Abstract
Metastasis underlies cancer morbidity and accounts for disease progression and significant death rates generally and in non-small cell lung cancer (NSCLC) particularly. Therefore, it is critically important to understand the molecular events that regulate metastasis. Accumulating data portray a central role for protein synthesis, particularly translation initiation (TI) factors eIF4E and eIF4G in tumorigenesis and patients' survival. We have published that eIF4E/eIF4GI activities and consequently NSCLC cell migration are modulated by bone-marrow mesenchymal stem cell secretomes, suggesting a role for TI in metastasis. Here, we aimed to expand our understanding of the TI factors significance to NSCLC characteristics, particularly epithelial-to-mesenchymal transition (EMT) and migration, supportive of metastasis. In a model of NSCLC cell lines (H1299, H460), we inhibited eIF4E/eIF4GI's expressions (siRNA, ribavirin) and assessed NSCLC cell lines' migration (scratch), differentiation (EMT, immunoblotting), and expression of select microRNAs (qPCR). Initially, we determined an overexpression of several TI factors (eIF4E, eIF4GI, eIF4B, and DHX29) and their respective targets in NSCLC compared with normal lung samples (70–350%↑, P<0.05). Knockdown (KD) of eIF4E/eIF4GI in NSCLC cell lines (70%↓, P<0.05) also manifested in decreased target levels (ERα, SMAD5, NFkB, CyclinD1, c-MYC, and HIF1α) (20–50%↓, P<0.05). eIF4E/eIF4GI KD also attenuated cell migration (60–75%↓, P<0.05), EMT promoters (15–90%↓, P<0.05), and enhanced EMT suppressors (30–380%↑, P<0.05). The importance of eIF4E KD to NSCLC phenotype was further corroborated with its inhibitor, ribavirin. Changes in expression of essential microRNAs implicated in NSCLC cell migration concluded the study (20–100%, P<0.05). In summary, targeting eIF4E/eIF4GI reduces migration and EMT, both essential for metastasis, thereby underscoring the potential of TI targeting in NSCLC therapy, especially the already clinically employed agents (ribavirin/4EGI). Comparison of these findings with previously reported effects of eIF4E/eIF4GI KD in multiple myeloma suggests a collective role for these TI factors in cancer progression.
Similar content being viewed by others
Main
Lung cancer is the leading cause of cancer-related death worldwide. Non-small cell lung cancer (NSCLC) accounts for 85% of all lung cancer cases. Despite several novel therapeutic approaches, the majority of NSCLC patients present locally advanced or metastatic disease, which are at present incurable.1, 2 Thus, there is great need for new therapies. The key factor of NSCLC-associated mortality is attributed to metastasis marking its underlying mechanisms as potentially important targets in the quest for effective treatment.3, 4 Indeed, during metastasis, tumor cells need to survive detachment from their primary microenvironment, acquire migratory characteristics, invade neighboring tissues, intravasate and survive circulation, extravasate into new distant microenvironments, settle and eventually start proliferating to establish new tumor sites. A major process that allows cancer cells to metastasize is epithelial-to-mesenchymal transition (EMT) where polarized epithelial cells lose cell–cell adhesion and cell polarity and acquire a motile and invasive phenotype.5 Moreover, it was demonstrated that EMT is involved in treatment resistance, acquisition of characteristics of cancer stem-like cells, and progression of many tumors types.5 Unraveling the molecular events that promote EMT and metastasis along with improving the means of foretelling their development is a major goal of current clinical cancer research, NSCLC included.
Control of protein translation is a crucial aspect of cancer development and progression.6 Translational control includes the regulation of global protein synthesis rate as well as selective translation of specific mRNAs that promote tumor cell survival, angiogenesis, invasion, and metastasis. Deregulation of protein translation has been observed in various human malignancies with both elevated global translation and increased synthesis of proteins involved in malignant characteristics.7 Translation initiation (TI) is the most regulated step of protein synthesis and the rate limiting phase of the process.8 In concordance, high expression of the TI factors, eukaryotic translation initiation factor 4E (eIF4E), and eukaryotic translation initiation factor 4GI (eIF4GI) was reported in various tumors9 including lung cancer.10, 11
Overexpression and activity of eIF4E are associated with tumor formation, metastatic disease, and increased tumor invasion. Moreover, higher levels of eIF4E were demonstrated in metastatic cells compared with primary tumor cells.12 It is established that eIF4E is rate limiting to 5' cap-dependent translation typical of 90% of cellular proteins.13, 14 Also well-known, eIF4G (I, II) is a key initiator of eIF4F-complex assembly15 and eIF4GI is the major isoform in mammalian cells (>85%).16 Of note, accumulating data suggest that eIF4E and eIF4GI may dictate not only rate of protein synthesis but also its quality17 thereby contributing to cell phenotype design. Interestingly, additional TI factors are deregulated in cancers.18, 19 For instance, eIF4B, a factor critical for recruitment of mRNA to the ribosome and assistance in unwinding mRNAs secondary structures necessary for ribosome scanning is often overexpressed.19 DHX29 is an ATP-dependent RNA helicase implicated in cap-dependent TI and promotion of cell proliferation, and tumorigenesis was also reported to be elevated in several cancer models.18 Previous publications by others and us6, 20, 21, 22 have demonstrated the critical role of protein translation in lung cancer cells23 with specific emphasis on translational machinery.24 We have investigated different aspects of proteostasis in NSCLC and showed that its manipulation by proteasome inhibition or induction of ER stress damaged lung cancer cells.21 We have also shown manipulation of NSCLC TI by microenvironmental constituents, ie, mesenchymal stem cells (MSCs) affected the cancer cells' migration.25 Finally, in a multiple myeloma (MM) cancer model we have shown that eIF4E and eIF4GI are critically modulated by microenvironmental components and important to the malignant cells' phenotype and protein repertoire.26
On the basis of all these accumulating data, we wanted to expand our understanding of TI to NSCLC characteristics and fate. We concentrated on the role of eIF4E/eIF4GI in the design of NSCLC cells' proteome and phenotype, particularly traits that promote metastasis.
Materials and methods
NSCLC Cell Lines and Samples
H1299 and H460 were cultured in RPMI-1640 with 10% heat-inactivated fetal bovine serum and antibiotics (Biological Industries, Israel). NSCLC and the corresponding normal lung tissue specimens were obtained from therapy-naive patients who undergo lung surgery for newly diagnosed NSCLC (complying with Helsinki regulations). Samples were minced into 1 mm3 pieces digested and proteins were extracted.
Trypan Blue
Total cell counts as well as the respective proportion of viable and dead cells were enumerated by Trypan blue dye exclusion using a hematocytometer and a phase-contrast microscope as described before.20
Cell Count, Viability, Cycle, and Death
Cells were counted with ADVIA 120 Automated Hematology Analyzer (GMI, Minneapolis, MN, USA) and assayed for viability with WST1 Cell Proliferation Reagent (Roche, Basel, Switzerland) according to the manufacturer's instructions. Cell death was determined by flow cytometry. Cells were harvested and stained with annexin V-PE (250 μg/ml) (BioVision, CA, USA) and 7AAD (0.05 μg/μl) (eBioscience, CA, USA). Annexin V+/7AAD− cells were considered apoptotic and annexin V+/7AAD+ cells were considered necrotic or late apoptotic cells. For cell cycle harvested cells were exposed to 40 μg/ml propidium iodide (PI) and 100 μg/ml Ribonuclease A (Sigma, St. Louis, MO, USA) in PBS for 30 min at room temperature in the dark and analyzed by FACS.
Immunoblotting
NSCLC cells were lysed and western blot was performed as described elsewhere.27 The proteins were detected with rabbit/mouse anti-human: peIF4E(Ser209)/total eIF4E, peIF4GI(Ser1108)/total eIF4GI, peIF4B (Ser422)/total eIF4B, DHX29, CyclinD1 (Cell Signaling, Danvers, MA, USA); SMAD5, HSC70 (Epitomics, CA, USA); and c-Myc, HIF1α, NFkB (Santa-Cruz, CA, USA); ERα (Millipore, Billerica, MA, USA). Bound antibodies were visualized using peroxidase-conjugated secondary goat anti-rabbit or mouse antibody (Jackson ImmunoResearch Laboratories, West Grove, PA, USA), followed by enhanced chemiluminescence (ECL) detection (Millipore). Products were visualized with LAS3000 Imager (Fujifilm, Greenwood, SC, USA). Integrated optical densities of the immunoreactive protein bands were measured as arbitrary units employing Multi Gauge software v3 (Fujifilm).
Scratch Assay
Scratch assay was conducted as described before.20 Wound closure was monitored by microscopy immediately after cell scratching (0 h) and at 24 h post wounding.
Ribavirin Treatment
NSCLC cultured cells were treated with ribavirin (5 μM) (Sigma) diluted in water for 24 h.
siRNA Transfection
Validated Alexa-labeled AllStars negative control and anti-eIF4E (20 pmol) and combination of five different sequences anti-eIF4G (20 pmol: 4 pmol each, four designed by Qiagen and one published previously28) (Qiagen, Valencia, CA, USA) were delivered into H1299 and H460 NSCLC cell lines using Lipofectaime2000 (Invitrogen, Carlsbad, CA, USA). Fluorescence (≥10 000 events/analysis) was analyzed by flow cytometery (Navios, Beckman Coulter, USA) and determined in 95% of the cells (transfection efficiency). Silencing of eIF4E/eIF4G was detected at the RNA level by qPCR and at the protein level by western blot. In all, 24, 48, and 72 h post transfection time points were tested.
Quantitative Reverse Transcription PCR
Total RNA was extracted with RNeasy kit (Qiagen). RNA (1 μg) was reverse transcribed (GeneAmp RNA PCR, Applied Biosystems, Carlsbad, CA, USA) and amplified (Power SYBR Green, Applied Biosystems) according to the manufacturer's instructions. TI factors and their targets were assayed: eIF4E's - DOCK4, SERPINI1, IL23A, EGR1eIF4GI's - FBXO32, GAB2, BCL2, and ASS1. All primers' sequences were described by us before.29
microRNAs Quantitative Reverse Transcription PCR
RNA was extracted from with TRI Reagent and was converted to cDNA using the Quanta reverse transcription kit (Quanta-Bioscience) according to the manufacturer's instructions. Human small nucleolar RNA SNORD44 was amplified as an internal control. Amplification was performed using Power SYBR Green PCR Master Mix (Quanta-Bioscience).
Statistical Analysis
Student’s paired t-tests were used in analysis of differences between cohorts. Effects were considered as significant, when P-value is <0.05.
Results
NSCLC Cells Exhibit Elevated Levels of TI Factors Compared with Normal Lung Tissue
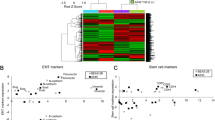
Primarily, we set out to examine the baseline expression levels of the major TI factors, already recorded to be aberrant in multiple malignancies but yet to be determined in NSCLC. Expressions of phosphorylated and total eIF4E, eIF4GI, eIF4B, and DHX29 were examined in NSCLC cell lines (H1299, H460, and A549) and primary tumor samples (n=6). Findings were compared with expression levels in normal lung tissue specimens that were obtained from therapy-naive patients who underwent lung surgery for newly diagnosed NSCLC (complying with Helsinki regulations) (n=6). In concordance with observations in other cancer models, we determined elevated expression of all four assayed TI factors in NSCLC samples (lines-70–320%↑, primary-90–280%↑; P<0.05) (Figures 1a–c). To test whether elevated eIF4E and eIF4GI levels translated into increased activity, we studied the expression levels of their established targets (eIF4E: CyclinD1; eIF4GI: SMAD5). Again, we compared expression levels in NSCLC cells with normal lung cells. Indeed, significantly higher levels were observed in the NSCLC cells (lines and primary tumors) compared with control (cell lines: 125–350%↑, NSCLC samples: 70%↑; P<0.05) (Figure 1d). The elevated TI factors' expression in NSCLC cells suggests that they may have a role in the malignant process.
NSCLC cells exhibit elevated levels of translation initiation factors compared with normal lung tissue. Graphic presentations of the baseline levels expression of phosphorylated (p) and total (t) (a) eIF4E; (b) eIF4GI (c) eIF4B and DHX29; (d) CyclinD1 and SMAD5; in NSCLC cell lines (H1299, H460, and A549), NSCLC tumor samples (n=6) and normal lung tissue samples (n=6) (X axis). Results were detected by immunoblotting and expressed as arbitrary units (mean±s.e.) of immunoblotting densitometry. Each sample (normalized to HSC70) is indicated by an empty circle and group averages are depicted by line. Statistically significant differences of NSCLC cell lines vs. normal samples (*P<0.05) and NSCLC tumor samples vs normal samples ($P<0.05) are indicated.
Knockdown of eIF4E/eIF4GI TI Factors
To directly investigate whether eIF4E/eIF4GI TI factors' activities are indeed critical to NSCLC progress, we introduced anti-eIF4E/eIF4GI siRNAs into H1299 and H460 NSCLC cell lines (knockdown, KD). Negative siRNA was delivered as a control. Ninety-five percent of the cells were transfected (validated by FACS) (data not shown). Factors' silencing was validated 24 h post transfection at the transcript level (>70%↓ for si-eIF4GI/eIF4E, P<0.01) and 48 h post transfection at the protein level (70%↓ for si-eIF4GI/eIF4E, P<0.01) without reciprocal effects (Figure 2a). The KD was deliberately of limited extent in order to avoid dramatic cell damage due to total protein synthesis shutdown that might mask the nuances of the relative TI factors. Having established that the expressions of eIF4E and eIF4GI were decreased we wanted to validate that the decreased levels also affected eIF4E/eIF4GI's activities. Thus, we assayed the levels of targets previously shown to depend on these TI factors for their expression (functional assay).26, 29, 30, 31, 32, 33, 34 Significant attenuation of NFkB and CyclinD1 levels was observed in both cell lines 72 h post transfection, demonstrating that eIF4E activity was indeed compromised (20–50%↓; P<0.05) (Figure 2b). Reduced SMAD5 and ERα levels demonstrated that eIF4GI activity was also attenuated (25–35%↓; P<0.05) (Figure 2b). Finally, expression of eIF4E/eIF4GI common targets, HIF1α and c-Myc, was also decreased in the KD models (40–70%↓; P<0.05) (Figure 2b). Again, established eIF4E targets were not affected by the eIF4GI silencing and vice versa (data not shown). Of note, these results are in concordance with similar KD model we established previously in MM cells.29 The similarity in eIF4E/eIF4GI KD in two different cancer models underscores the authenticity of these observations and their general relevance. Subsequently, we assessed the effect of the KDs on NSCLC cells’ viability, proliferation, cell cycle, and death. No changes in any survival or proliferative characteristics were observed (Supplementary Figure 1).
Knockdown of eIF4E/ eIF4GI translation initiation factors. H1299 and H460 cells were transfected with control negative siRNA (si-NEG) or anti-eIF4E siRNA (si-E) or anti-eIF4GI siRNA (si-G). (a) Following 48 h, the cells were immunoblotted for eIF4E and eIF4GI expression levels. (b) Following 72 h, the cells were also immunoblotted for eIF4E/eIF4GIs' targets. Graphic (left) and (right) representative immunoblots are presented (mean±s.e.). Protein quantities were normalized to HSC70 that served as a loading control. Statistically significant differences (*P<0.05; **P<0.01) are depicted.
Knockdown of eIF4E/eIF4GI TI Factors Inhibited NSCLC Cell Lines' Migration
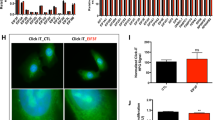
In a previous study,25 we demonstrated that bone-marrow MSCs' secretome downregulated the eIF4E/eIF4GI expression and cells' migration. Here, we wanted to assess directly the role of eIF4E/eIF4GI in NSCLC cells' migration. Thus, we knocked down the factors (separately) in NSCLC cell lines (H1299 and H460) and tested our premise by applying the scratch assay. Results indicated that eIF4E or eIF4GI KDs significantly retarded the cells' scratch closure after 24 h compared with negative control cells (negative siRNA) (70–75%↓ and 60%↓, respectively; P<0.05) (Figures 3a and b). In concurrence with the cells' altered migration, we also observed changes in the cells' morphology. Indeed, while control migrating NSCLC cells exhibited a more fibroblast-like spindle appearance, the KD cells displayed an epithelial-like more round morphology (Figure 3c).
eIF4E/eIF4GI KD inhibited NSCLC cell lines' migration. NSCLC cell lines were transfected with control negative siRNA (si-NEG) or anti-eIF4E siRNA (si-E) or anti-eIF4GI siRNA (si-G), and the effect on cell migration was assessed by scratch assay. Scratch closure was photographed immediately (0 h) and after 24 h (magnification, × 40). Results are presented as (a) percent of the scratch closure in bar graphs (mean±s.e., n≥3) (the change percentages of each silencing compare with negative control are depict above the lines between cohorts) and (b) representative images. Statistically significant differences between cohorts (*P<0.05, **P<0.01) are indicated. (c) Morphology changes of si-E/si-G-silenced NSCLC cells in comparison with negative control (representative images). Control cells displayed spindle-shaped and fibroblast-like shape, while si-E/si-G cells reserved their epithelial morphology.
Competitive Inhibition of eIF4E with Ribavirin Negatively Affected NSCLC Cell Lines’ Migration
Our findings so far indicate that eIF4E and eIF4GI diminution deleteriously affects NSCLC cells' migration but the delivery of siRNA is currently clinically difficult. Therefore, we examined the effect of the clinically employed anti-viral drug, ribavirin previously shown to act as a mimic for the 5' cap present on most human transcripts thereby competing with eIF4E binding.35, 36, 37, 38 Ribavirin applied for 24 h did not affect the treated NSCLC cell lines' viability/proliferation/death thereby indicating that under these conditions there was no toxic effect (data not shown). Yet, as hypothesized, ribavirin (5 μM) decreased the cells' (H1299 and H460) migration compared with untreated cells (65–75%↓, 24 h; P<0.05) (Figures 4a and b). Here too, we observed changes in the cells' morphology. While control migrating cells showed fibroblast-like spindle appearance the ribavirin-treated cells displayed an epithelial-like more round morphology (Figure 4c).
Competitive inhibition of eIF4E with ribavirin negatively affected NSCLC cell lines’ migration. NSCLC cell lines were treated with the eIF4E inhibitor ribavirin (5 μM) for 24 h. The effect of ribavirin on NSCLC cells’ migration was assessed by scratch assay. Scratch closure was photographed immediately (0 h) and after 24 h (magnification, × 40) of cultured cells’ area. Results are presented as (a) percent of the scratch closure in bar graphs (mean±s.e., n≥3) (the change percentages of ribavirin treatment compared with control are depicted above the lines between cohorts) and (b) representative images. Statistically significant differences between cohorts (*P<0.05) are indicated. (c) Morphology changes of treated cells in comparison with negative control (representative images). Control cells displayed spindle-shaped and fibroblast-like shape, while ribavirin-treated cells reserved their epithelial morphology.
In summary, our results so far indicate that manipulation of TI pathways indeed affects NSCLC cells' migration.
Knockdown of eIF4E/eIF4GI TI Factors Downregulated EMT of NSCLC Cell Lines
Studies using ribosome profiling have revealed that oncogenic eIF4E activity downstream of mTOR signaling has a striking effect on the translational landscape of the cancer genome, particularly in the context of metastasis.39 This study has functionally characterized a novel subset of translationally regulated mRNAs associated with cancer cell invasion and metastasis. Moreover, previous data demonstrate that eIF4E has a prominent role in EMT by controlling the translation of a pool of mRNAs critical for EMT.40, 41, 42 Our observations so far indicate that eIF4E and eIF4GI have a role in the regulation of NSCLC cell migration. Since, cell migration may be a symptom of EMT, we hypothesized that EMT will be downregulated in eIF4E/eIF4GI-silenced NSCLC cells (ie, the cells will undergo mesenchymal-to-epithelial transition (MET)). Thus, we set out to examine the expression of EMT/MET markers by immunoblotting eIF4E/eIF4GI KD NSCLC cell lines (H1299 and H460) 48 h post siRNA transfection. First, we examined changes in cell–cell adhesion molecules E-cadherin/N-cadherin known to have a critical role in the EMT/MET. E-cadherin is considered as an active suppressor of invasion and growth of many epithelial cancers5, 43 and the term ‘cadherin switching’ refers to a switch from expression of E-cadherin to expression of N-cadherin.44 In concordance with our hypothesis, we determined elevated levels of E-cadherin (100–290%↑; P<0.05) and reduced expression of N-cadherin (40–90%↓; P<0.05) (Figures 5a and b). Next, we assayed the tight junction-associated proteins claudin-1 and zona occludens-1 (ZO-1) that affect cell–cell junctions and migration.45 Indeed, we observed elevated expression of claudin-1 (30–380%↑; P<0.05) and ZO-1 (40–220%↑; P<0.05), which is consistent with decreased EMT (Figures 5a and b). Finally, we determined reduced levels of two transcription factors known to promote EMT: Slug a transcriptional repressor of integrin that causes decreased cell adhesion46 (40–70%↓; P<0.05) and ZEB1 that suppresses E-cadherin44 (40–60%↓; P<0.05) (Figures 5a and b).
eIF4E/eIF4GI KD downregulated epithelial-to-mesenchymal transition in NSCLC cell lines. H1299 and H460 cells were transfected with control negative siRNA (si-NEG) or anti-eIF4E siRNA (si-E) or anti-eIF4GI siRNA (si-G). Following 48 h, the cells were immunoblotted for EMT markers expression levels: N-cadherin, E-cadherin, Claudin-1, ZO-1, Slug, and ZEB1. (a) Graphic and (b) representative immunoblots are presented (mean±s.e.). Protein quantities were normalized to HSC70 that served as a loading control. (c) RNA was extracted 24 h post transfection and tested for EMT markers (BMP7, FN1) at the transcript level. Statistically significant differences (*P<0.05; **P<0.01) are depicted.
Subsequently, we examined EMT markers at the transcriptional level by qPCR and here too determined changes consistent with reduced EMT and induction of MET. Specifically, we observed elevated levels of the EMT inducer bone morphogenetic protein 7 (BMP7)47 (30–60%↑; P<0.05), as well as decreased fibronectin (FN1)48 (15–35%↓; P<0.05) (Figure 5c).
Altogether, our data indicate that eIF4E and eIF4GI promote NSCLC cells EMT and that inhibition of their function facilitates EMT/MET switch and consequently cell migration and metastatic potential.
eIF4E/eIF4GI KDs Affected MicroRNAs with a Possible Role in NSCLC Migration
Accumulating data underscore the importance of microRNAs in intercellular crosstalk and modulation of major cellular functions such as survival and migration.49 Furthermore, several microRNAs are known to be regulated by eIF4E/eIF4GI factors.50, 51 Therefore, we tested the expression of specific microRNAs in eIF4E/eIF4GI-silenced cells that may be involved in NSCLC migration and EMT.
Previously, we showed that eIF4E/eIF4GI silencing in MM cells affected MIR-125a expression using bioinformatics tools and qPCR validations.29 Interestingly, published data reported that MIR-125a suppresses NSCLC cell proliferation, migration, and invasion.52 Thus, we tested the expression of MIR-125a in eIF4E/eIF4GI KD NSCLC cell lines by qPCR. Both NSCLC cell lines displayed elevated levels of MIR-125a 48 h post eIF4E/eIF4GI silencing (45–65%↑; P<0.05) (Figure 6a). A recent study depicts a role for MIR-29b in NSCLC metastatic suppression via inhibition of MMP2 expression.53 In our KD cells, we observed induction of MIR-29b levels after 48 h (35–55%↑; P<0.05) (Figure 6b). Moreover, we also tested the levels of MMP2 as an MIR-29b probable downstream tumor metastasis-related gene. Indeed, NSCLC-transfected cells displayed decreased levels of MMP2 mRNA (35–45%↓; P<0.05) (Figure 6b). Finally, we examined the expressions of MIR-148b and MIR-23a in the eIF4E/eIF4GI KDs, both previously implicated in NSCLC migration.54, 55 MIR-148b has been shown to function as a tumor suppressor with anti-proliferation, migration, and EMT activity.54 On the contrary, MIR-23a acts as an oncogene and regulates TGF-β-induced EMT by targeting E-cadherin in NSCLC.55 In concordance with our accumulating observations and work hypothesis, we observed opposite changes in expressions of MIR-148b (20–100%↑; P<0.05) and MIR-23a (40%↓; P<0.05) (Figure 6a).
microRNAs important to NSCLC migration were affected by eIF4E/eIF4GI KDs. H1299 and H460 cells were transfected with control negative siRNA (si-NEG) or anti-eIF4E siRNA (si-E) or anti-eIF4GI siRNA (si-G). Following 48 h, the cells were harvested and microRNAs were extracted. The expression levels of (a) MIR-125a, MIR-148b, MIR-23a, (b) MIR-29b, (c) MIR-96, MIR-146, and (b) MMP2 were tested by qPCR. Statistically significant differences (*P<0.05; **P<0.01) are depicted.
Our results so far indicate that similar microRNAs were affected by both eIF4E and eIF4GI KDs. We wondered whether there are microRNAs that uniquely depend on a specific TI factor for their expression and may influence NSCLC migration. According to previous publications MIR-96 is regulated by eIF4E and promotes growth and motility of NSCLC cells.51 Moreover, its’ expression levels in NSCLC tissues were higher than in normal lung tissues, which was significantly associated with lymph-node metastasis.29, 56 MIR-146 is regulated by eIF4GI and inhibits cell growth, migration and induces apoptosis in NSCLC.50 Upon examination of these microRNAs in our research model, we indeed witnessed that eIF4E KD affected MIR-96 expression in the NSCLC cell lines (25–45%↓; P<0.05) and eIF4GI KD affected MIR-146 expression (60–70%↑, respectively; P<0.05) with no reciprocal effects between the KDs (Figure 6c).
In summary, our results depict significant changes in multiple microRNAs consistent with decreased NSCLC migration and EMT. These observations may afford a mechanistic explanation for the eIF4E/eIF4GI KDs influence on the NSCLC cell phenotype.
eIF4E/eIF4GI KDs Differentially Affected Transcription Factors Important to the Tumorigenic Phenotype of NSCLC Cells
In a previous publication,29 we silenced eIF4E and eIF4GI in MM cells and assembled the TIMLIP screening list (‘Translation Initiation Mode Litmus Paper’) in order to establish an objective, high throughput assay for distinguishing between the initiation factors' imprints. Indeed, the TIMLIP revealed that there are distinct and significant differences between eIF4E/eIF4GI TI modes.
One of the TIMLIP facets consisted of changes in transcription factors’ targets at the mRNA level. As in the MM model here too we assayed four targets for each TI factor using qPCR (eIF4E: DOCK4, IL23A, EGR1, SERPINI; eIF4GI: FBXO32, GAB2, BCL2, ASS1). Interestingly, all assayed transcription factors' targets were significantly affected in the same manner as observed in our MM model (Table 1). Again, we demonstrated a differential imprint of eIF4E and eIF4GI silencing.
Discussion
In this study, we systematically addressed the expression levels of a panel of TI factors in NSCLC cells (cell lines and patients) in comparison with normal lung cells. Our findings demonstrated that eIF4E; eF4GI, DHX29, and eIF4B are significantly overexpressed in this malignancy and unsurprisingly most elevated in the cell lines, which typically represent the more advanced disease. To the best of our knowledge, this is the first report to describe elevated expression levels and function of eIF4GI, eIF4B, and DHX29 in NSCLC.
Next, we investigated the significance of this increased expression by silencing eIF4E and eIF4GI in select NSCLC cell lines. Our major findings depict a considerable effect on the cells' differentiation (EMT) and migration. Further influence was registered in the altered expressions of select microRNAs and transcription factors, all noted for their role in cell phenotype and fate.52, 53, 54, 55 We further corroborated the utility and feasibility of TI inhibition by the application of the clinically employed eIF4E inhibitor and anti-viral drug ribavirin.35, 36, 37, 38 Generally, all of our observations are in sync with published data. It was reported that elevated eIF4E expression is significantly associated with short survival of NSCLC patients57 and eIF4GI was shown to correspond with cancer progression in other types of cancers.58 Moreover, In vitro experiments with eIF4E-specific antisense oligonucleotides (ASOs) effectively inhibited the growth of cancer xenografts in mice with minimal toxicity.59 An additional study that used antisense of eIF4E in Ras-transformed rat embryo fibroblasts showed that cells with reduced levels of eIF4E were less invasiveness and metastatic.60 Taken together, accumulating data argue for the necessity of developing inhibitors of eIF4E for clinical use. Indeed, we showed that the use of ribavirin may afford such a strategy and a comparable observation was also reported by us previously with eIF4E-eIF4GI association inhibitor 4EGI-1.25
The potential of targeting TI factors in lung cancer treatment is also supported by other observations. It is well established that lung cancer patients receive more and more targeted therapies that induce complete disease remissions yet most eventually develop drug resistance. For instance, this is the case with anti-epidermal growth factor receptor (EGFR) therapy that at first provides exceptional benefit, but over time tumors develop resistance, causing recurrence.61 Interestingly, several publications have associated the elevated eIF4E levels in NSCLC cells with acquired erlotinib (EGFR inhibitor) resistance.10, 62 Specifically, they showed that EGFR-TKI- (tyrosine kinase inhibitors) resistant NSCLC cells possess increased capacity of eIF4F assembly and elevated expression of oncogenic proteins known to be regulated by the cap-dependent translation mechanism (eg, HIF1α, c-Myc, and Mcl-1). They have also shown that KD of eIF4E/eIF4G enhanced erlotinib anti-proliferative effect in resistant cells. Unsurprisingly, combination of erlotinib and 4EGI-1 synergistically inhibited cell growth. Finally, another publication demonstrated that EMT pathways are also associated with erlotinib resistance and that 'EGFR-addicted' cancer cell lines undergoing EMT become erlotinib-resistant in vitro.63 Overcoming EMT-associated anti-cancer drug resistance is a major clinical goal and our results mark eIF4E and/or eIF4GI as potential therapeutic targets for treatment of erlotinib-resistant NSCLC patients.
Previously, we assembled an imprint list using an objective, high throughput assay enabling distinction between eIF4E and eIF4GI-silenced MM cells (ie, 'TIMLIP'). Our TIMLIP lists addressed several facets: transcription factors, microRNAs, and phenotype. Here, we assessed the TIMLIP lists in eIF4E/eIF4GI-silenced NSCLC and determined similar trends (Table 1). The similarity of eIF4E/eIF4GI KD outputs in both cancer models underscores the legitimacy of our TIMLIP model and indicates that eIF4E and eIF4GI may have individual influence on the cells' proteome26, 30, 64, 65 despite their recognized association and function in the eIF4F complex.66 Most importantly, our findings support the emerging understanding that the conventional paradigm of cap binding translation should be revised67 and that the eIF4E/eIF4GI TI modes control the cellular ‘translatomes’ (genome-wide pools of translated mRNA).
It is now well established that microRNAs also contribute to proteome design. Indeed, this level of regulation was also modulated in our model of NSCLC eIF4E/eIF4GI KDs. Specifically, elevated levels of MIR-29b were evident and according to published data correspond with migration inhibition. Obviously, this marks MIR-29b as a potential therapeutic target capable of slowing down NSCLC metastasis. Mechanistically, it was shown that MIR-29b is a tumor metastasis suppressor that suppresses NSCLC cell metastasis via MMP2 attenuation.53 Indeed, MIR-29b is downregulated in lung carcinoma tissues68 causing high MMP2 expression and serving as a poor prognostic factor closely related to clinical stage, pathological grade, lymphatic metastasis, and prognosis.69
The benefit of targeting TI lies with its capacity to simultaneously affect the expression of multiple oncogenic proteins to which NSCLC cells are addicted and that converge on proliferative and migratory pathways. Taken together, we propose that this paradigm represents a solid rational for continued research with the hope of developing small molecule inhibitors of TI with anti-NSCLC activity.
Further study is needed to examine additional facets of TIMLIP screening model. Moreover, we are currently conducting studies aimed at uncovering additional and different pathways unique to eIF4E/eIF4GI in NSCLC cells.
References
Sun S, Schiller JH, Spinola M et al. New molecularly targeted therapies for lung cancer. J Clin Invest 2007;117:2740–2750.
Higgins MJ, Ettinger DS . Chemotherapy for lung cancer: the state of the art in 2009. Expert Rev Anticancer Ther 2009;9:1365–1378.
Huang J, Song H, Liu B et al. Expression of Notch-1 and its clinical significance in different histological subtypes of human lung adenocarcinoma. J Exp Clin Cancer Res 2013;32:84.
Steeg PS . Metastasis suppressors alter the signal transduction of cancer cells. Nat Rev Cancer 2003;3:55–63.
Moreno-Bueno G, Portillo F, Cano A . Transcriptional regulation of cell polarity in EMT and cancer. Oncogene 2008;27:6958–6969.
Silvera D, Formenti SC, Schneider RJ . Translational control in cancer. Nat Rev Cancer 2010;10:254–266.
Barnhart B, Simon M . Taking aim at translation for tumor therapy. J Clin Invest 2007;117:2385–2388.
Thornton S, Anand N, Purcell D et al. Not just for housekeeping: protein initiation and elongation factors in cell growth and tumorigenesis. J Mol Med 2003;81:536–548.
Agnelli L, Fabris S, Bicciato S et al. Upregulation of translational machinery and distinct genetic subgroups characterise hyperdiploidy in multiple myeloma. Br J Haematol 2007;136:565–573.
Li Y, Fan S, Koo J et al. Elevated expression of eukaryotic translation initiation factor 4E is associated with proliferation, invasion and acquired resistance to erlotinib in lung cancer. Cancer Biol Ther 2012;13:272–280.
Bauer C, Brass N, Diesinger I et al. Overexpression of the eukaryotic translation initiation factor 4G (eIF4G-1) in squamous cell lung carcinoma. Int J Cancer 2002;98:181–185.
Pettersson F, Del Rincon SV, Emond A et al. Genetic and pharmacologic inhibition of eIF4E reduces breast cancer cell migration, invasion, and metastasis. Cancer Res 2015;75:1102–1112.
Polunovsky V, Rosenwald I, Tan A et al. translational control of programmed cell death: eukaryotic translation initiation factor 4E blocks apoptosis in growth-factor-restricted fibroblasts with physiologically expressed or deregulated Myc. Mol Cell Biol 1996;16:6573–6581.
Lin C-J, Cencic R, Mills J et al. c-Myc and eIF4F are components of a feedforward loop that links transcription to translation. Cancer Res 2008;68:5326–5334.
Ramirez-Valle F, Braunstein S, Zavadil J et al. eIF4GI links nutrient sensing by mTOR to cell proliferation and inhibition of autophagy. J Cell Biol 2008;181:293–307.
Ramirez-Valle F, Braunstein S, Zavadil J et al. eIF4GI links nutrient sensing by mTOR to cell proliferation and inhibition of autophagy. J Cell Biol 2008;181:293–307.
Cenci S, Sitia R . Managing and exploiting stress in the antibody factory. FEBS Lett 2007;581:3652–3657.
Parsyan A, Shahbazian D, Martineau Y et al. The helicase protein DHX29 promotes translation initiation, cell proliferation, and tumorigenesis. Proc Natl Acad Sci USA 2009;106:22217–22222.
Shahbazian D, Parsyan A, Petroulakis E et al. eIF4B controls survival and proliferation and is regulated by proto-oncogenic signaling pathways. Cell Cycle 2010;9:4106–4109.
Zismanov V, Drucker L, Gottfried M . ER homeostasis and motility of NSCLC cell lines can be therapeutically targeted with combined Hsp90 and HDAC inhibitors. Pulm Pharmacol Ther 2013;26:388–394.
Zismanov V, Drucker L, Gottfried M . Combined inhibition of Hsp90 and the proteasome affects NSCLC proteostasis and attenuates cell migration. Anticancer Drugs 2014;25:998–1006.
Meric F, Hunt KK . Translation initiation in cancer: a novel target for therapy. Mol Cancer Ther 2002;1:971–979.
Legrier ME, Yang CP, Yan HG et al. Targeting protein translation in human non small cell lung cancer via combined MEK and mammalian target of rapamycin suppression. Cancer Res 2007;67:11300–11308.
Jacobson BA, Alter MD, Kratzke MG et al. Repression of cap-dependent translation attenuates the transformed phenotype in non-small cell lung cancer both in vitro and in vivo. Cancer Res 2006;66:4256–4262.
Attar-Schneider O, Zismanov V, Drucker L et al. Secretome of human bone marrow mesenchymal stem cells: an emerging player in lung cancer progression and mechanisms of translation initiation. Tumour Biol 2015;37:4755–4765.
Attar-Schneider O, Drucker L, Zismanov V et al. Bevacizumab attenuates major signaling cascades and eIF4E translation initiation factor in multiple myeloma cells. Lab Invest 2012;92:178–190.
Zismanov V, Lishner M, Tartakover-Matalon S et al. Tetraspanin-induced death of myeloma cell lines is autophagic and involves increased UPR signalling. Br J Cancer 2009;101:1402–1409.
Welnowska E, Castello A, Moral P et al. Translation of mRNAs from vesicular stomatitis virus and vaccinia virus is differentially blocked in cells with depletion of eIF4GI and/or eIF4GII. J Mol Biol 2009;394:506–521.
Attar-Schneider O, Pasmanik-Chor M, Tartakover-Matalon S et al. eIF4E and eIF4GI have distinct and differential imprints on multiple myeloma's proteome and signaling. Oncotarget 2015;6:4315–4329.
Attar-Schneider O, Drucker L, Zismanov V et al. Targeting eIF4GI translation initiation factor affords an attractive therapeutic strategy in multiple myeloma. Cell Signal 2014;26:1878–1887.
Braunstein S, Karpisheva K, Pola C et al. A hypoxia-controlled cap-dependent to cap-independent translation switch in breast cancer. Mol Cell 2007;28:501–512.
Kaiser C, Dobrikova EY, Bradrick SS et al. Activation of cap-independent translation by variant eukaryotic initiation factor 4G in vivo. RNA 2008;14:2170–2182.
Shiroki K, Ohsawa C, Sugi N et al. Internal ribosome entry site-mediated translation of Smad5 in vivo: requirement for a nuclear event. Nucleic Acids Res 2002;30:2851–2861.
Baird SD, Turcotte M, Korneluk RG et al. Searching for IRES. RNA 2006;12:1755–1785.
Assouline S, Culjkovic B, Cocolakis E et al. Molecular targeting of the oncogene eIF4E in acute myeloid leukemia (AML): a proof-of-principle clinical trial with ribavirin. Blood 2009;114:257–260.
Matassa DS, Amoroso MR, Agliarulo I et al. Translational control in the stress adaptive response of cancer cells: a novel role for the heat shock protein TRAP1. Cell Death Dis 2013;4:e851.
Issur M, Bougie I, Despins S et al. Enzymatic synthesis of RNAs capped with nucleotide analogues reveals the molecular basis for substrate selectivity of RNA capping enzyme: impacts on RNA metabolism. PLoS ONE 2013;8:e75310.
Tan K, Culjkovic B, Amri A et al. Ribavirin targets eIF4E dependent Akt survival signaling. Biochem Biophys Res Commun 2008;375:341–345.
Hsieh AC, Liu Y, Edlind MP et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 2012;485:55–61.
Robichaud N, del Rincon SV, Huor B et al. Phosphorylation of eIF4E promotes EMT and metastasis via translational control of SNAIL and MMP-3. Oncogene 2015;34:2032–2042.
Ghosh B, Benyumov AO, Ghosh P et al. Nontoxic chemical interdiction of the epithelial-to-mesenchymal transition by targeting cap-dependent translation. ACS Chem Biol 2009;4:367–377.
Smith KA, Zhou B, Avdulov S et al. Transforming Growth Factor-beta1 induced epithelial mesenchymal transition is blocked by a chemical antagonist of translation factor eIF4E. Sci Rep 2015;5:18233.
Christofori G . Changing neighbours, changing behaviour: cell adhesion molecule-mediated signalling during tumour progression. EMBO J 2003;22:2318–2323.
Aigner K, Dampier B, Descovich L et al. The transcription factor ZEB1 (deltaEF1) promotes tumour cell dedifferentiation by repressing master regulators of epithelial polarity. Oncogene 2007;26:6979–6988.
Polakis P . The oncogenic activation of beta-catenin. Curr Opin Genet Dev 1999;9:15–21.
Turner FE, Broad S, Khanim FL et al. Slug regulates integrin expression and cell proliferation in human epidermal keratinocytes. J Biol Chem 2006;281:21321–21331.
Na YR, Seok SH, Kim DJ et al. Bone morphogenetic protein 7 induces mesenchymal-to-epithelial transition in melanoma cells, leading to inhibition of metastasis. Cancer Sci 2009;100:2218–2225.
Drabsch Y, ten Dijke P . TGF-beta signaling in breast cancer cell invasion and bone metastasis. J Mammary Gland Biol Neoplasia 2011;16:97–108.
Phuah NH, Nagoor NH . Regulation of microRNAs by natural agents: new strategies in cancer therapies. Biomed Res Int 2014;2014:804510.
Chen G, Umelo IA, Lv S et al. miR-146a inhibits cell growth, cell migration and induces apoptosis in non-small cell lung cancer cells. PLoS One 2013;8:e60317.
Guo H, Li Q, Li W et al. MiR-96 downregulates RECK to promote growth and motility of non-small cell lung cancer cells. Mol Cell Biochem 2014;390:155–160.
Zhang H, Zhu X, Li N et al. miR-125a-3p targets MTA1 to suppress NSCLC cell proliferation, migration, and invasion. Acta Biochim Biophys Sin (Shanghai) 2015;47:496–503.
Wang H, Guan X, Tu Y et al. MicroRNA-29b attenuates non-small cell lung cancer metastasis by targeting matrix metalloproteinase 2 and PTEN. J Exp Clin Cancer Res 2015;34:59.
Liu GL, Liu X, Lv XB et al. miR-148b functions as a tumor suppressor in non-small cell lung cancer by targeting carcinoembryonic antigen (CEA). Int J Clin Exp Med 2014;7:1990–1999.
Cao M, Seike M, Soeno C et al. MiR-23a regulates TGF-beta-induced epithelial-mesenchymal transition by targeting E-cadherin in lung cancer cells. Int J Oncol 2012;41:869–875.
Li J, Li P, Chen T et al. Expression of microRNA-96 and its potential functions by targeting FOXO3 in non-small cell lung cancer. Tumour Biol 2015;36:685–692.
Zhang B, Zhu C, Chen B et al. [Expression and its clinical significance of eIF4E in non-small cell lung cancer]. Zhongguo Fei Ai Za Zhi 2010;13:1132–1135.
Silvera D, Arju R, Darvishian F et al. Essential role for eIF4GI overexpression in the pathogenesis of inflammatory breast cancer. Nat Cell Biol 2009;11:903–908.
Graff JR, Konicek BW, Vincent TM et al. Therapeutic suppression of translation initiation factor eIF4E expression reduces tumor growth without toxicity. J Clin Invest 2007;117:2638–2648.
Graff JR, Boghaert ER, De Benedetti A et al. Reduction of translation initiation factor 4E decreases the malignancy of ras-transformed cloned rat embryo fibroblasts. Int J Cancer 1995;60:255–263.
Gazdar AF . Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 2009;28:S24–S31.
Bonanno L, Jirillo A, Favaretto A . Mechanisms of acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors and new therapeutic perspectives in non small cell lung cancer. Curr Drug Targets 2011;12:922–933.
Lin Y, Wang X, Jin H . EGFR-TKI resistance in NSCLC patients: mechanisms and strategies. Am J Cancer Res 2014;4:411–435.
Mamane Y, Petroulakis E, Martineau Y et al. Epigenetic activation of a subset of mRNAs by eIF4E explains its effects on cell proliferation. PLoS One 2007;2:e242.
Zismanov V, Attar-Schneider O, Lishner M et al. Multiple myeloma proteostasis can be targeted via translation initiation factor eIF4E. Int J Oncol 2015;46:860–870.
Yanagiya A, Svitkin YV, Shibata S et al. Requirement of RNA binding of mammalian eukaryotic translation initiation factor 4GI (eIF4GI) for efficient interaction of eIF4E with the mRNA cap. Mol Cell Biol 2009;29:1661–1669.
Shatsky IN, Dmitriev SE, Andreev DE et al. Transcriptome-wide studies uncover the diversity of modes of mRNA recruitment to eukaryotic ribosomes. Crit Rev Biochem Mol Biol 2014;49:164–177.
Fabbri M, Garzon R, Cimmino A et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 2007;104:15805–15810.
Qian Q, Wang Q, Zhan P et al. The role of matrix metalloproteinase 2 on the survival of patients with non-small cell lung cancer: a systematic review with meta-analysis. Cancer Invest 2010;28:661–669.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Metastasis underlies non-small cell lung cancer (NSCLC) cancer morbidity and accounts for disease progression. In the current study, the authors demonstrate that inhibition of eIF4E/eIF4GI translation initiation factors reduce migration and epithelial to mesenchymal transition, both essential for metastasis. These findings highlight the potential of translation initiation targeting in NSCLC therapy.
Supplementary information
Rights and permissions
About this article
Cite this article
Attar-Schneider, O., Drucker, L. & Gottfried, M. Migration and epithelial-to-mesenchymal transition of lung cancer can be targeted via translation initiation factors eIF4E and eIF4GI. Lab Invest 96, 1004–1015 (2016). https://doi.org/10.1038/labinvest.2016.77
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2016.77
This article is cited by
-
A CRISPR knockout screen reveals new regulators of canonical Wnt signaling
Oncogenesis (2021)
-
Eukaryotic Translation Initiation Factor 4 Gamma 1 (EIF4G1): a target for cancer therapeutic intervention?
Cancer Cell International (2019)
-
Eukaryotic Translation Initiation Factor 4 Gamma 1 (eIF4G1) is upregulated during Prostate cancer progression and modulates cell growth and metastasis
Scientific Reports (2018)
-
The effect of mesenchymal stem cells' secretome on lung cancer progression is contingent on their origin: primary or metastatic niche
Laboratory Investigation (2018)
-
Fatty Acid Synthase induced S6Kinase facilitates USP11-eIF4B complex formation for sustained oncogenic translation in DLBCL
Nature Communications (2018)