Abstract
Pathologists have had increasing responsibility for quantitating immunohistochemistry (IHC) biomarkers with the expectation of high between-reader reproducibility due to clinical decision-making especially for patient therapy. Digital imaging-based quantitation of IHC clinical slides offers a potential aid for improvement; however, its clinical adoption is limited potentially due to a conventional field-of-view annotation approach. In this study, we implemented a novel solely morphology-based whole tumor section annotation strategy to maximize image analysis quantitation results between readers. We first compare the field-of-view image analysis annotation approach to digital and manual-based modalities across multiple clinical studies (~120 cases per study) and biomarkers (ER, PR, HER2, Ki-67, and p53 IHC) and then compare a subset of the same cases (~40 cases each from the ER, PR, HER2, and Ki-67 studies) using whole tumor section annotation approach to understand incremental value of all modalities. Between-reader results for each biomarker in relation to conventional scoring modalities showed similar concordance as manual read: ER field-of-view image analysis: 95.3% (95% CI 92.0–98.2%) vs digital read: 92.0% (87.8–95.8%) vs manual read: 94.9% (91.4–97.8%); PR field-of-view image analysis: 94.1% (90.3–97.2%) vs digital read: 94.0% (90.2–97.1%) vs manual read: 94.4% (90.9–97.2%); Ki-67 field-of-view image analysis: 86.8% (82.1–91.4%) vs digital read: 76.6% (70.9–82.2%) vs manual read: 85.6% (80.4–90.4%); p53 field-of-view image analysis: 81.7% (76.4–86.8%) vs digital read: 80.6% (75.0–86.0%) vs manual read: 78.8% (72.2–83.3%); and HER2 field-of-view image analysis: 93.8% (90.0–97.2%) vs digital read: 91.0 (86.6–94.9%) vs manual read: 87.2% (82.1–91.9%). Subset implementation and analysis on the same cases using whole tumor section image analysis approach showed significant improvement between pathologists over field-of-view image analysis and manual read (HER2 100% (97–100%), P=0.013 field-of-view image analysis and 0.013 manual read; Ki-67 100% (96.9–100%), P=0.040 and 0.012; ER 98.3% (94.1–99.5%), p=0.232 and 0.181; and PR 96.6% (91.5–98.7%), p=0.012 and 0.257). Overall, whole tumor section image analysis significantly improves between-pathologist’s reproducibility and is the optimal approach for clinical-based image analysis algorithms.
Similar content being viewed by others
Main
The association of immunohistochemistry (IHC) expression to hormone and targeted therapy response in breast cancer care is well established for an array of biomarkers.1, 2, 3, 4 Pathologists have had increasing responsibility for quantifying IHC biomarkers such as estrogen receptor (ER), progesterone receptor (PR), human epithelial growth factor receptor-2 (HER2), Ki-67, and p53.5, 6, 7 During this time, guideline bodies (eg, College of American Pathologists) and other groups have attempted to bring pre-analytic, analytic, and post-analytic clinical standardization to the field of IHC quantitation with particular focus on the assessment of breast cancer biomarkers.7, 8 Some earlier studies evaluating manual scoring using conventional microscopy (manual read) across pathologist readers or central vs local review, have highlighted inherent between-pathologist reproducibility and within-pathologist repeatability challenges.4, 9, 10, 11, 12, 13 Some of these challenges have been in standardizing the post-analytic component of IHC quantitation including the interpretation approach, selection of representative regions (hot spots) to score, cellular expression/intensity thresholding, binning, overall positive/negative slide rating, and cut-offs. While training and various quality systems have increased pathologists’ scoring repeatability, reproducibility, and accuracy, there is still significant room for improvement.14, 15, 16, 17
Computational quantitation of IHC stained slides using digital image analysis algorithms offers the potential to improve reader precision performance.18, 19 Quantitation digitally can be assessed either using a digital read method, in which the pathologist evaluates the digital image of the glass slide on a computer monitor without the use of image analysis, or an image analysis method, which uses a digital image and a computational algorithm to provide a score (eg, 0, 1+, 2+ or 3+ or 25.43%). Recently, studies have shown incremental improvement using field-of-view image analysis over manual read in the context of HER2 IHC quantitation20 but with limited adoption clinically. Significantly increasing between-reader reproducibility could improve image analysis’s medical value proposition for clinical use.
In this paper, we culminate pathologists’ scored data across five clinical studies to understand the impact of field-of-view image analysis in the evaluation of HER2, PR, ER, Ki-67, and p53 IHC biomarkers. We then evaluate a subset of the same cases (HER2, PR, ER, and Ki-67) using a novel whole tumor section annotation approach showing superiority over field-of-view as an ideal annotation strategy for clinical quantitative image analysis.
Materials and methods
Samples and Protocol Approach
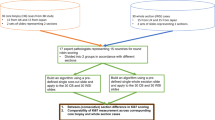
Studies reviewed here were conducted as multicenter comparative trials in the context of Food and Drug Administration regulatory submissions with three participants per biomarker who performed manual read, digital read and image analysis scorings. Each participant pathologist is American Board of Pathology certified and for any particular study, at most 1 industry pathologist reader was involved with 1–2 additional readers as non-industry pathologists (academic and/or community). All studies were performed under protocol, each of which was reviewed externally by an institutional review board. In each study, ~120 clinically obtained de-identified archived whole tissue breast carcinoma slides from Ventana Medical Systems, Inc.’s (VMSI) retrospective clinical archives (Ki-67 and p53) or from previous clinical study 510 K submissions (ER, PR, and HER2) were stained at VMSI or at study sites for the identification of the appropriate IHC biomarker receptor. Each of these assembled biomarker cohorts is considered a clinical study cohort.
Samples were binned into scoring categories targeting an approximately equal distribution across each bin. For ER, PR, Ki-67, and p53, the scoring categories were 0 to 0.99%, 1 to 10%, and >=10% (~33% each); and for HER2, they were 0, 1+, 2+, 3+ (~25% each). After binning, if there were additional samples available, they were randomly selected and added to the appropriate scoring category. For the whole tumor section sub-study, ~40 cases each from the HER2, ER, PR, and Ki-67 studies were randomly sourced from the FDA submission studies noted earlier in the same proportion of negative, borderline, and positive cases. Slides were scanned and accessioned into the respective software (VENTANA iSCAN Coreo and Virtuoso software, VMSI, Tucson, AZ, USA) at VMSI by a qualified technician, along with the hematoxylin and eosin and negative reagent control slides for each specimen. The glass slides were sent to the reader participants in order to perform manual reads.
In all cases, the participants were trained in study biomarker scoring, computer technology, and clinical study procedures before beginning. The participants binned scored each case, as positive or negative using specific thresholds (ER and PR: 1%; Ki-67 and p53: 10%, and HER2: 2+) across scoring modes (manual read, digital read and image analysis) with a 1- to 2-week washout period between modes. In all studies, the case order was randomized for each scoring session. All scoring was done in a blinded fashion with no access to the results from previous rounds of scoring. For the digital read and image analysis readings, as is the case when using a microscope, the participants had the ability to navigate freely around the images at various magnifications, select fields of view for scoring, and, for image analysis only, determine the score for the given test sample with the assistance of respective image analysis algorithms (VMSI, Tucson, AZ, USA). The participants selected a minimum of three fields-of-view per sample to assess the biomarker staining score using field-of view image analysis.
Immunohistochemistry
For each biomarker clinical study, tissue slides were stained in accordance with commercially available methods (Supplementary Tables 1–3). Tissue specimens were fixed in 10% neutral buffered formalin then dehydrated with alcohols and rinsed with reagents. Samples were then infiltrated and embedded with paraffin. Sections of each specimen were cut to the appropriate size, centered and picked up on glass slides. The tissue was either air-dried in ambient temperature overnight or placed in an oven at the recommended temperature and length of time. Positive and negative tissue run controls were implemented for each staining run. For each case, a negative reagent immunoglobulin control was also included.
Image Analysis
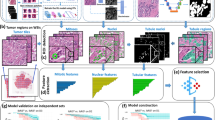
To obtain the image analysis scores in ER, PR, Ki-67, and p53 studies, biomarker specific image analysis algorithms were used to analyze each of the selected fields-of-view and whole tumor section annotations from the digitized whole slide image. Based on the computed cellular intensity and morphometric attributes, detected nuclei are classified into different cell types (ie, tumor, stroma, and lymphocyte) and subsets of specific biomarker positive and negative cells. For the HER2 study, IHC image analysis algorithms were used to detect tumor nuclei and their membrane outlines and to quantify the DAB membrane stain completeness and strength. The image analysis algorithms are machine learning-based and pre-trained on a set of independent and representative whole slide images to account for biological and stain variability. The algorithms use a single and fixed set of pre-determined parameters to analyze all the images for a specific biomarker. For visual review, the pathologists are provided with the cell counts, percentage positive score and a visual overlay of the detected biomarker’s positive and negative cells.
Statistical Analysis
For the ER and PR studies, a positive score was ≥1% and a negative score was 0–0.99%. For HER2, a positive score was 2+ or 3+ and a negative score was 0 or 1+. For Ki-67 and p53, a positive score was ≥10% and negative score was 0–9.99%. To evaluate between-reader agreement, the overall percent agreement (OPA), average positive agreement (APA), average negative agreement (ANA), and their associated confidence intervals, were calculated for each reader-pair (Supplementary Statistical Methods).
Results
Conventional Modalities
In order to understand the baseline between-pathologists’ reproducibility for manual read, digital read, and field-of-view image analysis (Supplementary Figures 1–3) modalities, we evaluated these metrics across ER, PR, HER2, Ki-67, and p53 IHC biomarker-based cohorts with three pathologists scoring all cases in each cohort. Within the ER and PR studies, scored cases showed similar overall percent agreements (OPAs) using each mode including field-of-view image analysis (Figure 1 and Table 1). For the HER2 study, using a 2+ threshold, OPA concordance between the three reader pairs was statistically higher for only field-of-view image analysis (93.8%) compared against manual read (87.2%, p=0.035).
While ER and PR IHC is a universal standard of care for clinical diagnostic analysis in breast cancer specimens, Ki-67 and p53 IHC are used less often. For both biomarkers, controversy exists with regard to clinical utility, analytic scoring methodology (hot-spot vs whole slide average), and cut-point thresholding. Within our studies, readers used a whole slide average and a threshold of 10%. For the Ki-67 study, digital read OPA (76.6%) underperformed both manual read (85.6%, P=0.166) and field-of-view image analysis (86.8%, P=0.034), while the p53 study noted similar performance across all modalities (Figure 2).
Example cases of pathologists’ fields-of-view selection and potential variability for image analysis. Scored fields-of-view are noted per case (high-power images) with image analysis overlay indicating positive (red) and negative (green) cells and individual fields-of-view quantitative scores determined by the respective algorithm (% or X+). Fields-of-view are noted in relation to the overall tumor location (green rectangles, low-power image) and culminated quantitative case score. Ground truth for each case was determined by an independent consensus panel of three pathologists.
Reviewing across all studies, the pathology participants were represented by 1 industry, 2 academic, and 3 community American Board of Pathology-certified pathologists. When using percent positivity across modalities and biomarkers as a comparative metric to understand trends between these three pathologist groups, it was observed community practice pathologists tend to have the highest percent positive calls in this study (Supplementary Table 4); however, it is important to note this group was over-represented in the overall comparison by 1 pathologist compared to the industry category.
Whole Tumor Section Annotation Methodology
Because of the limited between-reader reproducibility gain using field-of-view image analysis, we wanted to improve on this concept by implementing a novel whole tumor section approach. With the whole tumor section approach, we instructed pathologists to annotate the entire area of invasive tumor for computational calculation instead of annotating representative areas as in the fields-of-view strategy (Figure 3 upper panel). We randomly sourced approximately 40 cases each from previously evaluated ER, PR, HER2, and Ki-67 studies while maintaining ratios of negative, borderline, and positive cases for each marker. Three pathologists independently drew whole tumor section annotations for all 161 cases, and between-reader reproducibility OPAs were derived using previously noted categorical positive/negative bins (Figure 3 bottom panel). The three scoring pathologists averaged 1.3 min per case to draw a whole tumor section annotation. For HER2, Ki-67, ER, and PR, respectively, whole tumor section approach improved between-pathologists’ reproducibility as compared to field-of-view image analysis (P=0.013, 0.040, 0.232, and 0.012), digital read (P=0.008, 0.034, 0.076, and 0.249), and manual read (P=0.013, 0.012, 0.181, and 0.257). Out of a total of 161 cases, 1 ER and 2 PR cases showed disagreement. All discordant cases were intact ductal carcinoma masses without a lobular component. Review of discordant cases indicates differences in the morphologic assessment of relevant invasive tumor areas (Figure 4). For the discrepant ER case (ER809PSVRun1), pathologist 2 chose not to include a positive staining area that the other two pathologists included, reducing the slide score below the 1% threshold. For PR discrepant case 1(A089PSVRun1), pathologist 1 chose to include a tissue fold that the algorithm assessed as additional negative tumor nuclei, reducing the cut-point below the 1% threshold. For PR discrepant case 2 (C361PSVRun1), all three pathologists chose different scoring areas that impacted the overall slide score. As opposed to binned scores and assessing individual stained and unstained cell counts across the entire tumor for all scored cases, continuous quantitative scaled scores are highly reproducible indicating similar annotated areas (Supplementary Figure S4).
Whole tumor section approach. Morphologic differences between field-of-view and whole tumor section image analysis annotation approaches (top panel). Fields-of-view are drawn to scale. Whole tumor section captures relevant scoring areas, in this case, all invasive tumor cells. Overall percent agreement vs absolute difference between average positive and negative between-reader agreements for whole tumor section annotated cases (bottom panel). For each biomarker, the three dots represent the three reader (R) pairs (R1 vs R2, R1 vs R3, and R2 vs R3). For Ki-67 and HER2, all three dots for each marker are overlapping at 100%; for ER at 98.3%; for PR at 98.3% for two dots.
In terms of impact of biomarker calls around a specific threshold, average total discrepancy rates of positive and negative biomarker status flips between pathologist pairs and scoring modality showed improvement with whole tumor section image analysis (1.2% average discrepancy rate per pathologist pair) as opposed to field-of-view image analysis (9.7%), digital read (13.3%), and manual read (11.9%). Because HER2 calls have particularly significant impact to patient care and financially to healthcare systems, average discrepancy rates for this specific biomarker were none for whole tumor section image analysis as opposed to field-of-view image analysis (6.2%), digital read (9.0%), and manual read (12.8%).
Discussion
These studies used a series of pathologists with various levels of clinical diagnostic experience. For almost all of the pathologist participants, this experience was their first time utilizing digital pathology. Given this background, these results demonstrate comparability of digital read and field-of-view image analysis to manual read across all studies. In the HER2 study, the results support the notion field-of-view image analysis incrementally increases between-reader reproducibility over manual read, which echoes findings seen in another study.20
There are different ideas as to why between-pathologist variation occurs across these three modalities. The variations may be due to differences in each pathologist’s clinical experience and technological competence. As expected, familiarity comes with increased usage of image analysis technology. With the field-of-view image analysis method, since the algorithmic computational flow and parameters are fixed for algorithm use on any given slide, a major source of between-reader variability is thought to be due to readers’ selection of fields-of-view. When using image analysis, the reader is asked to select representative fields-of-view of the tumor which the algorithm analyzes to yield a score intended to be representative of the whole invasive tumor score. Choices for readers include using fields-of-view averaging and/or ratiometric strategies when annotating. This fields-of-view placement heterogeneity amongst users is seen in our study leading to reproducibility limitations on the potential advantages of image analysis. Similarly, in the manual setting, pathologists may use the same approach by selecting high magnification fields to generate a representative whole invasive tumor average. As expected, this approach can be challenging for cases with heterogeneous expression of any particular biomarker. Not surprisingly, when the initial scoring goal is clearly defined (ie, strong staining areas) such as in HER2 IHC, readers tend to select similar fields-of-view leading to similar image analysis results and potentially improvement over manual scoring.
In order to fully leverage computational quantitative consistency for between-reader reproducibility, it is critical to remove variation between users when annotating for image analysis. This can be done in potentially two ways including drawing fields-of-view until they cover the entire relevant area or through a whole tumor section approach. We chose to implement a whole tumor section approach for user efficiency including selection of relevant morphologic areas. From the subset analysis we performed using this methodology, our results show highly concordant between-reader results significantly improving over manual read, digital read, and field-of-view image analysis.
For markers requiring hot-spot scoring or another type of complicated scoring algorithmic approach such as spatial location of immune markers, geographic scoring, etc, whole tumor section approach can also be leveraged in these scenarios as the algorithm development within the annotation is not limited; the computer can be programmed to perform any of these potentially complicated tasks. For example in the case of hot spots, the computer can be programmed to identify cell staining densities in any area scored. Since the annotations would be expected to be highly similar between pathologists using a whole tumor section approach, the hot-spot or density metrics would expected to be similar. Interestingly, this approach may further be improved with automatic detection of tumor cells.21 Some of these findings are echoed in additional studies we performed using the whole tumor section approach with PD-L1 scoring in both bladder and NSCLC samples.22, 23
Limitations of this study include a smaller cohort as compared to the fields-of-view-based studies. While whole tumor section cases were randomly selected and positive, negative, and borderline cases were proportionately represented for each biomarker, a total cohort comparison may be more accurate and give better insights into the morphological selection differences between pathologists. These differences may be exacerbated in situations where morphology concordance is challenging and/or annotations are laborious to draw such as excluding multiple areas of in-situ carcinoma. Other challenging situations with limited visibility in this study are tumors with fragmented/patchy growth pattern or histologic considerations such as lobular carcinoma. These types of tumor morphologies could lead to a higher discordant rate and potentially increased time to annotate. In our study, only a total of eight cases were of lobular histology, and while none of these cases were discordant using the whole tumor section approach, these ideas need to be examined further.
Another study limitation is a complete understanding of workflow time and value-based metrics. While we did not specifically gear this study to evaluate timing metrics for manual read, digital read, and field-of-view image analysis studies, we did track a manageable time (average 1.3 min per case per annotation) using the whole tumor section approach. Because computation time across the whole slide can be lengthy (4–30 min depending on the computation power and speed), we would envision placing this task in the laboratory as an automatic pre-processing step. When this pre-processing step is complete, the case would be ready for the pathologist to score, and results would be available immediately at the time of pathologist’s annotation to review. In addition when we evaluated the positive/negative case call rates between pathologist pairs across all biomarkers and modalities, we noted a drastic decline in discrepancies using the whole tumor section approach (average 1.2% discrepancy rate) as compared with the other modalities averaged (11.6%). This is particularly significant with biomarkers such as HER2, which has a large impact for patient care and healthcare systems financially. Of course, imaging and scanning technology to enable whole tumor section approach can be of significant cost, and these considerations need to be put in context for any particular clinical need in terms of value. In order to better understand comparative impact across modalities and value, these ideas need to be examined in future studies.
This is the first comprehensive study to culminate pathologists’ derived quantitation differences in relation to four modalities and across multiple IHC clinically-used biomarkers. While field-of-view image analysis shows limited improvement for between-reader reproducibility as compared to manual read, whole tumor section image analysis clearly shows significant advantages and is an optimized approach for clinically-based quantitative image analysis algorithms.
References
Wolff AC, Hammond ME, Hicks DG et al. American Society of Clinical Oncology; College of American Pathologists. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. Arch Pathol Lab Med 2014;138:241–256.
Hammond ME, Hayes DF, Dowsett M et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. Arch Pathol Lab Med 2010;134:907–922.
Dowsett M, Nielsen TO, A'Hern R et al. International Ki-67 in Breast Cancer Working Group. Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer working group. J Natl Cancer Inst 2011;103:1656–1664.
Ciocca DR, Elledge R . Molecular markers for predicting response to tamoxifen in breast cancer patients. Endocrine 2000;13:1–10.
Prichard JW . Overview of automated immunohistochemistry. Arch Pathol Lab Med 2014;138:1578–1582.
"Paving the way for personalized medicine: FDA's role in a new era of medical product development". US Food and Drug Adminitration, October 2013. Available at: https://www.fda.gov/downloads/scienceresearch/specialtopics/personalizedmedicine/ucm372421.pdf.
Fitzgibbons PL, Bradley LA, Fatheree LA et al. Arch Pathol Lab Med 2014;138:1432–1443.
Terry J, Torlakovic EE, Garratt J et al. Implementation of a Canadian external quality assurance program for breast cancer biomarkers: an initiative of Canadian Quality Control in immunohistochemistry (cIQc) and Canadian Association of Pathologists (CAP) National Standards Committee/Immunohistochemistry. Appl Immunohistochem Mol Morphol 2009;17:375–382.
Dyhdalo KS, Fitzgibbons PL, Goldsmith JD et al. Laboratory compliance with the American Society of Clinical Oncology/College of American Pathologists human epidermal growth factor receptor 2 testing guidelines: a 3-year comparison of validation procedures. Arch Pathol Lab Med. 2014;138:876–884.
Badve SS, Baehner FL, Gray RP et al. Estrogen- and progesterone-receptor status in ECOG 2197: comparison of immunohistochemistry by local and central laboratories and quantitative reverse transcription polymerase chain reaction by central laboratory. J Clin Oncol 2008;26:2473–2481.
Gelber RD, Gelber S . International Breast Cancer Study Group; Breast International Group. Facilitating consensus by examining patterns of treatment effects. Breast 2009;18 (Suppl 3):S2–S8.
Viale G, Regan MM, Maiorano E et al. Prognostic and predictive value of centrally reviewed expression of estrogen and progesterone receptors in a randomized trial comparing letrozole and tamoxifen adjuvant therapy for postmenopausal early breast cancer: BIG 1-98. J Clin Oncol 2007;25:3846–3852.
Reisenbichler ES, Lester SC, Richardson AL et al. Interobserver concordance in implementing the 2010 ASCO/CAP recommendations for reporting ER in breast carcinomas: a demonstration of the difficulties of consistently reporting low levels of ER expression by manual quantification. Am J Clin Pathol 2013;140:487–494.
Nielsen S . External quality assessment for immunohistochemistry: experiences from NordiQC. Biotech Histochem 2015;90:331–340.
Terrenato I, Arena V, Pizzamiglio S et al. External Quality Assessment (EQA) program for the preanalytical and analytical immunohistochemical determination of HER2 in breast cancer: an experience on a regional scale. J Exp Clin Cancer Res 2013;32:58.
Torlakovic EE, Francis G, Garratt J et al. International Ad Hoc Expert Panel. Standardization of negative controls in diagnostic immunohistochemistry: recommendations from the international ad hoc expert panel. Appl Immunohistochem Mol Morphol 2014;22:241–252.
Lin F, Chen Z . Standardization of diagnostic immunohistochemistry: literature review and geisinger experience. Arch Pathol Lab Med 2014;138:1564–1577.
Ghaznavi F, Evans A, Madabhushi A et al. Digital imaging in pathology: whole-slide imaging and beyond. Annu Rev Pathol 2013;8:331–359.
Rexhepaj E, Brennan DJ, Holloway P et al. Novel image analysis approach for quantifying expression of nuclear proteins assessed by immunohistochemistry: application to measurement of oestrogen and progesterone receptor levels in breast cancer. Breast Cancer Res 2008;10:R89.
Nassar A, Cohen C, Agersborg SS et al. Trainable immunohistochemical HER2/neu image analysis: a multisite performance study using 260 breast tissue specimens. Arch Pathol Lab Med 2011;135:896–902.
Yu KH, Zhang C, Berry GJ et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat Commun 2016;7:12474.
Barnes M, Bai I, Nguyen K et al. Pathologist agreement rates of PD-L1 tumor and immune cell quantitation using digital read, field-of-view, and whole tumor image analysis. J Thorac Oncol 2016;12:S811–S812.
Barnes M, Bai I, Nguyen K et al. Pathologists' accuracy and reproducibility rates for PD-L1 immune cell quantitation using a digital pathology image analysis algorithm in urothelial carcinoma samples (abstract 2114). Mod Pathology 2017;30:523–538.
Acknowledgements
We thank the reading participant pathologists for their time and commitment to this project as well as thank Patricia Maes for her critical evaluation of this text.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
All authors are employees of Roche. This project was entirely funded by Roche.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
It is challenging for pathologists' quantitative scoring of immunohistochemistry-stained slides to meet high between-pathologist reproducibility. Technologies such as computational image analysis are available; however, its adoption is limited clinically. To increase image analysis's value proposition, the authors instituted a novel whole tumor section annotation approach across multiple modalities and biomarkers. This approach showed superiority compared to other modalities.
Supplementary information
Rights and permissions
About this article
Cite this article
Barnes, M., Srinivas, C., Bai, I. et al. Whole tumor section quantitative image analysis maximizes between-pathologists’ reproducibility for clinical immunohistochemistry-based biomarkers. Lab Invest 97, 1508–1515 (2017). https://doi.org/10.1038/labinvest.2017.82
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2017.82
This article is cited by
-
Association of artificial intelligence-powered and manual quantification of programmed death-ligand 1 (PD-L1) expression with outcomes in patients treated with nivolumab ± ipilimumab
Modern Pathology (2022)
-
Can AI-assisted microscope facilitate breast HER2 interpretation? A multi-institutional ring study
Virchows Archiv (2021)
-
Quantitative digital imaging analysis of HER2 immunohistochemistry predicts the response to anti-HER2 neoadjuvant chemotherapy in HER2-positive breast carcinoma
Breast Cancer Research and Treatment (2020)
-
Systems pathology by multiplexed immunohistochemistry and whole-slide digital image analysis
Scientific Reports (2017)