 2014;106:7.
Sahin IH, Kazmi SMA, Yorio JT et al. Mutually exclusive: a report of three cases of concomitant KRAS and braf mutation and a review of the literature. J Cancer 2013;4:320–322.
Rajagopalan H, Bardelli A, Lengauer C et al. RAF/RAS oncogenes and mismatch-repair status. Nature 2002;418:934.
Ahlquist T, Bottillo I, Danielsen SA et al. RAS signaling in colorectal carcinomas through alteration of RAS, RAF, NF1, and/or RASSF1A. Neoplasia 2008;10:680–686.
Amado RG, Wolf M, Peeters M et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 2008;26:1626–1634.
Popat S, Hubner R, Houlston RS . Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol 2005;23:609–618.
Le DT, Uram JN, Wang H et al. PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 2015;372:2509–2520.
Rosty C, Young JP, Walsh MD et al. Colorectal carcinomas with KRAS mutation are associated with distinctive morphological and molecular features. Mod Pathol 2013;26:825–834.
Koochak A, Rakhshani N, KarbalaieNiya MH et al. Mutation analysis of KRAS and BRAF genes in metastatic colorectal cancer: a first large scale study from Iran. Asian Pac J Cancer Prev 2016;17:603–608.
Yamauchi M, Morikawa T, Kuchiba A et al. Assessment of colorectal cancer molecular features along bowel subsites challenges the conception of distinct dichotomy of proximal versus distal colorectum. Gut 2012;61:847–854.
Kawazoe A, Shitara K, Fukuoka S et al. A retrospective observational study of clinicopathological features of KRAS, NRAS, BRAF and PIK3CA mutations in Japanese patients with metastatic colorectal cancer. BMC Cancer 2015;15:258.
Cortes U, Guilloteau K, Rouvreau M et al. Development of pyrosequencing methods for the rapid detection of RAS mutations in clinical samples. Exp Mol Pathol 2015;99:207–211.
Nitsche U, Stögbauer F, Späth C et al. Right sided colon cancer as a distinct histopathological subtype with reduced prognosis. Dig Surg 2016;33:157–163.
Hamilton SR, Rubio CA, Vogelstein B et al. Carcinoma of the colon and rectum. In: Hamilton SR, Aaltonen LA (eds). World Health Organization classification of tumours. Tumours of the digestive system. Lyon, France: IARC Press, 2000, pp 101–119.
Bosman F, Carneiro F, Hruban RH et al. WHO classification of tumours of the digestive system. WHO Classif Tumours Dig Syst 2010;4:417.
Imamura Y, Lochhead P, Yamauchi M et al. Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review. Mol Cancer 2014;13:135.
Baldus SE, Schaefer K-L, Engers R et al. Prevalence and heterogeneity of KRAS, BRAF, and PIK3CA mutations in primary colorectal adenocarcinomas and their corresponding metastases. Clin Cancer Res 2010;16:790–799.
Tol J, Nagtegaal ID, Punt CJA . BRAF mutation in metastatic colorectal cancer. N Engl J Med 2009;361:98–99.
Boleij A, Tack V, Taylor A et al. RAS testing practices and RAS mutation prevalence among patients with metastatic colorectal cancer: results from a Europe-wide survey of pathology centres. BMC Cancer 2016;16:825.
Jung S-B, Lee H-I, Oh H-K et al. Clinico-pathologic parameters for prediction of microsatellite instability in colorectal cancer. Cancer Res Treat 2012;44:179–186.
Sinicrope FA, Rego RL, Halling KC et al. Prognostic impact of microsatellite instability and DNA ploidy in human colon carcinoma patients. Gastroenterology 2006;131:729–737.
Knox RD, Luey N, Sioson L et al. Medullary colorectal carcinoma revisited: a clinical and pathological study of 102 cases. Ann Surg Oncol 2015;22:2988–2996.
Samara M, Kapatou K, Ioannou M et al. Mutation profile of KRAS and BRAF genes in patients with colorectal cancer: association with morphological and prognostic criteria. Genet Mol Res 2015;14:16793–16802.
Zhang J, Zheng J, Yang Y et al. Molecular spectrum of KRAS, NRAS, BRAF and PIK3CA mutations in Chinese colorectal cancer patients: analysis of 1,110 cases. Sci Rep 2015;5:18678.
Fleming M, Ravula S, Tatishchev SF et al. Colorectal carcinoma: pathologic aspects. J Gastrointest Oncol 2012;3:153.
Rosty C, Young JP, Walsh MD et al. PIK3CA activating mutation in colorectal carcinoma: associations with molecular features and survival. PLoS ONE 2013;8:e65479.
Yamauchi M, Lochhead P, Morikawa T et al. Colorectal cancer: a tale of two sides, or a continuum? Gut 2012;61:794–797.
Li W, Qiu T, Zhi W et al. Colorectal carcinomas with KRAS codon 12 mutation are associated with more advanced tumor stages. BMC Cancer 2015;15:340.
Clancy C, Burke JP, Kalady MF et al. BRAF mutation is associated with distinct clinicopathological characteristics in colorectal cancer: a systematic review and meta-analysis. Colorectal Dis 2013;15:e711–e718.
Acknowledgements
This work was provided by the Ligue contre le Cancer of Vienne, Deux-Sèvres, Charente et Charente-Maritime Departments, and the ‘Sport et Collection’ foundation. We thank J Arsham, an American translator, for having reviewed and revised the original English-language text. We thank V Le Berre, a research secretary, for her help in editing and formatting the manuscript. We also thank Pierre Rivet, Sébastien Martin, Mélanie Rouvereau, Céline Marquant, and Marine Martel for their technical involvement as well as Gwenaelle Merrien.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Rimbert, J., Tachon, G., Junca, A. et al. Association between clinicopathological characteristics and RAS mutation in colorectal cancer. Mod Pathol 31, 517–526 (2018). https://doi.org/10.1038/modpathol.2017.119
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.119
This article is cited by
-
New Developments in Treating RAS-Mutated Metastatic Colorectal Cancer
Current Treatment Options in Oncology (2023)
-
Clinicopathological and molecular characteristics of RSPO fusion-positive colorectal cancer
British Journal of Cancer (2022)
-
Upregulated NLGN1 predicts poor survival in colorectal cancer
BMC Cancer (2021)
-
Regulation of KRAS protein expression by miR-544a and KRAS-LCS6 polymorphism in wild-type KRAS sporadic colon adenocarcinoma
Human Cell (2021)
-
Mutation Status and Prognostic Value of KRAS and BRAF in Southeast Iranian Colorectal Cancer Patients: First Report from Southeast of Iran
Journal of Gastrointestinal Cancer (2021)