Abstract
We previously showed that histone deacetylase inhibitor (HDACi) and 5-azacytidine (AZA) treatment selectively induced cell death of esophageal cancer cells. The mechanisms of cancer selectivity, however, remained unclear. Here we examined whether the cancer selectivity of HDACi/AZA treatment is mediated by the thioredoxin (Trx) system and reactive oxygen species (ROS) in esophageal cancer cells. For this, we first analyzed human tissue specimens of 37 esophageal cancer patients by immunohistochemistry for Trx, Trx-interacting protein (TXNIP) and Trx reductase (TXNRD). This revealed a loss or at least reduction of nuclear Trx in esophageal cancer cells, compared with normal epithelial cells (P<0.001). Although no differences were observed for TXNIP, TXNRD was more frequently expressed in cancer cells (P<0.001). In the two main histotypes of esophageal squamous cell carcinomas (ESCCs, n=19) and esophageal adenomcarcinomas (EAC, n=16), similar Trx, TXNIP and TXNRD expression patterns were observed. Also in vitro, nuclear Trx was only detectable in non-neoplastic Het-1A cells, but not in OE21/ESCC or OE33/EAC cell lines. Moreover, the two cancer cell lines showed an increased Trx activity, being significant for OE21 (P=0.0237). After treatment with HDACi and/or AZA, ROS were exclusively increased in both cancer cell lines (P=0.048–0.017), with parallel decrease of Trx activity. This was variably accompanied by increased TXNIP levels upon AZA, MS-275 or MS-275/AZA treatment for 6 or 24 h in OE21, but not in Het-1A or OE33 cells. In summary, this study evaluated Trx and its associated proteins TXNIP and TXNRD for the first time in esophageal cancers. The analyses revealed an altered subcellular localization of Trx and strong upregulation of TXNRD in esophageal cancer cells. Moreover, HDACi and AZA disrupted Trx function and induced accumulation of ROS with subsequent apoptosis in esophageal cancer cells exclusively. Trx function is hence an important cellular mediator conferring non-neoplastic cell resistance for HDACi and/or AZA.
Similar content being viewed by others
Main
Esophageal cancer constitutes the two main histotypes of esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC). As epigenetic alterations are frequently detected in both subtypes,1 treatment strategies including epigenetic-acting drugs may be considered as feasible novel therapeutic approach. Indeed, epigenetic modifiers such as histone deacetylases (HDACs) or DNA (cytosine-5)-methyltransferase 1 (DNMT1) are frequently expressed in esophageal cancers.2, 3, 4 Hence, HDAC inhibitors (HDACi) as well as the DNMT1 inhibitor 5-azacytidine (AZA) may be useful tools for targeting esophageal cancer cells. The selectivity of combined HDACi and AZA treatment was recently described by us, whereby the underlying mechanisms were partially addressed by RNA transcriptome analyses.5 These RNA transcriptome data suggested a possible role of the redox environment. Indeed, in lung (cancer) cells selectivity of HDACi was shown before to be mediated by disturbance of the thioredoxin (Trx) system and subsequent increase of reactive oxygen species (ROS).6
Trx is a small 12-kDa protein with thiol reductase activity. The function of Trx is complex and is linked to cell proliferation, DNA synthesis or inhibition of apoptosis.7 Cellular function of Trx depends on its localization in the nucleus or the cytoplasm.8 Trx is also regulating the activity of various transcription factors, including eg, NFκB,8, 9 Nrf-2,10 the glucocorticoid receptor11 or p53.12 This often involves a direct interaction of Trx and the transcription factor in the nucleus.8, 10 Together with Trx reductase (TXNRD) and NADPH, the so called Trx system is formed. In addition, activity of Trx is also regulated by its endogenous inhibitor, the Trx-interacting protein (TXNIP). Moreover, Trx acts as potent antioxidant by scavenging of ROS.13 Trx is overexpressed in different tumor entities such as breast,14 lung,15 gastric16 or colorectal17 cancer. Moreover, Trx overexpression correlated with poorer survival/prognosis16, 18, 19 and was an independent prognostic factor in colorectal cancer.19 However, Trx and its associated proteins TXNIP and TXNRD have not yet been evaluated in esophageal cancers so far.
Thus, we here examined the expression of Trx, TXNRD and TXNIP in human tissue specimens of esophageal cancers. In order to evaluate whether or not ROS induction and regulation of Trx, TXNIP or TXNRD is involved in the esophageal cancer cell selective responses to interference with epigenetic modifiers, they were analyzed in vitro after treatment with HDACi and/or AZA.
MATERIALS AND METHODS
Tissue Specimens of Human Esophageal Cancer
Formalin-fixed and Paraffin-embedded tissue specimens of 37 patients with esophageal cancers undergoing primary resection at the University Medical Center were examined. The study was approved by institutional ethics regulations (no. 435/11; Ethik Kommission, Albert-Ludwigs-Universität Freiburg, Germany). Median age of patients was 60.7 years (range 39–89 years). The female/male ratio was 10/27. All cases were received as primary resection specimens without previous neoadjuvant radiotherapy/chemotherapy. Histological re-classification was according to the WHO Classification of Tumours of the Digestive System.20 This study included 19 ESCC cases, 16 EAC cases, 1 adenosquamous carcinoma and 1 carcinoid.
Immunohistochemistry
Immediate serial sections (3 μm) of human tissue specimens were subjected to deparaffinization, antigen retrieval and immunohistochemical staining with appropiate antibodies (Trx: 1:5000 from abcam/ab133524; TXNIP: 1:5000 from Novus Biologicals/clone JY2; TXNRD: 1:800 from Abcam/ab124954). IHC was evaluated for nuclear as well as cytoplasmic positivity in normal esophageal epithelial cells and invasive tumor cells with the following scores: score 0=negative (<10% positive cells), score 1=weak and/or heterogeneous protein expression (from 10 to 70% positive cells), and score 2=strong expression and homogeneous in >70% cells. For TXNIP and TXNRD, only basal layers were considered for scoring in normal esophageal epithelia.
Cell Culture and Inhibitor Treatment
Esophageal cell lines Het-1A, OE21 and OE33 (European Collection of Cell Cultures, Salisbury, UK) were cultured in GIBCO RPMI 1640 medium (Invitrogen, Karlsruhe, Germany) supplemented with 10% FBS and 2 mM GIBCO L-Glutamine (Invitrogen) at 37 °C in a 5% CO2 atmosphere as described before.5, 21 All three cell lines were verified by DNA fingerprinting/STR analysis for authenticity via the Leibniz-Institute DSMZ (July 2013; data available upon request). All drugs were purchased from Selleckchem (Munich, Germany) and dissolved in DMSO. For inhibitor treatment, cells were seeded overnight and inhibitor or vehicle (DMSO) was added on the next day. Cells were then incubated for the indicated time points.
Indirect Immunofluorescence
As reported by us before,22 cells were incubated with appropriate antibodies (Trx: 1 : 250 from Abnova/H00007295-M01; TXNIP: 1:120 from Abcam/ab86983; TXNRD: 1:100 from Abcam/ab124954) overnight at 4 °C. On the next day, cells were washed in PBS and incubated with 1:200 diluted fluorescently labeled secondary antibodies from Invitrogen for 1 h at RT. Finally, cells were rinsed with PBS, and cell nuclei were counterstained with DAPI (Vector Laboratories, Burlingame, USA).
Immunoblotting
Immunoblotting was performed as described before.22 Following antibodies and dilutions were used: Trx: 1:5000 from abcam/ab133524, TXNIP: 1:1000 from Novus Biologicals/clone JY2 and TXNRD: 1:10 000 from Abcam/ab124954. Bands were quantified by densitometry with ImageJ Version 1.46r.
Quantification of Trx Activity
Activity of Trx was measured by the Thioredoxin Activity Fluorescent Assay Kit (FkTRX-02, IMCO, Stockholm) according to the manufacturer’s instruction. For this, 2.5 × 105 cells were seeded into six-well plates and processed on the next day. To evaluate the impact of HDACi on Trx activity, cells were treated with vehicle/DMSO or HDACi for 24 h. Data are presented normalized to non-neoplastic Het-1A cells or vehicle (=DMSO)-treated cells for HDACi experiments.
Quantification of ROS
Cells were incubated for 24 h in phenol red-free RPMI with vehicle or HDACi/AZA before quantification of ROS. Staining of ROS was performed as followed: cells were trypsinized, pelleted, and resuspended in phenol red-free RPMI containing 1 μM CM-H2DCFDA (Invitrogen). After incubation for 30 min at 37 °C, cells were washed once in PBS and then resuspended in 500 μl phenol red-free RPMI. Stained cells were then analyzed by flow cytometry. Data are presented as the percentage of 10 000 recorded cells.
Quantification of Apoptosis
Apoptosis was assessed 72 h after inhibitor addition, as described before by us.21 Cells were trypsinized, pooled with the cell culture medium fraction and stained by FITC Annexin V Apoptosis Detection Kit I (BD Pharmingen, Heidelberg, Germany) according to the manufacturer’s instruction. Data are presented as the percentage of 10 000 recorded cells.
Quantitative RT-PCR
Total RNA isolation and cDNA synthesis was performed as described before by us.5 qRT-PCR was performed using established protocols23 and mRNA expression levels were calculated by the comparative Ct method, normalized to the housekeeping gene β-actin.5 Following primers were used for TXNIP: forward 5′-GGCTAAAGTGCTTTGGATGC-3′ and reverse 5′-TGATCACCATCTCATTCTCACC-3′.
Detection of Acetylated TXNIP by Immunoprecipitation
After 24 h incubation with MS-275 and/or AZA, cells were washed with PBS and lysed with RIPA containing cOmplete ultra protease inhibitor cocktail (Roche, Basle, Switzerland). Protein concentrations were determined by the DC Protein Assay (Bio-Rad, Munich, Germany). For each IP reaction, 100 μg protein were subjected to incubation at 4 °C under rotary agitation overnight with either anti-acetyl-lysine antibody (Cell Signaling Technology, Danvers, USA; 1:100 dilution, corresponding to 0.449 μg antibody per 100 μl IP reaction) or IgG. On the next day, protein–antibody complexes were captured with Protein-G Dynabeads (Invitrogen). After three washes with 0.1%Tween/PBS, bead–antibody complexes were denatured in NuPage LDS Sample Buffer (Invitrogen) and analyzed by immunoblotting via anti-TXNIP. Bands were analyzed by densitometry with ImageJ Version 1.46r. Data are represented as the percentage of control (vehicle/DMSO=100%).
Proximity Ligation Assay (PLA)
To visualize acetylated TXNIP, cells were seeded onto eight-well culture slides and incubated with MS-274 and/or AZA for 24 h. PLA was performed according to our previously published protocol.21 Following antibodies were used for anti-acetyl-lysine (rabbit, Cell Signaling Technology) and for anti-TXNIP (mouse-clone JY2, Novus Biologicals). Control cells were incubated with only anti-TXNIP antibody or with isotype IgG for control. Cells were imaged (about 100 cells/condition) and PLA signals were quantified using the BlobFinder V3.2. Data are represented as signals/cell.
Statistics
Statistical analysis for data of immunohistochemistry was performed using the software package SPSS v 18 (SPSS, Chicago, USA). Fisher’s exact test, two-sided for proportions, was applied to compare normal and cancer tissues, and P-values<0.05 were considered statistically significant. For all in vitro analyses, at least three independent experiments were performed and in the corresponding diagrams the mean±s.e.m. is shown. Statistical testing for in vitro data was performed by Student’s t-test and obtained P-values were represented as *0.05–0.01, **≤0.01–0.001 and ***≤0.001.
RESULTS
Esophageal Cancers Show a Loss of Nuclear Trx
As Trx, TXNIP and/or TXNRD have not been analyzed in esophageal cancers so far, human tissue specimens of esophageal cancers with case-matched normal esophageal epithelium (NEE) were stained by immunohistochemistry (Figure 1a for IHCs, Table 1, Supplementary Figures S1A–C for quantification and diagrams). The majority of all NEE cells showed strong cytoplasmic (97%, score 2) and nuclear localization (94%, score 2) of Trx. Cytoplasmic localization and expression levels of Trx were maintained in esophageal cancers. In contrast, nuclear expression of Trx in esophageal cancers was reduced in around 60% or lost in 30% of all cases (Fisher’s exact test; P<0.001). Only 11% of tumors showed similar nuclear expression (=score 2) as compared with NEE. Moreover, this loss of nuclear Trx was particularly observed in the transition zones from non-invasive to invasive epithelial cells in ESCC as well as in EAC (Figure 1b).
Esophageal cancer cells show loss of nuclear Trx and higher TXNRD expression. (a) The panels show representative serial sections of stained case-matched normal esophageal epithelium and ESCC or EAC tissue specimens for Trx, TXNIP and TXNRD. Refer to Table 1 for results of all cases. (b) Please note loss of nuclear Trx in transition zones of esophageal cancers. Bars represent 200 μm.
TXNIP was expressed only in the basal layers of NEE and esophageal cancers tended to show reduced staining compared with the basal layers of NEE, with a decrease from 60% down to 37% of all cases for score 2 (Figure 1a and Table 1).
Importantly, the majority of NEE cells (>50%) were completely negative (=score 0; Figure 1a in NEE of ESCC case) for TXNRD and 45% of NEE cells showed only very faint staining in the basal layer (=score 1). However, the majority of cancer cases (80%; score 2) showed a frequent and strong upregulation of cytoplasmic TXNRD expression in tumor cells (Fisher’s exact test; P<0.001). Interestingly, some cases also showed TXNRD expression in the tumor surrounding stromal cells (Figure 1, EAC case). In addition, nuclear TXNRD expression was found exclusively in cancer cells (Supplementary Figure S1C). Regarding the two main histotypes of esophageal cancers, ESCC and EAC cases did not show major differences of expression levels or localization of Trx or TXNIP (see Table 1B). In contrast, the observed strong nuclear TXNRD expression (score 2) was more frequent in EAC (37.5%) than in ESCC (5.2%) cases (see Table 1B and Supplementary Figure S1D for one example), albeit not reaching statistical significance (P=0.064).
In summary, both main histotypes of esophageal cancer showed an alteration of the Trx system with loss of nuclear Trx and strong upregulation of (nuclear) TXNRD in esophageal cancer cells compared with NEE cells.
The Pattern of Trx Expression is Mirrored in Esophageal Cancer Cell Lines
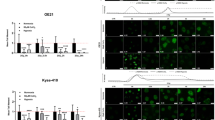
Next, esophageal cell lines were characterized for Trx, TXNIP and TXNRD (Figure 2). Immunofluorescence stainings revealed that non-neoplastic Het-1A cells showed the same expression pattern of Trx and its associated proteins as NEE in situ, with cytoplasmic and nuclear Trx as well as cytoplasmic localization of TXNIP and TXNRD. Interestingly, the esophageal cancer cell lines OE21 (ESCC) and OE33 (EAC) showed no nuclear Trx as observed before in situ (Figure 2a). Comparison of protein levels (Figures 2b and c) revealed a small but significant increase of Trx for OE21 (P=0.00091) and OE33 (P=0.0009). TXNIP was strongly decreased in both cancer cell lines compared with Het-1A (P<0.0001). For TXNRD, no significant differences between Het-1A and esophageal cancer cells were observed. Finally, the baseline Trx activity was measured, revealing higher Trx activity in both cancer cell lines (P=0.02 for OE21) compared with non-neoplastic Het-1A cells (Figure 2d).
Characterization of esophageal cell lines for Trx, TXNIP and TXNRD. (a) Subcellular localization of Trx and its associated proteins TXNIP and TXNRD was analyzed by immunofluorescence stainings. Shown are 3D image stacks and, for Trx, one central image through the nucleus. Note cytoplasmic and nuclear Trx expression in Het-1A cells, but loss of nuclear Trx in both cancer cell lines OE21 and OE33 (see inserts of nuclear sections). Bar represents 20 μm. (b and c) Protein levels of Trx, TXNIP and TNXRD levels were compared by immunoblotting with subsequent (b) quantification by ImageJ. (c) Shown is one representative immunoblot of three independent experiments. Note slight Trx increase and strong TXNIP decrease in both cancer cell lines. (d) Quantification of baseline Trx activity with higher Trx activity in OE21 and, to a lesser extent, also in OE33 compared with non-neoplastic Het-1A cells. Shown is the mean±s.e.m. for three independent experiments. Significance levels are represented as *0.05–0.01 and ***≤0.001.
Thus, these data show that esophageal cancer cell lines mirror the observed in situ situation with loss of nuclear Trx. Moreover, downregulated TXNIP protein levels may explain the observed increased Trx activity in esophageal cancer cell lines.
HDACi Induce ROS Exclusively in Esophageal Cancer Cell Lines
To examine whether ROS accumulation is involved in HDACi selectivity toward esophageal cancer cells,5, 6 ROS levels were quantified 24 h after different HDACi/AZA combinations, using suberoylanilide hydroxamic acid/SAHA as broad HDACi as well as MS-275 and FK228 as selective class I HDACi.
Indeed, non-neoplastic Het-1A cells showed no increased ROS levels after HDACi and/or AZA treatments (Figure 3a). In contrast, both esophageal cancer cell lines OE21/OE33 showed an increased ROS production, mainly after AZA addition, but to a lesser extent also after HDACi. In OE21 cells, ROS levels increased up to 30%, especially for the combination of MS-275 and AZA with slightly enhanced effects compared with single substance treatments (P=0.048, Figure 3b). In OE33 cells, ROS induction was less extensive than in OE21. However, MS-275/AZA showed again the strongest effect with an increase of ROS-positive cells up to 16.5% (P=0.028, Figure 3c).
Reactive oxygen species (ROS) are exclusively increased in esophageal cancer cell lines after HDACi/AZA but not in non-neoplastic cells. (a–c) ROS levels were determined by flow cytometry 24 h after HDACi/AZA treatment. This revealed no changes of ROS levels in non-neoplastic Het-1A cells (a). In contrast, both esophageal cancer cell lines (OE21 (b) and OE33 (c)) showed an increase of ROS-positive cells after HDACi/AZA treatment, especially for combination of MS-275 and AZA. (d) In parallel, at 24 h after MS-275/AZA treatment Trx activity decreased in OE21 and OE33 cells. Shown is the mean±s.e.m. for three independent experiments. Significance levels are represented as *0.05–0.01.
To evaluate whether increased ROS in esophageal cancer cells was linked to apoptosis induction by HDACi/AZA treatment as reported before by us,5 ROS accumulation was attenuated by pretreatment with the ROS scavenger Trolox, a substance proven to be effective as antioxidant in esophageal cancer cells.24 Non-neoplastic Het-1A cells were unaffected by Trolox pretreatment in their response to AZA and/or HDACi, whereas OE21 and OE33 showed slightly decreased ROS levels (Supplementary Figure S2A) as well as slightly reduced apoptosis induction by AZA or to a lesser extent by AZA/HDACi treatment (Supplementary Figure S2B).
These data support a role of ROS in treatment responses of esophageal cancer cells to AZA and/or HDACi treatment.
Trx Activity is Decreased Exclusively in Cancer Cell Lines after MS-275/AZA
As Trx is involved in ROS scavenging,13 the Trx activity was quantified after treatment with MS-275 and/or AZA at 24 h. This time point was chosen in view of our previous experiments, demonstrating increased ROS levels for this time point. For these experiments, treatment by MS-275/AZA showed the strongest effect on ROS levels (see Figures 3a–c). This revealed that both cancer cell lines displayed a decrease of Trx activity after MS-275 and/or AZA (Figure 3d). Thereby, OE21 cells showed a reduction of Trx activity for AZA and MS-275, which was even further reduced upon combination of MS-275/AZA treatment. In OE33 cells, decreased Trx activity was mainly mediated by AZA treatment. In contrast, non-neoplastic Het-1A showed a variable and slight increase of Trx activity.
Regulation of Trx, TXNIP and TXNRD after MS-275/AZA
Different response of normal and transformed cells to HDACi are mediated by different regulation of Trx6 and TXNIP25 in response to HDACi. Thus Trx, TXNIP and TXNRD protein levels were analyzed 6 and 24 h after HDACi/AZA treatment (Figure 4). For this, only MS-275 and/or AZA were analyzed, as this combination showed the strongest induction of ROS. This revealed that Het-1A and OE33 cells showed similar Trx, TXNIP and TXNRD protein levels. However, in OE21 cells, a variable increase of TXNIP was observed 24 h after addition of the MS-275/AZA combination.
TXNIP is increased in OE21 cells after MS-275/AZA treatment. For analysis of Trx, TXNIP and TXNRD levels after MS-275 and/or AZA treatment, proteins were isolated 6 and 24 h after addition of inhibitors and evaluated by (a) immunoblotting with (b) quantification by densitometry. For Het-1A and OE33 cells, no differences for Trx, TXNIP and TXNRD were detectable. However, OE21 cells displayed an increase of TXNIP, especially for combination of MS-275 and AZA after 24 h. Shown is one representative immunoblot for the target and control protein of each the same protein lysate. The bar diagrams represent the mean±s.e.m. for three independent experiments.
To further investigate the mechanism of the TXNIP increase in OE21 cells, TXNIP mRNA levels were analyzed, revealing its downregulation (Supplementary Figure S3A). Thus elevated TXNIP levels may not be explained by increased transcription of its gene.
As protein acetylation interfere with ubiquitination and thus proteasomal degradation,26 we analyzed the acetylation level of TXNIP after MS-275/AZA. For this, immunoprecipitation for acetyl-lysine was performed after MS-275/AZA treatment and the fraction of TXNIP was then quantified by immunoblotting. This revealed a weak increase of TXNIP after MS-275/AZA (Supplementary Figure S3B). Finally, in situ PLAs were performed to detect acetylated TXNIP in OE21 cells in situ. This revealed a five-fold increase of acetylated TXNIP, respective PLA signals, after MS-275/AZA (P=0.01, Supplementary Figures S3C and D).
Thus the increase of TXNIP protein after combination of MS-275 and AZA in OE21 cells was independent of mRNA TXNIP levels and was at least in part mediated by its acetylation.
DISCUSSION
Recently, we revealed that combinations of different HDACi with AZA act selectively on esophageal cancer cell lines.5 However, the mechanisms of this cancer cell selectivity remained unclear. In lung cancer cells, disruption of the Trx system with subsequent ROS induction appears to mediate cancer cell selectivity of HDACi.6 Thus this study analyzed the potential involvement of Trx and ROS in HDACi-based esophageal cancer cell responses.
First, Trx and its associated proteins TXNIP and TXNRD were examined in human tissue specimens of cases with ESCC and EAC, including case-matched NEE. This revealed a loss of nuclear, but preserved cytoplasmic Trx expression in esophageal cancer cells compared with NEE. Indeed, Trx is known to mediate different functions depending on its subcellular localization: for example, in keratinocytes, cytoplasmic Trx blocked degradation of IκB kinase, retarding NFκB in the cytoplasm. In contrast, nuclear Trx enhanced the DNA binding of NFκB and thus its transcriptional activity.8 In addition, Trx upregulated the protease MMP-9 via NFκB, facilitating tumor cell invasion.27 In fact, distinct subcellular localizations of Trx were already described for cancer as compared with normal tissues in other tumor entities.28, 29 In gallbladder carcinomas, patients with nuclear Trx showed shorter survival than patients without nuclear Trx.28 Hence, cancer cells seem to have higher levels of nuclear Trx, which associate with proliferation,29 shorter survival28 and p53 expression.29 However, there are also contradictory findings for nuclear Trx. Patients with ovarian cancer showed better progression-free survival with higher nuclear Trx levels.30 Similar findings have been obtained for early-stage breast cancers.31 Thus nuclear Trx may mediate different cellular effects depending on the tumor entity and thus different pathways of carcinogenesis. Interestingly, the role of Trx in esophageal cancers seems to be different, since the presented data shows a frequent loss of nuclear Trx in cancer cells, which may enhance the cytoplasmic pool of Trx. In the cytoplasm, Trx counteracts apoptosis by directly inhibiting the apoptosis signal-regulating kinase 1 (ref. 32) and therefore (potentially trapped/sequestered) cytoplasmic Trx might protect esophageal cancer cells against cell death. But clearly, the underlying mechanisms of nuclear and cytoplasmic Trx localization in esophageal cancer cells have to be further evaluated.
In addition, TXNRD was highly upregulated in esophageal cancers and here nuclear localization appeared to be a more prominent feature of EACs than ESCCs. Actually, upregulation of TXNRD was also observed in many other entities, such as breast cancer,29 squamous cell carcinoma of the tongue,18 oral squamous cell carcinoma (OSCC)33 and thyroid cancer.34 In OSCC patients, TXNRD expression correlated with lymph node metastasis and clinical stage.33 Thus TXNRD may facilitate carcinogenesis and may therefore be associated with an aggressive phenotype. Actually, TXNRD levels as well as TXNRD activity were increased in drug-resistant cells,35, 36 and high TXNRD levels actually interfered with drug-specific cytotoxic efficacy in vitro.37 In the present study, some esophageal cancer cases also showed TNXRD expression in tumor surrounding stromal cells, which may be related to inflammation. Indeed, similar observations were made for macrophages and reactive fibroblasts in breast cancer.29 Moreover, nuclear expression of TXNRD was frequently found in EAC cases in this study, being the only major difference between the two cancer subtypes. Nuclear TXNRD expression has been also described for breast carcinoma.29, 31 However, functional consequences of nuclear TXNRD are unknown. Its frequent overexpression in cancers may, however, qualify TXNRD as a novel therapeutic target.
As selectivity of HDACi on transformed cells has been reported to be mediated by ROS and disrupted Trx function in lung cells,6 we evaluated this for the first time in esophageal (cancer) cells in vitro.
Interestingly, characterization of esophageal cell lines revealed that both cancer cell lines showed a lack of nuclear Trx compared to non-neoplastic Het-1A cells, thus providing a suitable in vitro cell line system to study the potential link of HDACi/AZA and Trx. In addition, both cancer cell lines showed higher Trx activity than Het-1A cells, which may be explained by the parallel decreased expression of endogenous Trx inhibitor TXNIP, which was also observed in human tissue specimens of esophageal cancer. The TXNRD upregulation, specifically in the nucleus of esophageal cancer cells in vitro, was not as prominent as in situ. This may reflect the selection of the long-term cultured cancer cell lines.
In fact, ROS were induced by different HDACi/AZA combinations exclusively in both esophageal cancer cells, whereas non-neoplastic Het-1A cells were unaffected. Moreover, combination of MS-275 and AZA showed the strongest induction of ROS in both cell lines. Interestingly, ROS induction after HDACi has been frequently described in cancer cells after HDACi.6, 25, 38, 39 Moreover, potentiation of ROS after combination of HDACi/AZA was suggested as a marker for synergistic cytotoxicity.40
Our data support previous reports, which found frequent alteration of Trx or TXNIP after HDACi treatment.6, 25, 38, 39 Interestingly, cancer cells seem to show an upregulation of TXNIP25, 38, 39 and downregulation of Trx,39 whereas normal cells exhibit an increase of Trx6 after HDACi. In the present study, upregulation of TXNIP was only detectable in OE21 cells but not in OE33 cells.
Finally, quantification of Trx activity after MS-275/AZA revealed that both cancer cell lines showed a decreased Trx activity, especially in OE21 cells. In contrast, non-neoplastic Het-1A cells showed an increased activity of Trx.
The in vitro data suggest the following role of Trx/ROS in the cancer cell selective mechanisms of action of HDACi/AZA treatment: in non-neoplastic Het-1A cells, HDACi/AZA treatment triggers a higher Trx activity. This may enable the scavenging of (potential) ROS accumulation after HDACi/AZA treatment, preventing subsequent cell death (=cell survival). In contrast, HDACi/AZA treatment decreases Trx activity in esophageal cancer cells, which limits the Trx-associated mechanisms for controlling ROS accumulation and hence leads to cancer cell death. One potential (further) player for HDACi/AZA-associated loss of Trx activity may be upregulation of TXNIP (as seen in ESCC/OE21 cells), which counteracts Trx activity.
In summary, the present study evaluated Trx and its associated proteins TXNIP and TXNRD for the first time in esophageal cancers. This revealed an altered subcellular localization of Trx and strong upregulation and nuclear localization of TXNRD in cancer cells as compared with normal cells. Our in vitro data further demonstrate Trx function as a central mediator of cancer cell selectivity of HDACi/AZA in esophageal cancer cells. Thereby, the Trx function is disrupted with decreased Trx activity and ROS accumulation after combined HDACi/AZA treatment. In contrast, non-neoplastic cells are protected by increased Trx activity. In conclusion, our study hence provides novel evidence for the involvement of Trx activity in treatment responses of esophageal cancer cells to HDACi/AZA.
References
Ahrens TD, Werner M, Lassmann S . Epigenetics in esophageal cancers. Cell Tissue Res 2014;356:643–655.
Toh Y, Yamamoto M, Endo K et al. Histone H4 acetylation and histone deacetylase 1 expression in esophageal squamous cell carcinoma. Oncol Rep 2003;10:333–338.
Langer R, Mutze K, Becker K et al. Expression of class I histone deacetylases (HDAC1 and HDAC2) in oesophageal adenocarcinomas: an immunohistochemical study. J Clin Pathol 2010;63:994–998.
Zhao SL, Zhu ST, Hao X et al. Effects of DNA methyltransferase 1 inhibition on esophageal squamous cell carcinoma. Dis Esophagus 2011;24:601–610.
Ahrens TD, Timme S, Hoeppner J et al. Selective inhibition of esophageal cancer cells by combination of HDAC inhibitors and Azacytidine. Epigenetics 2015;10:431–445.
Ungerstedt JS, Sowa Y, Xu WS et al. Role of thioredoxin in the response of normal and transformed cells to histone deacetylase inhibitors. Proc Natl Acad Sci USA 2005;102:673–678.
Arner ES, Holmgren A . Physiological functions of thioredoxin and thioredoxin reductase. Eur J Biochem 2000;267:6102–6109.
Hirota K, Murata M, Sachi Y et al. Distinct roles of thioredoxin in the cytoplasm and in the nucleus. A two-step mechanism of redox regulation of transcription factor NF-kappaB. J Biol Chem 1999;274:27891–27897.
Hayashi T, Ueno Y, Okamoto T . Oxidoreductive regulation of nuclear factor kappa B. Involvement of a cellular reducing catalyst thioredoxin. J Biol Chem 1993;268:11380–11388.
Hansen JM, Watson WH, Jones DP . Compartmentation of Nrf-2 redox control: regulation of cytoplasmic activation by glutathione and DNA binding by thioredoxin-1. Toxicol Sci 2004;82:308–317.
Makino Y, Yoshikawa N, Okamoto K et al. Direct association with thioredoxin allows redox regulation of glucocorticoid receptor function. J Biol Chem 1999;274:3182–3188.
Ueno M, Masutani H, Arai RJ et al. Thioredoxin-dependent redox regulation of p53-mediated p21 activation. J Biol Chem 1999;274:35809–35815.
Nordberg J, Arner ES . Reactive oxygen species, antioxidants, and the mammalian thioredoxin system. Free Radic Biol Med 2001;31:1287–1312.
Cha MK, Suh KH, Kim IH . Overexpression of peroxiredoxin I and thioredoxin1 in human breast carcinoma. J Exp Clin Cancer Res 2009;28:93.
Kakolyris S, Giatromanolaki A, Koukourakis M et al. Thioredoxin expression is associated with lymph node status and prognosis in early operable non-small cell lung cancer. Clin Cancer Res 2001;7:3087–3091.
Lim JY, Yoon SO, Hong SW et al. Thioredoxin and thioredoxin-interacting protein as prognostic markers for gastric cancer recurrence. World J Gastroenterol 2012;18:5581–5588.
Berggren M, Gallegos A, Gasdaska JR et al. Thioredoxin and thioredoxin reductase gene expression in human tumors and cell lines, and the effects of serum stimulation and hypoxia. Anticancer Res 1996;16:3459–3466.
Zhu X, Huang C, Peng B . Overexpression of thioredoxin system proteins predicts poor prognosis in patients with squamous cell carcinoma of the tongue. Oral Oncol 2011;47:609–614.
Noike T, Miwa S, Soeda J et al. Increased expression of thioredoxin-1, vascular endothelial growth factor, and redox factor-1 is associated with poor prognosis in patients with liver metastasis from colorectal cancer. Hum Pathol 2008;39:201–208.
Bosman FT, Carneiro F, Hruban RH et al. WHO Classification of Tumours of the Digestive System; Vol 3, 4 (edn). IARC: Lyon, France, 2010.
Fichter CD, Timme S, Braun JA et al. EGFR, HER2 and HER3 dimerization patterns guide targeted inhibition in two histotypes of esophageal cancer. Int J Cancer 2014;135:1517–1530.
Fichter CD, Herz C, Munch C et al. Occurrence of multipolar mitoses and association with Aurora-A/-B kinases and p53 mutations in aneuploid esophageal carcinoma cells. BMC Cell Biol 2011;12:13.
Timme S, Ihde S, Fichter CD et al. STAT3 expression, activity and functional consequences of STAT3 inhibition in esophageal squamous cell carcinomas and Barrett's adenocarcinomas. Oncogene 2014;33:3256–3266.
Whibley CE, McPhail KL, Keyzers RA et al. Reactive oxygen species mediated apoptosis of esophageal cancer cells induced by marine triprenyl toluquinones and toluhydroquinones. Mol Cancer Ther 2007;6:2535–2543.
Ungerstedt J, Du Y, Zhang H et al. In vivo redox state of human thioredoxin and redox shift by the histone deacetylase inhibitor suberoylanilide hydroxamic acid (SAHA). Free Radic Biol Med 2012;53:2002–2007.
Caron C, Boyault C, Khochbin S . Regulatory cross-talk between lysine acetylation and ubiquitination: role in the control of protein stability. Bioessays 2005;27:408–415.
Farina AR, Cappabianca L, DeSantis G et al. Thioredoxin stimulates MMP-9 expression, de-regulates the MMP-9/TIMP-1 equilibrium and promotes MMP-9 dependent invasion in human MDA-MB-231 breast cancer cells. FEBS Lett 2011;585:3328–3336.
Nagano M, Hatakeyama K, Kai M et al. Nuclear expression of thioredoxin-1 in the invasion front is associated with outcome in patients with gallbladder carcinoma. HPB (Oxford) 2012;14:573–582.
Turunen N, Karihtala P, Mantyniemi A et al. Thioredoxin is associated with proliferation, p53 expression and negative estrogen and progesterone receptor status in breast carcinoma. APMIS 2004;112:123–132.
Woolston CM, Deen S, Al-Attar A et al. Redox protein expression predicts progression-free and overall survival in ovarian cancer patients treated with platinum-based chemotherapy. Free Radic Biol Med 2010;49:1263–1272.
Woolston CM, Storr SJ, Ellis IO et al. Expression of thioredoxin system and related peroxiredoxin proteins is associated with clinical outcome in radiotherapy treated early stage breast cancer. Radiother Oncol 2011;100:308–313.
Saitoh M, Nishitoh H, Fujii M et al. Mammalian thioredoxin is a direct inhibitor of apoptosis signal-regulating kinase (ASK) 1. EMBO J 1998;17:2596–2606.
Iwasawa S, Yamano Y, Takiguchi Y et al. Upregulation of thioredoxin reductase 1 in human oral squamous cell carcinoma. Oncol Rep 2011;25:637–644.
Lincoln DT, Al-Yatama F, Mohammed FM et al. Thioredoxin and thioredoxin reductase expression in thyroid cancer depends on tumour aggressiveness. Anticancer Res 2010;30:767–775.
Marzano C, Gandin V, Folda A et al. Inhibition of thioredoxin reductase by auranofin induces apoptosis in cisplatin-resistant human ovarian cancer cells. Free Radic Biol Med 2007;42:872–881.
Liu JJ, Liu Q, Wei HL et al. Inhibition of thioredoxin reductase by auranofin induces apoptosis in adriamycin-resistant human K562 chronic myeloid leukemia cells. Pharmazie 2011;66:440–444.
Eriksson SE, Prast-Nielsen S, Flaberg E et al. High levels of thioredoxin reductase 1 modulate drug-specific cytotoxic efficacy. Free Radic Biol Med 2009;47:1661–1671.
Lee JH, Jeong EG, Choi MC et al. Inhibition of histone deacetylase 10 induces thioredoxin-interacting protein and causes accumulation of reactive oxygen species in SNU-620 human gastric cancer cells. Mol Cells 2010;30:107–112.
Butler LM, Zhou X, Xu WS et al. The histone deacetylase inhibitor SAHA arrests cancer cell growth, up-regulates thioredoxin-binding protein-2, and down-regulates thioredoxin. Proc Natl Acad Sci USA 2002;99:11700–11705.
Gao S, Mobley A, Miller C et al. Potentiation of reactive oxygen species is a marker for synergistic cytotoxicity of MS-275 and 5-azacytidine in leukemic cells. Leuk Res 2008;32:771–780.
Acknowledgements
We thank the expert technical support of Fabian Beier and Anja Schöpflin. The study is part of a project grant from the Deutsche Forschungsgemeinschaft (SFB992 project C3 to SL). SL and MW thank the Mushett Family Foundation (Chester, NJ, USA) for supporting long-term collaboration on gastrointestinal cancers with the Department of Pathology at MSKCC, NY, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Histone deacetylase inhibitor (HDACi) and 5-azacytidine (AZA) treatment selectively induce cell death of esophageal cancer cells. This study reveals that thioredoxin and reactive oxygen species are involved in cancer cell-selective responses to HDACi and AZA treatment in these cells. In situ and in vitro analyses disclose altered regulation of thioredoxin expression, localization and activity.
Supplementary information
Rights and permissions
About this article
Cite this article
Ahrens, T., Timme, S., Ostendorp, J. et al. Response of esophageal cancer cells to epigenetic inhibitors is mediated via altered thioredoxin activity. Lab Invest 96, 307–316 (2016). https://doi.org/10.1038/labinvest.2015.148
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2015.148
This article is cited by
-
Droxinostat sensitizes human colon cancer cells to apoptotic cell death via induction of oxidative stress
Cellular & Molecular Biology Letters (2018)
-
Metabolic inhibitors accentuate the anti-tumoral effect of HDAC5 inhibition
Oncogene (2017)