Abstract
Tumor exomes and RNASeq data were originally intended for obtaining tumor mutations and gene expression profiles, respectively. However, recent work has determined that tumor exome and RNAseq read files contain reads representing T-cell and B-cell receptor (TcR and BcR) recombinations, presumably due to infiltrating lymphocytes. Furthermore, the recovery of immune receptor recombination reads has demonstrated correlations with specific, previously appreciated aspects of tumor immunology. To further understand the usefulness of recovering TcR and BcR recombinations from tumor exome files, we developed a scripted algorithm for recovery of reads representing these recombinations from a previously described mouse model of lung tumorigenesis. Results indicated that exomes representing lung adenomas reveal significantly more TcR recombinations than do exomes from lung adenocarcinomas; and that exome files representing high mutation adenomas, arising from chemical mutagens, have more TcR recombinations than do exome files from low mutation adenomas arising from an activating Kras mutation. The latter results were also consistent with a similar analysis performed on human lung adenocarcinoma exomes. The mouse and human results for obtaining TcR recombination reads from tumor specimen exomes are consistent with human tumor biology results indicating that adenomas and high mutation cancers are sites of high immune activity. The results indicate hitherto unappreciated opportunities for the use of tumor specimen exome files, particularly from experimental animal models, to study the connection between the adenoma stage of tumorigenesis, or high cancer mutation rates, and high level lymphocyte infiltrates.
Similar content being viewed by others
Main
To obtain a tumor-exome-based assessment of T-cell lymphocyte infiltrates into mouse adenomas and adenocarcinomas (Supplementary Tables S1–S3),1 we developed a scripted algorithm modeled from, and similar to previous algorithms for mining human exome files for TcR recombination reads,2, 3, 4, 5 using mouse TcR-α, β, γ, δ sequences (imgt.org).6
Methods
Detailed methods are at the beginning of the SOM, before Supplementary Tables S1–S21. A copy of the computer script and related material is provided in the supporting online material (Supplementary Tables S4-S12 and S14, SOM), and is a duplicate of the script used for refs 4, 7 except for the use of mouse TcR sequences in this study (Supplementary Table S14). Briefly, the algorithm is designed to recover candidate V-regions, which are then searched for perfect matches to J-regions, accounting for N-region diversity at the V(D)J junctions. The resulting matches are further processed using the Immune GeneTics (IMGT) web tool6 for detecting both productive and unproductive TcR recombinations.6
Results
We processed exome files representing 86 lung adenomas and 19 lung adenocarcinomas from ref. 1. The lung adenomas were derived from three basic approaches: (i) methyl-nitrosourea treatment of wild-type mice and mice lacking one copy of Kras; (ii) urethane treatment of wild-type mice and mice lacking one copy of Kras; and (iii) untreated mice expressing an oncogenic, KrasG12D mutant allele.1 All of the adenocarcinomas were the result of urethane treatment of wild-type mice.1 The exome files were downloaded from the European Nucleotide Archive (ENA) (www.ebi.ac.uk/ena/) and were processed for recovery of the mouse TcR-α, β, γ, δ, with results summarized in Table 1. Because of the very large number of candidate TcR-α reads recovered from the 83 adenoma samples, only 19 adenoma files underwent further individual-inspection for validation of the TcR-α reads (ie, using the IMGT web tool), to match the 19 sample total for adenocarcinoma. Only reads representing an IMGT6 assessment that evaluated a minimum of 20 nucleotides, for both the V- and J-regions, were further considered in this report, but all IMGT reported TcR recombinations are available in the SOM for the candidate, recombined TcR-α and –γ reads.
Reads representing all four TcR gene recombinations were detected (Figure 1 and Supplementary Table S14). As with all previous analyses of human tumor exomes,4, 7 mouse TcR-α reads were more commonly detected in all categories, compared to TcR-β reads. This is likely reflective of a previous report that, for a many T-cells, both TcR-α alleles undergo recombination.8 Also, a recombined TcR-α allele, whether productive or unproductive, requires only one successful recombination event, whereas recombination of a TcR-β allele requires two recombination events to join the V-, D-, and J-segments. Likewise, again as in the case with human tumor exome files,4, 7 mouse TcR-γ recombinations were more commonly obtained than TcR-δ recombinations. The TcR-γ recombinations were relatively abundant, compared to previous reports for non-lung human tissues, not surprising considering the extensive documentation of γ/δ T-cells in the mouse lung.9
Example mouse TcR reads recovered from the exomes representing the adenomas in ref. 1.
Statistical comparisons of the TcR-β and TcR-δ recoveries were not possible, due to the lower levels of recoveries of reads representing these two genes. However, the combined sample counts, and combined read totals for productive and unproductive TcR-α (Figure 2a) and –γ (Figure 2b) indicated more T-cell infiltration in the adenomas versus the adenocarcinomas, consistent with high level of T-cell infiltrates reported previously for human adenomas.10, 11, 12 Similar results were obtained when considering only the productive TcR-α (Figure 2c) and –γ (Figure 2d) read counts, raising questions regarding potential sources of antigenicity in the adenomas and potential mechanisms of the apparent blunting of the immune response in the adenocarcinomas.
(a) Numbers of samples (dark column) and reads (light column) representing productive and unproductive recombinations (combined) of the TcR-α gene detected in the mouse lung adenoma and adenocarcinoma samples. In both adenoma and adenocarcinoma, 19 samples were analyzed. For the differences in the number of samples where there was detection of either productive or unproductive TcR-α recombinations, P-value<0.001 (single asterisk); for read count differences P<0.005 (double asterisk). (b) Fraction of samples (dark column) and average number of read counts per sample (light column) representing the detection of either productive or unproductive (combined) TcR-γ recombinations in the 83 adenoma or 19 adenocarcinoma samples, as indicated, with no significant differences between adenoma and adenocarcinoma. (c) Total number of reads representing productive TcR-α recombinations in 19 samples of adenoma and adenocarcinoma, respectively (P<0.04) (single asterisk). (d) Average number of reads per sample representing productive TcR-γ recombinations, 83 adenoma and 19 adenocarcinoma samples, respectively (P<0.007) (single asterisk).
Numerous reports have indicated that, in cancer patients, high mutation levels lead to more robust T-cell infiltrates, presumably due to the increased numbers of neoantigens resulting from the coding region mutations.13, 14, 15, 16, 17, 18 Thus, we assessed the detection of the TcR-α and TcR-γ recombination reads in the high mutation, mouse adenomas resulting from chemical mutagen treatment versus the adenomas resulting from KrasG12D. Results indicated a significant increase in detection of TcR-α (Figure 3a) and and TcR-γ (Figure 3b) in samples representing mutagen-derived, high mutation adenomas.
(a) Fraction of samples for the mutagen induced, high mutation frequency adenomas, or the Kras mutation induced adenomas, representing detection of a productive TcR recombination, for the indicated TcR genes, combined (P<3.0 E−06) (asterisk). (b) Average number of reads for the mutagen induced, high mutation frequency adenomas, or the Kras mutation induced adenomas, representing detection of a productive TcR recombination, for the indicated TcR genes, combined (P<0.002) (asterisk).
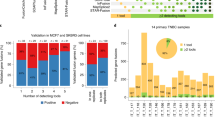
To determine whether a similar relationship could be obtained for human lung adenocarcinoma, we obtained the TCGA-LUAD exome set from the genome data commons (approved dbGaP project #6300); and processed the exome files for TcR V(D)J recombination reads. The recovered reads, along with annotations, are listed in the SOM (Supplementary Tables S15–18; for TcR-α and TcR-β, only productive reads are listed in the indicated tables). In contrast to the mouse results, the recovery of TcR-γ VJ recombination reads was minimal, with only one productive TcR-γ and one unproductive TcR-γ read recovered from 571 LUAD samples; and productive TcR-δ VDJ reads were recovered from 43 LUAD samples. Similar to the mouse, many productive TcR-α reads were recovered. However, in contrast to the above mouse data, there was no correlation of detection of TcR-α recombination reads and mutation rates. The 571 LUAD samples yielded 356 samples with TcR-β VDJ recombination reads. Thus, the use of productive TcR-α and –γ recombination reads, as a surrogate for T-cell infiltration, for determining whether there was a correlation with mutation counts, was not possible in the human LUAD data set. However, a previous study with a bladder cancer data set indicated that a favorable outcome was associated with detection of TcR-β recombination reads.4 And, overall survival is significantly better, based on the detection of TcR-β recombination reads in the TCGA melanoma data set7 (Supplementary Table S19, P<0.024). Keeping in mind that, in general, T-cell infiltrates are associated with better survival, and that T-cell infiltrates via conventional pathological assessments are associated with high mutation counts in certain human cancers, we obtained the mutation count average for the LUAD samples with productive reads for TcR-β for comparison with all the remaining LUAD samples. Results indicated that, with a statistically significant difference, TcR-β, recombination read-positive samples had, on average more mutations (Table 2 and Supplementary Table S20). This correlation also applied to the detection of TcR-δ recombination reads, ie, the mutation rate was higher among the LUAD samples with TcR-δ recombination rates (Table 2 and Supplementary Table S21). However, almost all of the samples with TcR-δ recombination reads also had TcR-β recombination reads. In sum, the recovery of TcR recombination reads in both the human and mouse settings can correlate with higher mutation rates, consistent with higher T-cell infiltrate percentages correlating with higher human mutation rates, but there is currently no explanation for why there is a difference, between human and mouse, in which TcR gene is the best surrogate for T-cell infiltration.
Discussion
The above results indicate that two aspects of mouse T-cell infiltrates, as detected by mining tumor specimen exome files, simulate natural human characteristics of cancer development: robust T-cell infiltration of adenomas and of high mutation cancers. The availability of the mouse for the study of these natural human disease states offers the opportunity to pinpoint detailed features of human cancer development, particularly related to the anti-cancer immune response, which has largely been studied via model systems that do not completely reflect complexities of the human cancers being targeted by the human immune system. For example, a current high priority for cancer treatment development is to integrate immunotherapy with conventional therapies, many of which are mutagenic and have the potential benefit of creating neoantigens in DNA repair-deficient cancers. The opportunity to use the connection between mouse mutation rates and detection of TcR recombinations will allow testing of applications of the integrative therapy approach, that is, the testing of recovered TcR recombinations for generation of TcR-based therapies in response to neoantigens developed during chemotherapy. The opportunity to pursue such goals relying on only tumor specimen exome files, with TcR recombination reads potentially serving as surrogates for T-cell infiltration quantification, is particularly attractive due to the current, extensive use of these files in patient care. That is, it is conceivable that immunotherapies based on mutation rates, or even specific mutations, could be designed with little added expense for obtaining the cognate immune receptors. The mouse model now represents an opportunity to lay the ground work for such a goal.
A more immediate application of the above approach, of using tumor specimen exome files to assess the immune-microenvironment, could be related to immune checkpoint based therapies. In particular, recent work has indicated a strong correlation between the co-detection of TcR-α and TcR-β recombination reads and the high level expression of PD-1, in melanoma.7 This result suggests that tumor specimen exome files alone could be used to justify anti-immune checkpoint therapy. While this justification may not be needed in settings of robust, direct testing for PD-1 expression in patient tumors, for making therapy decisions, the use of tumor specimen exomes for justifying such therapy, when TcR recombination reads can be detected in those exomes, may represent the kind of cost efficiency that could impact settings where direct testing of PD-1 expression is effectively cost-prohibitive. Again, the mouse model offers an opportunity to understand the details of the relationship between PD-1 expression and recovery of recombination reads from tumor specimen exome files such that the use of the exomes for making anti-PD-1 therapy decisions is done with the highest possible success rates.
References
Westcott PM, Halliwill KD, To MD,et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 2015; 517: 489–492.
Gill TR, Samy MD, Butler SN et al, Detection of productively rearranged TcR-alpha V-J sequences in TCGA exome files: implications for tumor immunoscoring and recovery of antitumor T-cells. Cancer Inform 2016; 15: 23–28.
Levy E, Marty R, Garate Calderon V et al, Immune DNA signature of T-cell infiltration in breast tumor exomes. Sci Rep 2016; 6: 30064.
Samy MD, Tong WL, Yavorski JM et al, T cell receptor gene recombinations in human tumor specimen exome files: detection of T cell receptor-beta VDJ recombinations associates with a favorable oncologic outcome for bladder cancer. Cancer Immunol Immunother 2017; 66: 403–410.
Tong WL, Tu YN, Samy MD et al, Identification of immunoglobulin V(D)J recombinations in solid tumor specimen exome files: evidence for high level B-cell infiltrates in breast cancer. Hum Vaccin Immunother 2017; 13: 501–506.
Alamyar E, Duroux P, Lefranc MP et al, IMGT((R)) tools for the nucleotide analysis of immunoglobulin (IG) and T cell receptor (TR) V-(D)-J repertoires, polymorphisms, and IG mutations: IMGT/V-QUEST and IMGT/HighV-QUEST for NGS. Methods Mol Biol 2012; 882: 569–604.
Tu YN, Tong WL, Samy MD et al, Assessing microenvironment immunogenicity using tumor specimen exomes: co-detection of TcR-alpha/beta V(D)J recombinations correlates with PD-1 expression. Int J Cancer 2017; 140: 2568–2576.
Padovan E, Casorati G, Dellabona P et al, Expression of two T cell receptor alpha chains: dual receptor T cells. Science 1993; 262: 422–424.
Lafont V, Sanchez F, Laprevotte E et al, Plasticity of gammadelta T cells: impact on the anti-tumor response. Front Immunol 2014; 5: 622.
Dennis KL, Wang Y, Blatner NR et al, Adenomatous polyps are driven by microbe-instigated focal inflammation and are controlled by IL-10-producing T cells. Cancer Res 2013; 73: 5905–5913.
McLean MH, Murray GI, Stewart KN et al, The inflammatory microenvironment in colorectal neoplasia. PLoS One 2011; 6: e15366.
Bedossa P, Poynard T, Bacci J et al, Expression of histocompatibility antigens and characterization of the lymphocyte infiltrate in hyperplastic polyps of the large bowel. Hum Pathol 1990; 21: 319–324.
Snyder A, Makarov V, Merghoub T et al, Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 2014; 371: 2189–2199.
Bouffet E, Larouche V, Campbell BB et al, Immune checkpoint inhibition for hypermutant glioblastoma multiforme resulting from germline biallelic mismatch repair deficiency. J Clin Oncol 2016; 34: 2206–2211.
Westdorp H, Fennemann FL, Weren RD et al, Opportunities for immunotherapy in microsatellite instable colorectal cancer. Cancer Immunol Immunother 2016; 65: 1249–1259.
Maletzki C, Schmidt F, Dirks WG et al, Frameshift-derived neoantigens constitute immunotherapeutic targets for patients with microsatellite-instable haematological malignancies: frameshift peptides for treating MSI+ blood cancers. Eur J Cancer 2013; 49: 2587–2595.
Howitt BE, Shukla SA, Sholl LM et al, Association of polymerase e-mutated and microsatellite-instable endometrial cancers with neoantigen load, number of tumor-infiltrating lymphocytes, and expression of PD-1 and PD-L1. JAMA Oncol 2015; 1: 1319–1323.
Gros A, Robbins PF, Yao X et al, PD-1 identifies the patient-specific CD8(+) tumor-reactive repertoire infiltrating human tumors. J Clin Invest 2014; 124: 2246–2259.
Acknowledgements
We acknowledge the support of the USF research computing facility and the taxpayers of the State of Florida and Dr Peter Westcott, for helpful discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
It is challenging for pathologists' quantitative scoring of immunohistochemistry-stained slides to meet high between-pathologist reproducibility. Technologies such as computational image analysis are available; however, its adoption is limited clinically. To increase image analysis's value proposition, the authors instituted a novel whole tumor section annotation approach across multiple modalities and biomarkers. This approach showed superiority compared to other modalities.
Supplementary information
Rights and permissions
About this article
Cite this article
Tu, Y., Tong, W., Fawcett, T. et al. Lung tumor exome files with T-cell receptor recombinations: a mouse model of T-cell infiltrates reflecting mutation burdens. Lab Invest 97, 1516–1520 (2017). https://doi.org/10.1038/labinvest.2017.80
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2017.80
This article is cited by
-
An age-based, RNA expression paradigm for survival biomarker identification for pediatric neuroblastoma and acute lymphoblastic leukemia
Cancer Cell International (2019)
-
Immune receptor recombinations from breast cancer exome files, independently and in combination with specific HLA alleles, correlate with better survival rates
Breast Cancer Research and Treatment (2019)
-
T-cell receptor-α CDR3 domain chemical features correlate with survival rates in bladder cancer
Journal of Cancer Research and Clinical Oncology (2019)
-
Merkel cell polyomavirus-specific immune responses in patients with Merkel cell carcinoma receiving anti-PD-1 therapy
Journal for ImmunoTherapy of Cancer (2018)
-
T-cell receptor-β V and J usage, in combination with particular HLA class I and class II alleles, correlates with cancer survival patterns
Cancer Immunology, Immunotherapy (2018)