Abstract
Overnutrition is the major cause of nonalcoholic fatty liver disease (NAFLD) and its advanced form nonalcoholic steatohepatitis (NASH). We aimed to develop and characterize a murine model, which resembles both the pathology and nutritional situation, of NASH patients in Western societies. Mice were fed with a NASH-inducing diet (ND) containing sucrose, cholesterol and fats rich in saturated fatty acids in a composition, which mimics Western food. After 12 weeks, ND-fed mice revealed obesity and impaired glucose tolerance. In the liver, ND-feeding led to marked steatosis, hepatocellular damage, inflammation and beginning fibrosis. Transcriptome-wide gene expression analysis and search for over-represented transcription factor target sites among the differentially expressed genes identified activator protein-1 (AP-1) as the most likely factor to cause the transcriptional changes in ND livers. Combining differentially expressed gene and protein–protein interaction network analysis identified c-Jun as hub in the largest connected deregulated sub-network in ND livers. Accordingly, ND livers revealed c-Jun-phosphorylation and nuclear translocation. Moreover, hepatic c-Jun expression was enhanced in ND-fed mice. Combined tissue microarray technology and immunohistochemical analysis confirmed enhanced hepatic c-Jun levels in NAFLD patients, which correlated with inflammation, and notably, with the degree of hepatic steatosis. In summary, our new mouse model shows important pathological changes also found in human NASH and indicates c-Jun/AP-1 activation as critical regulator of hepatic alterations. Abundance of c-Jun in NAFLD likely facilitates development and progression of NASH.
Similar content being viewed by others
Main
Nonalcoholic fatty liver disease (NAFLD) is considered as the hepatic manifestation of the metabolic syndrome. In the majority of patients, NAFLD is associated with metabolic risk factors such as obesity, insulin resistance and dyslipidemia. Early stages of NAFLD are histologically categorized into simple steatosis and nonalcoholic steatohepatitis (NASH). Simple steatosis is defined as the presence of hepatic steatosis with no evidence of further hepatocellular injury (eg, ballooning of the hepatocytes). NASH is defined as the presence of hepatic steatosis and inflammation with hepatocellular injury. NASH may progress to fibrosis, cirrhosis and hepatocellular carcinoma.1 The prevalence of NAFLD in Western society is 20–30% and particularly because of a sedentary lifestyle and overnutrition, its incidence is still rising. In addition to the quantity of consumed calories, certain nutrients and food compounds have profound effects on development and/or progression of NAFLD.2
Pure high-fat diets result in hepatic steatosis as well as obesity and insulin resistance in rodents, but do not reliably induce significant hepatic inflammation and fibrosis.3 In addition to the amount, also the type of fat critically affects NAFLD progression. Fat predominantly composed of saturated fatty acids, which are frequent in Western diets, exhibits more adverse health effects than fat types with a higher amount of mono- or polyunsaturated fatty acids.4 Further, also cholesterol has a crucial role in NASH development.5 As reviewed by Farrell et al5, free cholesterol accumulation in the liver is one of the few reliable factors, which discriminate steatohepatitis from simple fatty liver. In addition to lipids, also the impact of fructose on NAFLD development and progression gets increasing attention.2, 6 Fructose is mainly consumed with added sugars (sucrose and high fructose corn syrup), and represents up to 10% of the total energy intake in Western countries.6 Fructose consumption induces insulin resistance and hepatic steatosis as well as steatohepatitis in rodents, and epidemiological studies revealed an association between fructose intake and NAFLD development and progression in men.7
The aim of this study was to establish a dietary murine NASH model, which closely mimics the nutritional situation of NAFLD patients in Western countries. We applied a diet enriched with sucrose, cholesterol and a high concentration of fats, rich in saturated fatty acids in a composition, which mimics Western food. Feeding of this diet caused obesity, insulin resistance and significant hepatic steatosis, inflammation and fibrosis in mice. The mouse model faithfully reproduced most alterations of hepatic gene expression that were previously observed in humans. Surprisingly, we found enhanced c-Jun levels, and this observation was confirmed in human tissues, too. The activity of c-Jun N-terminal kinases (JNKs) and herewith c-Jun activation have been implicated in NASH progression,8, 9 but abundance of c-Jun in NAFLD has not been describe before.
MATERIALS AND METHODS
Animals and Dietary Model of NASH
Male C57BL/6N mice were purchased from Charles River Laboratories (Sulzfeld, Germany) at 12 weeks of age and housed in a 22 °C controlled room under a 12-h light–dark cycle with free access to food and water. After 2 weeks of acclimatization, mice were fed either with standard diet (control) or a NASH-inducing diet (ND), enriched with pork lard (15%), beef tallow (15%), palmitic acid (4%), stearic acid (4%), cholesterol (0.2%) and sucrose (30%). Both chows were prepared by Ssniff (Soest, Germany). After 12 weeks of feeding, animals were killed by heart puncture under deep ketamine/xylazine anesthesia, and blood samples and liver tissues were collected for further analyses. All procedures followed the University Regensburg guidelines for the care and use of laboratory animals.
Quantification of Serum Adipokines
Adipokines in murin serum samples were measured by enzyme-linked immunosorbent assay (ELISA) analysis using the adiponectin and leptin DuoSet ELISA Development kits from R&D Systems (Minneapolis, MN, USA) according to the manufacturer’s instructions.
Glucose Tolerance Test
After 12-h fasting, mice were injected intraperitoneally with glucose (50% w/v solution, 6 μl/kg body weight). Blood glucose concentrations were measured in samples taken from the tail vein before and 15, 30, 60 and 90 min after glucose administration using an accutrend glucometer (Roche, Mannheim, Germany). For quantitative evaluation of the blood glucose curve, the area under the curve was determined.
Western Blotting
Protein extraction and western blotting analysis were performed as described10 applying anti-rabbit antibodies against phospho-AKT (#4058), AKT (#9272), phospho-p65 (#3033), phospho-JNK (#9251), c-Jun (#9165) or phospho-c-Jun (#3270), all from Cell Signaling Technology (Danvers, MA, USA; all diluted 1:1000). Furthermore, antibodies against alpha smooth muscle actin (α-SMA) (ab5694 from Abcam, Cambridge, UK; 1:1000) or actin (MAB1501 from Merck Millipore, Billerica, MA, USA; 1:500) were applied.
Quantification of Hepatic Lipids
Hepatic lipids were extracted using the method of Bligh and Dyer with slight modifications.11 Hepatic triglyceride, free fatty acid (FFA) and cholesterol levels were quantified using the GPO-triglyceride kit (Sigma, Deisenhofen, Germany), the cholesterol/cholesteryl ester quantification kit (BioVision, Mountain View, CA, USA) and the FFAs half micro test (Roche), respectively, as described.12, 13
Quantitative Real-Time-PCR Analysis
RNA isolation from liver tissue and reverse transcription were performed as described.14 Quantitative real-time-PCR was performed applying LightCycler technology (Roche)14 and the following pairs of primers: murine Mcp-1 (forward: 5′-TGGGCCTGCTGTTCACA-3′; reverse: 5′-TCCGATCCAGGTTTTTAATGTA)-3′, murine collagen type I (forward: 5′-CGGGCAGGACTTGGGTA-3′; reverse: 5′-CGGAATCTGAATGGTCTGACT-3′), murine cyclin D1 (forward: 5′-GACACCAATCTCCTCAACGAC-3′; reverse: 5′-TCCTCTTCGCACTTCTGCCTC-3′) and human c-Jun (forward: 5′-TTGCCAGAGCCCTGTTGCGG-3′; reverse: 5′-AAGGTCCGCTCTCGGACGGG-3′). All other mRNA expression analyses were performed using QuantiTect Primer Assays according to the manufacturer’s instructions (Qiagen, Hilden, Germany). Amplification of cDNA derived from 18s rRNA (forward: 5′-AAACGGCTACCACATCCAAG-3′; reverse: 5′-CCTCCAATGGATCCTCGTTA-3′) was used for normalization.
Hepatic Hydroxyproline Content
Hepatic hydroxyproline content was quantified as described previously.13 Briefly, liver tissues (200 mg) were hydrolyzed in 6 M HCl at 110 °C for 16 h. In all, 50 μl of each sample were incubated with chloramine T (2.5 mM) for 5 min and Ehrlich’s reagent (410 mM) for 30 min at 60 °C. Absorption was measured in triplicates at 560 nm.
Immunohistological Analysis
For immunohistological analysis, tissue sections (5 μm) of paraffin-embedded tissue specimens were deparaffinized with xylene and stained as described previously10 applying the following antibodies: anti-cJun (#9165), anti-phospho-cJun (#3270; both from Cell Signaling Technology, diluted 1:100), anti-α-SMA actin (ab5694 from Abcam, diluted 1:200), anti-Ki67 (ab16667 from Abcam, diluted 1:50), anti-CD3 (C7930 from Sigma-Aldrich, diluted 1:500), anti-active caspase-3 (1476-1 from Epitomics, Burlingame, CA, USA; diluted 1:100) and anti-4 hydroxynonenal (HNE11-S from Alpha Diagnostics Int., San Antonio, TX, USA; diluted 1:500).
Microarray Gene Expression Analysis
The Mouse Gene 1.1 ST array from Affymetrix (Affymetrix, High Wycombe, UK) was used to assess transcriptome-wide expression profiles of four mice fed with the ND and four mice fed with regular diet. Microarrays were hybridized and scanned following the standard protocol of the Center for Fluorescent Bioanalytics (KFB; Regensburg, Germany).
Normalization of raw intensity values from CEL files was performed using variance stabilization,15 and the median polish was used to summarize individual probes to one relative expression level per gene. Genes were defined using a custom chip description file based on entrez gene identifiers (version 13.0.0).16 Raw intensities and normalized gene expression data were publicly available at the NCBI Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/) under accession GSE52748. Differential gene expression between ND-fed and control mice was estimated using limma.17 Microarray weights according to Ritchie18 were applied for analysis of differential gene expression, giving higher weights to microarrays, which better fit the linear model for individual genes. Genes with a false discovery rate (FDR) below 0.01 were considered highly significant differentially expressed and genes with an FDR <0.05 were considered significant differentially expressed. FDR values are denoted as q-values in the text. All analyses were performed within the statistical programming environment R19 and using Bioconductor packages.20
Gene Set Analysis and Network Analysis
Gene Set Analysis was performed using the hypergeometric test within the Bioconductor package HTSanalyzeR.21 Genes meeting the FDR threshold of 0.01 were selected for testing significant enrichment of Gene Ontology (GO) terms.22
To search for over-represented transcription factor target sites among the highly significant differentially expressed genes, mouse genes were first mapped to human genes using the homology mapping from NCBI (http://www.ncbi.nlm.nih.gov/projects/homology/maps) and then the transcription factor-binding sites collection from the MSigDB database (collection C3, transcription factor targets) was used for hypergeometric testing within HTSanalyzeR.23 For both GO term and transcription factor-binding site enrichment, terms and binding sites with an FDR<0.05 were considered significant. Motives that cannot be assigned to any known transcription factor were excluded.
To integrate differentially expressed genes with protein–protein interactions, the Bioconductor package BioNet was used.24 Mouse gene symbols were mapped to human gene symbols as before to allow using the human interactome from the BioGRID data base.25 BioNet was then used to find the highest-scoring protein–protein interaction subnetwork from the significant differentially expressed genes with an absolute log2 fold change larger than 0.5.
Human Tissues and Tissue Microarray (TMA)
A TMA was constructed out of 110 paraffin-embedded human hepatic tissue specimens with different degree of steatosis and inflammation.26 All cases were reviewed by a surgical pathologist (AH). A modified histological NAFLD/NASH score according to the recommendations of the Nonalcoholic Steatohepatitis Clinical Research Network was used.27 Briefly, four histological features were evaluated semiquantitatively: steatosis (0–3), lobular inflammation (0–3) and hepatocellular ballooning (0–2). Steatohepatitis was diagnosed when the sum of the scores was 4 or more.
In addition to formalin-fixed liver tissue for TMA construction, 24 liver specimens, which had been immediately snap frozen after surgical resection and stored at −80 °C, were available.
Human liver tissue was obtained and experimental procedures were performed according to the guidelines of the charitable state controlled foundation HTCR, with the informed patients’ consent, and the study was approved by the local ethics committee of the University Regensburg.
Cells, Cell Culture and In Vitro Experiments
Primary human hepatocytes (PHHs) and hepatoma cell lines HepG2 (ATCC HB-8065) and Hep3B (ATCC HB-8064) were isolated and/or cultured as described previously.28, 29 Under the same conditions, cells were stimulated with indicated concentrations of the FFA palmitic acid for 24 h using a specifically prepared palmitic acid/bovine serum albumin (BSA) stock solution. Preparation of the palmitic acid/BSA stock solution was carried out as described previously.30 Briefly, a 100 mM palmitic acid stock solution (Cat#P0500; Sigma) was prepared in 0.1 mM NaOH by heating at 70 °C. A 10% (w/v) FFA-free BSA (Sigma) solution was prepared in ddH2O and maintained at 55 °C in a water bath. In all, 10 mM FFA/1% BSA solution was obtained by complexing the appropriate amount of palmitic acid stock solution to 10% BSA at 55 °C for another 30 min. The solution was then cooled to 25 °C, filter sterilized and stored at −20 °C until use. For siRNA-mediated gene knockdown experiments, Hep3B hepatoma cells were transfected 48 h before FFA treatment applying the HiPerfect transfection method (Qiagen) using JUN-siRNA (cat.# SI00300580 and SI00034678) or AllStar control-siRNA (cat.# 1748119) from Qiagen.
Statistical Analysis
Statistical analysis of all other experiments besides the microarray data was performed as follows: values are presented as mean±s.e.m. Comparison between groups was made using the Student’s unpaired t-test. Welch’s correction was performed if required. Contingency table analysis and the two-sided Fisher’s exact test were used to study the statistical association between clinicopathological and immunohistochemical variables. A P-value <0.05 was considered statistically significant. Calculations were performed using SPSS version 10.0 (SPSS, Chicago, IL, USA) and GraphPad Prism Software (GraphPad Software, Inc., San Diego, CA, USA).
RESULTS
Effect of the ND on Obesity and Other Elements of the Metabolic Syndrome
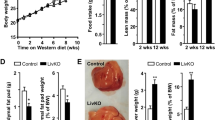
Feeding the ND led to significant body weight gain over time (Figure 1a), and after 12 weeks ND-fed mice appeared obese and showed a marked enlargement of both visceral and subcutaneous white adipose tissue compared with mice fed with standard control chow (Figure 1b). Serum concentration of leptin was significantly elevated, whereas adiponectin levels were moderately decreased in ND-fed compared with control mice (Figure 1c) mimicking an adipokine serum profile associated with obesity in humans.31 Furthermore, fasting blood glucose levels were elevated (Figure 1d), whereas glucose tolerance was impaired in ND-fed mice (Figure 1e) pointing to manifest insulin resistance. In line with this, decreased hepatic AKT phosphorylation in ND-fed mice compared with control mice (Figure 1f) indicated an impaired insulin response, a key pathological factor for NAFLD development and progression also in patients.32
Effect of the ND on obesity elements of the metabolic syndrome. Mice were fed with standard chow (CTR) or a ND. (a) Body weight control (CTR) and ND-fed (ND) mice during the experimental feeding period of 12 weeks. (b) Left panel: representative pictures of a control and a ND-fed mouse after 12 weeks of feeding. Right panel: laparotomy showing massively increased visceral (upper pictures) and subcutaneous (lower pictures) white adipose tissue depots of ND-fed compared with control mice. (c) Serum levels of adiponectin and leptin in control and ND-fed mice. (d) Fasting blood glucose levels of control and ND-fed mice. (e) Area under the curve (AUC) derived from a glucose tolerance test. (f) Western blot analysis of phosphorylated (Ser473) AKT in livers from control and ND-fed mice (left panel). Densitometric quantification of western blot data. (*P<0.05 compared with CTR).
Effect of the ND on Hepatic Lipid Metabolism and Oxidative Stress
ND-fed mice revealed remarkable hepatomegaly (Figure 2a) and a significantly increased liver to body weight ratio compared with control mice (7.5±1.5% vs 5.0±0.47%; P=0.02). Macroscopic liver appearance was indicative for steatosis (Figure 2a), and assessment of hepatic triglycerides confirmed significantly elevated levels in ND-fed mice (956±117 vs 24±11 mg/g protein; P<0.001). Histological analysis showed mixed micro- and macrovesicular steatosis (Figure 2b). Quantitative qPCR analysis revealed distinct alterations of the expression of several genes known to have a critical role in hepatic lipid metabolism in response to ND-feeding (Table 1) similar to pathological changes observed in NASH patients. Thus, the ND-diet induced a marked increase of expression of genes encoding proteins responsible for FFA uptake, binding and transport (Cd36, Fabp1), whereas the expression of genes involved in FA (de novo) synthesis (Acaca, Fasn, Scd1) were only slightly increased. These findings are similar as described in NASH patients26, 33 and indicate that significantly higher FFA levels in ND-fed mice compared with control mice (31.4±2.7 vs 23.9±2.4 μEq/g protein; P=0.007) are mainly caused by enhanced hepatic FFA uptake. Similar to human NASH,34 ND-feeding lead to a significant upregulation of expression of genes involved in mitochondrial (Acadl, Cpt1), peroxisomal (Acox1) and microsomal (Cyp2e1, Cyp4a10) FFA oxidation, which are known to cause increased reactive oxygen species (ROS) formation. Accordingly, a marked increase of hepatic Nrf2, Hmox1 and Nqo1 expression levels was detected in ND-fed mice (Figure 2c), which depicts an endogenous defense mechanism to oxidative stress. In accordance, immunohistochemical analysis for proteins conjugated with the reactive aldehyde 4-hydroxynonenal demonstrate ongoing processes of lipid peroxidation (Figure 2d). Peroxisome proliferator-activated receptor alpha is a strong regulator of hepatic energy combustion, while PPARγ has been shown to induce lipid synthesis and storage. Expression of both transcription factors was enhanced in ND-fed mice (Figure 2e) similar as described in human NASH35 indicative for a marked activation of the overall hepatic lipid metabolism.
Effect of the ND on hepatic lipid metabolism and oxidative stress. Mice were fed with standard chow (CTR) or a ND for 12 weeks. (a) Macroscopic and (b) histological (hematoxylin/eosin staining) images (of livers from control and ND-fed mice. (c) Measurement of hepatic Nrf2, Hmox1 and Nqo1 mRNA expression in control and ND-fed mice by quantitative RT-PCR. (d) Immunohistochemical staining of 4-hydroxynonenal (4-HNE) in liver sections from control and ND-fed mice. (e) Measurement of hepatic PPARα and PPARγ mRNA expression in livers from control and ND-fed mice by quantitative RT-PCR. (*P<0.05 compared with CTR).
It is increasingly recognized that also cholesterol accumulation in the liver has a role in the progression of NASH,5 and coherent with dietary cholesterol supplementation we found elevated hepatic cholesterol levels in ND-fed mice (21.1±4.9 vs 11.6±1.0 mg/g protein; P=0.01). In contrast, expression of genes involved in cholesterol de novo synthesis (Hmgcr, Cyp51) was reduced, whereas genes of cholesterol efflux (Abcg1, Abcg5, Abcg8) where slightly increased in livers of ND-fed mice (Table 1). Transcription of genes responsible for cholesterol conversion (into bile acids; Cyp7a1) or bile acid transportation (Abcb11, Slc10a1) was not significantly altered (Table 1).
Effect of the ND on Hepatocellular Damage and Inflammation
Enhanced hepatic cholesterol levels and oxidative stress have been shown to induce hepatocellular damage and apoptosis,36, 37 which are also present in patients with NASH.38 Also ND-fed mice showed increased alanine aminotransferase serum levels (122±41 vs 41±6 U/l; P=0.01) and elevated hepatocellular caspase-3 activity (Figure 3a) compared with control-fed mice. Increased hepatocellular damage was paralleled by increased regenerative proliferation as shown by an increase of Ki67-positive nuclei (Figure 3b) and increased hepatic cyclin D1 expression (Figure 3c). Furthermore, ND-feeding markedly induced expression of proinflammatory cytokines and chemokines (Figure 3e), as well as hepatic infiltration by inflammatory cells (Figure 3d). Elevated expression of NADPH oxidase complex genes, namely Ncf1, Cyba and Cybb, indicated high activity of macrophages and neutrophils in livers of ND-fed mice (Figure 3f), which presumably contributes to the observed increase of oxidative stress because of NADPH oxidase-mediated ROS formation. Oxidative stress is a strong inducer of the NFκB pathway, a regulator of proinflammatory gene expression and critical factor for the progression from simple steatosis to NASH. In line with this and similar as described in NASH patients,39 elevated levels of phosphorylated p65 indicated increased NFκB activation in the livers of ND-fed mice (Figure 3g). In accordance, NFκB target genes such as ICAM-1 and survivin were significantly higher in parenchymal and non-parenchymal liver cells isolated from ND-fed mice compared with cells from control mice (Supplementary Figure 1).
Effect of the ND on hepatocellular damage and inflammation. Mice were fed with standard chow (CTR) or a ND for 12 weeks. Representative images of paraffin-embedded liver sections from control and ND-fed mice immunohistochemically stained for (a) active caspase-3, (b) Ki-67 or (d) CD3. Measurement of mRNA expression levels of (c) cyclin D1, (e) Tnf, Il-1b, Cxcl1 (KC) and Ccl2 (Mcp1), and (f) the NADPH-oxidase complex genes Ncf1, Cyba and Cybb in livers from control and ND-fed mice by quantitative RT-PCR. (g) Western blot analysis and respective densitometric quantification of phosphorylated (Ser536) p65 in livers from control and ND-fed mice. (*P<0.05 compared with CTR).
Effect of the ND on Hepatic Fibrosis
Sustained hepatic injury and inflammation lead to the activation of hepatic stellate cells (HSCs), a key event of hepatic fibrosis. Also ND-fed mice revealed significantly heightened mRNA and protein levels of hepatic α-SMA, an established marker of activated HSC (Figures 4a–c).40 This was accompanied by a distinct profibrogenic transcriptional profile, characterized by significantly elevated hepatic levels of transforming growth factor beta (Tgf-β), tissue inhibitor of metalloproteases-1 (Timp-1) and plasminogen activator inhibitor-1 (Pai-1; Figure 4d), profibrogenic genes highly expressed in activated HSC.41 Accordingly, collagen type I (Coll-I) gene transcription was elevated (Figure 4d), resulting in increased hydroxyproline levels (Figure 4e) and extracellular matrix deposition (Figure 4f) in livers of ND-fed mice.
Effect of the ND on hepatic fibrosis. Mice were fed with standard chow (CTR) or a ND for 12 weeks. (a) Analysis of hepatic α-SMA mRNA expression by quantitative RT-PCR in control and ND-fed mice. (b) Western blot analysis and densitometric quantification of α-SMA expression in livers from control and ND-fed mice. (c) α-SMA immunostaining of liver sections from control and ND-fed mice. (d) Measurement of hepatic Tgf-β, Timp-1, Pai-1 and collagen type-I mRNA expression in control and ND-fed mice by quantitative RT-PCR. (e) Quantification of hepatic hydroxyproline-content and (f) representative images of sirius red/fast green-stained liver sections from control and ND-fed mice. (*P<0.05 compared to CTR).
Systemic Characterization of the Altered Hepatic Gene Expression Caused by Feeding the ND
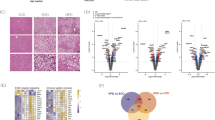
To further characterize this novel NASH model, we performed a transcriptome-wide gene expression analysis. Hepatic expression levels of ND-fed mice were compared with expression levels of mice fed with regular diet. Analysis of a total of 19 776 genes revealed significant (q<0.05) upregulation of 2305 genes (thereof 1030 highly significant; q<0.01) and significant downregulation of 713 genes (thereof 252 highly significant) in livers of ND-fed mice compared with control mice (Figure 5a). Classification of the highly significant differentially expressed genes in GO categories22 of biological processes showed significant over-representation of genes involved in metabolic and transport processes (Figure 5b). Also numerous genes related to signal transduction and transcriptional regulation as well as oxidation reduction processes were differentially expressed in ND-fed mice. Notably, among the 25 largest GO categories belonging to the biological processes branch, the categories ‘cell adhesion’, ‘lipid metabolic process’, ‘innate immune response’ and ‘inflammatory response’ carried significantly more differentially expressed genes than what would be expected by chance. Although alterations of ‘lipid metabolic processes’ reflect the dysregulated hepatic lipid metabolism, changes in the categories ‘innate immune response’ and ‘inflammatory response’ point to constant inflammatory processes in diet-induced NASH livers.
Effect of the NASH-inducing hepatic gene expression. (a) Heatmap of a transcriptome-wide gene expression array analysis of a total of 19 776 different genes revealing significant (q<0.05) upregulation of 2305 genes (thereof 1030 highly significant; q<0.01) and significant downregulation of 713 genes (thereof 252 highly significant) in livers of ND-fed mice (ND; n=4) compared with control mice (CTR; n=4). Here, the top 500 genes with smallest q-values are shown. (b) Classification of the highly significant differentially expressed genes in GO categories (the Gene Ontology Consortium, 2000) of biological processes. Among the 25 largest GO categories belonging to the biological processes branch, the categories ‘cell adhesion’, ‘lipid metabolic process’, ‘innate immune response’ and ‘inflammatory response’ carry significantly more differentially expressed genes than what would be expected by chance (white bars).
Effect of ND on c-Jun Expression
To identify possibly involved transcription factors responsible for the observed expressional alterations after ND feeding, we searched for over-represented transcription factor target sites among the differentially expressed genes. Mouse genes were first mapped to human genes using the homology mapping from NCBI (http://www.ncbi.nlm.nih.gov/projects/homology/maps). Then, the transcription factor-binding sites collection from the MSigDB database (collection C3, transcription factor targets)23 was used for hypergeometric testing within HTSanalyzeR. For both GO term and transcription factor-binding site enrichment, terms and binding sites with an FDR <0.05 were considered significant. Activator protein-1 (AP-1) binding sites were found to be significantly over-represented within the regulated genes (Table 2) indicating AP-1 as the most likely transcription factor to cause the transcriptional changes.
Furthermore, we looked for genes dysregulated in ND-fed mice, which interact on the protein level. Combining differentially expressed genes with a protein–protein interaction network, we searched for the largest connected dysregulated sub-network. Focusing on genes with an absolute log2-fold change of at least 0.5, we found a network of 23 connected genes, of which 22 were upregulated in ND-fed mice and one was repressed (Figure 6a). Interestingly, c-Jun (a component of the AP-1 complex) is a hub in this network, connecting other (transcription) factors, some of which have already been shown to be implicated in NASH development and progression.42, 43
Effect of ND on c-Jun and JNK expression. Mice were fed with standard chow (CTR) or a ND for 12 weeks. (a) Protein–protein interaction-based network of genes differentially expressed in ND-fed mice compared with control mice (green: upregulation, red: downregulation), focusing on genes with an absolute log2-fold change of at least 0.5 identified by transcriptome-wide gene expression analysis. Western blot analysis and respective densitometric quantification of (b) phosphorylated (Thr183/Tyr185) JNK, (c) phosphorylated (Ser73) c-Jun and (d) total c-Jun in control and ND-fed mice (*P<0.05 compared with CTR). (e) Representative images of paraffin-embedded liver sections from control (CTR) and ND-fed (ND) mice immunohistochemically stained for c-Jun (upper panel) and phosphorylated (Ser73) c-Jun (lower panel).
ND-fed mice showed significantly higher hepatic JNK activity (Figure 6b) and c-Jun-phosphorylation (Ser73) (Figure 6c) than control mice, indicative of c-Jun activation and subsequent transcriptional (AP-1) activity. Surprisingly, mRNA array analysis showed increased c-Jun expression (2.6-fold; FDR=0.02), which was confirmed by qPCR analysis (Supplementary Figure 2). Western blot (Figure 6d) and immunohistochemical analysis (Figure 6e) confirmed enhanced c-Jun expression on the protein level and identified mainly hepatocytes as cellular source of increased c-Jun levels in ND-fed livers. Here, the localization of c-Jun was mainly in nuclei (Figure 6e; upper panel) and corresponded with the immunohistochemical localization of phosphorylated (Ser73) c-Jun (Figure 6e; lower panel). Increased JNK activity and subsequent c-Jun phosphorylation have also been described in human NASH;32 however, higher c-Jun abundance in NAFLD has not been described so far in NAFLD models or human NAFLD tissues.
Expression of c-Jun in Human NAFLD
Notably, we observed also in human NAFLD tissue significantly enhanced c-Jun mRNA expression compared with normal liver tissue (Supplementary Figure 2). Furthermore, we applied TMA technology and immunohistochemistry (IH) to assess c-Jun expression in human liver tissue with different degree of steatosis and inflammation. IH was informative in 110 cases. The intensity of the immunohistochemical c-Jun signal revealed significant differences in individual patients, and thus, was systematically categorized into absent (score=0), weak (score=1) or strong (score=2) staining. Representative examples of steatotic liver tissues with different c-Jun staining intensities are depicted in Figure 7a. c-Jun staining intensity differed significantly (P<0.0001) between normal (n=38) and NAFLD tissues (n=72; Figure 7b). In simple steatotic livers, c-Jun IH showed a homogenous cytoplasmatic staining pattern of hepatocytes (Figure 7a). In NASH, c-Jun was localized in nuclei of hepatocytes and infiltrating inflammatory cells (Figure 7d) similarly as observed in our murine NASH model (Figure 6e). This nuclear staining pattern matched the one of phosphorylated (Ser73) c-Jun (Figure 7e) indicative of c-Jun activation. Activation of c-Jun induces c-Jun expression via a positive auto-regulatory loop, and inflammation is a known inducer of c-Jun expression.44 Accordingly, we observed significant correlation between the inflammatory score and the intensity of cytoplasmatic c-Jun expression (Table 3), while no association was found with age, gender, BMI class or ballooning of hepatocytes. However and surprisingly, we observed a remarkable correlation between the grade of hepatic steatosis and cytoplasmatic c-Jun staining (Figure 7c and Table 3). This finding indicates that hepatic steatosis leads to enhanced c-Jun levels independent of the presence of hepatic inflammation.
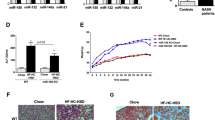
c-Jun expression pattern in liver tissues of patients with NAFLD. (a) Immunohistochemical c-Jun staining of a TMA comprising 110 non-tumorous human hepatic tissue specimens with normal histology (n=38) or NAFLD (n=72) different degree of steatosis and inflammation. Representative images of livers with NAFLD and with absent (score=0; left panel), weak (score=1; middle panel) or strong (score=2; right panel) immunoreactivity for c-Jun. (b) Percental distribution of livers with absent, weak or strong immunoreactivity for c-Jun in normal and NAFLD livers. (c) Percentage of steatotic hepatocytes in livers with absent, weak or strong cytoplasmatic staining for c-Jun (*P<0.05). Comparison between the staining patterns for c-Jun and phosphorylated c-Jun in tissue specimens of NAFLD livers applying TMA technology. Representative images of a NAFLD liver stained for (d) c-Jun and (e) phosphorylated (Ser73) c-Jun. Analysis of (f) c-JUN and (g) IL-8 mRNA expression levels by quantitative RT-PCR in PHHs stimulated with indicated concentrations of the FFA palmitic acid for 24 hours (*P<0.05 compared with FFA-free control; CTR) as well. (h, i) c-JUN and IL-8 mRNA expression in Hep3B hepatoma cells transfected with c-JUN-siRNA or control-siRNA and subsequent stimulation with 0.3 mM of the FFA palmitic acid for 24 h. (*P<0.05).
In line with this, we observed increased c-Jun expression in an in vitro model of hepatic steatosis, which we have recently described.11 Supplementation of increasing doses of FFAs to the cell culture medium led to a dose-dependent FFA accumulation and triglyceride accumulation (data not shown) and increased expression of c-Jun in both PHHs as well as a human hepatoma cell line (Figure 7f and Supplementary Figure 3A). It is noteworthy that this increase was flanked by increased expression of the proinflammatory cytokine interleukin 8 (IL-8; Figure 7g and Supplementary Figure 3B). Furthermore, we performed the according in vitro experiments with primary murine hepatocytes and found a similar induction of c-Jun expression in response to lipid accumulation as observed in PHHs indicating that this mechanism is not species dependent (data not shown). Transient transfection with siRNA against c-Jun significantly inhibited the FFA-induced c-Jun expression compared with cells transfected with control siRNA (Figure 7h). Moreover, inhibition of c-Jun impaired FFA-induced IL-8 expression (Figure 7i). Together, these findings suggest that steatosis induced c-Jun expression promotes inflammation in NAFLD.
DISCUSSION
We developed a new murine NASH model by feeding a diet that reproduces the nutritional situation of most NAFLD patients in Western countries, that is, chronic high caloric intake rich in saturated fats, cholesterol and fructose. After 12 weeks of feeding, this diet mice developed obesity and diabetes, key components of the metabolic syndrome and known factors driving human NAFLD development and progression. Furthermore, histological analysis revealed steatohepatitis that closely resembles human NASH. Importantly, ND-feeding also induced fibrosis, the most critical pathogenic factor for morbidity and mortality of NAFLD patients. Expression analysis revealed marked alterations in genes related to metabolic processes, transportation, signal transduction and oxidation reduction in the murine NASH model. AP-1 appeared as crucial transcriptional regulator of these alterations. Saturated FFAs activate this proinflammatory transcription factor in hepatocytes,45 and enhanced AP-1 activation has been demonstrated in obese patients with NASH.46 AP-1 is a homo- or heterodimer consisting of proteins belonging to the c-Jun, c-Fos, ATF and JDP families,47 with c-Jun being the best-characterized AP-1 component. The nuclear import of c-Jun is mediated by multiple mechanisms48, 49 and nuclear c-Jun levels correlate with AP-1 target gene activity.49 This further increases c-Jun protein abundance, as the jun proto-oncogene itself is activated by AP-1 in the manner of a positive auto-regulatory loop.44 The activity of c-Jun/AP-1 is markedly enhanced by phosphorylation of the transcriptional activation domain by JNKs.50 Sustained activation of the JNK pathway mediates the development and progression of experimental diet-induced NAFLD.51 Still, the role of different JNK-isoforms in NAFLD is not completely understood and it is becoming more and more evident that they may exhibit diverse effects in various cell types.8, 52
Apart from boosting c-Jun/AP-1 activity, JNK activation can also phosphorylate insulin receptor substrates (IRS)-1 and -2, which may lead to insulin resistance by blocking insulin receptor signal transduction.5 This is one mechanism by which fructose, regarded as a mostly insulin-independent sugar, can induce insulin resistance because of its JNK-activating properties.8 Further inducers of JNK activity are ROS, which are also known to have a role in NAFLD progression.36 In our model as well as in human NASH, ROS may arise from excessive β-oxidation of FFA, from mitochondrial damage caused by free cholesterol accumulation and crystallization,5 and from extensive activation of the NAD(P)H oxidase enzyme complex. The latter is activated by the adipokine leptin,53 and the levels of this adipokine are increased in our NASH model. Apart from the fact that JNK enhances c-Jun/AP-1 activity, Schreck et al49 demonstrated that the c-Jun itself promotes nuclear accumulation of JNK, further propelling AP-1 transcriptional activity. These various positive cross- and auto-regulatory loops may be co-responsible for the progression from simple steatosis to NASH. Factors that lead to a manifest derangement of the JNK/c-Jun/AP-1 regulatory system, or mechanisms by which the vicious circle of positive JNK/c-Jun/AP-1 auto-regulation could be interrupted, represent promising targets to interfere with NASH development and progression. Interestingly, in neurons JNK-mediated phosphorylation was shown to accelerate c-Jun degradation by allowing its recognition by the E3 ligase Fbw7-containing Skp/Cullin/F-box protein complex (SCFFbw7).54 Furthermore, activation of the JNK pathway promoted degradation of c-Jun (and JunB) through phosphorylation-dependent activation of the E3 ligase Itch in stimulated T cells.55 Analysis of these or similar mechanisms in hepatocytes are still elusive. It is important to distinguish interruption of hyperactivation of hepatocellular JNK/c-Jun/AP-1 signaling from a complete abolishment of JNK in hepatocytes, as mice with specific ablation of Jnk1 in hepatocytes surprisingly exhibit glucose intolerance, insulin resistance and hepatic steatosis.9 Moreover, we have shown that c-Jun abundance is a critical pathogenic factor in malignant melanoma independent of JNK activation.56 There, c-Jun levels are regulated at the post-transcriptional level by micro RNA (miR) 125b.57 Interestingly, miR-125b has also been described as a tumor suppressor in hepatocellular carcinoma,58 but its expression and function in NAFLD are unknown. Further studies are also required to unravel the mechanisms causing increased c-Jun levels in NAFLD.
In conclusion, our novel dietary murine NASH model induces important pathomechanisms also found in human NASH and indicates c-Jun/AP-1 activation as critical regulator of hepatic alterations. We newly discovered increased hepatic c-Jun expression in this model and confirmed this observation in NAFLD patients where we detected c-Jun abundance even in simple steatosis. It appears likely that higher c-Jun levels facilitate NASH development and progression. Thus, c-Jun may serve as prognostic marker and appears as attractive therapeutic target to prevent the development and progression of NASH.
Accession codes
References
Cohen JC, Horton JD, Hobbs HH . Human fatty liver disease: old questions and new insights. Science 2011;332:1519–1523.
de Wit NJ, Afman LA, Mensink M et al. Phenotyping the effect of diet on non-alcoholic fatty liver disease. J Hepatol 2012;57:1370–1373.
Collins S, Martin TL, Surwit RS et al. Genetic vulnerability to diet-induced obesity in the C57BL/6J mouse: physiological and molecular characteristics. Physiol Behav 2004;81:243–248.
de Wit NJ, Afman LA, Mensink M et al. Phenotyping the effect of diet on non-alcoholic fatty liver disease. J Hepatol 2012;57:1370–1373.
Farrell GC, van RD, Gan L et al. NASH is an inflammatory disorder: pathogenic, prognostic and therapeutic implications. Gut Liver 2012;6:149–171.
Tappy L, Le KA . Does fructose consumption contribute to non-alcoholic fatty liver disease? Clin Res Hepatol Gastroenterol 2012;36:554–560.
Lim JS, Mietus-Snyder M, Valente A et al. The role of fructose in the pathogenesis of NAFLD and the metabolic syndrome. Nat Rev Gastroenterol Hepatol 2010;7:251–264.
Seki E, Brenner DA, Karin M . A liver full of JNK: signaling in regulation of cell function and disease pathogenesis, and clinical approaches. Gastroenterology 2012;143:307–320.
Czaja MJ . JNK regulation of hepatic manifestations of the metabolic syndrome. Trends Endocrinol Metab 2010;21:707–713.
Gabele E, Dostert K, Dorn C et al. A new model of interactive effects of alcohol and high-fat diet on hepatic fibrosis. Alcohol Clin Exp Res 2011;35:1361–1367.
Wobser H, Dorn C, Weiss TS et al. Lipid accumulation in hepatocytes induces fibrogenic activation of hepatic stellate cells. Cell Res 2009;19:996–1005.
Dorn C, Kraus B, Motyl M et al. Xanthohumol, a chalcon derived from hops, inhibits hepatic inflammation and fibrosis. Mol Nutr Food Res 2010;54 (Suppl 2):S205–S213.
Patsenker E, Schneider V, Ledermann M et al. Potent antifibrotic activity of mTOR inhibitors sirolimus and everolimus but not of cyclosporine A and tacrolimus in experimental liver fibrosis. J Hepatol 2011;55:388–398.
Hellerbrand C, Muhlbauer M, Wallner S et al. Promoter-hypermethylation is causing functional relevant downregulation of methylthioadenosine phosphorylase (MTAP) expression in hepatocellular carcinoma. Carcinogenesis 2006;27:64–72.
Huber W, von Heydebreck A, S++ltmann H et al. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 2002;18 (Suppl 1):S96–104.
Dai M, Pinglang W, Boyd AD et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 2005;33:e175.
Smyth GK . Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 2004;3:1–25.
Ritchie M, Diyagama D, Neilson J et al. Empirical array quality weights in the analysis of microarray data. BMC Bioinformatics 2006;7:261.
Team RDC. R: A Language and Environment for Statistical Computing 2011.
Gentleman RC, Carey VJ, Bates DM et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 2004;5:R80.
Wang X, Terfve C, Rose JC et al. HTSanalyzeR: gene set over-representation, enrichment and network analyses for high-throughput screens. Bioinformatics 2011;27:879–880.
Consortium TGO. Gene ontology: tool for the unification of biology. Nat Genet 2000;25:25–29.
Subramanian A, Tamayo P, Mootha VK et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005;102:15545–15550.
Beisser D, Klau GW, Dandekar T et al. BioNet: an R-Package for the functional analysis of biological networks. Bioinformatics 2010;26:1129–1130.
Stark C, Breitkreutz BJ, Reguly T et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res 2006;34:D535–D539.
Dorn C, Riener MO, Kirovski G et al. Expression of fatty acid synthase in nonalcoholic fatty liver disease. Int J Clin Exp Pathol 2010;3:505–514.
Kleiner DE, Brunt EM, Van NM et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 2005;41:1313–1321.
Amann T, Bataille F, Spruss T et al. Reduced expression of fibroblast growth factor receptor 2IIIb in hepatocellular carcinoma induces a more aggressive growth. Am J Pathol 2010;176:1433–1442.
Bosserhoff AK, Moser M, Scholmerich J et al. Specific expression and regulation of the new melanoma inhibitory activity-related gene MIA2 in hepatocytes. J Biol Chem 2003;278:15225–15231.
Cousin SP, Hugl SR, Wrede CE et al. Free fatty acid-induced inhibition of glucose and insulin-like growth factor I-induced deoxyribonucleic acid synthesis in the pancreatic beta-cell line INS-1. Endocrinology 2001;142:229–240.
Ouchi N, Parker JL, Lugus JJ et al. Adipokines in inflammation and metabolic disease. Nat Rev Immunol 2011;11:85–97.
Ferreira DM, Castro RE, Machado MV et al. Apoptosis and insulin resistance in liver and peripheral tissues of morbidly obese patients is associated with different stages of non-alcoholic fatty liver disease. Diabetologia 2011;54:1788–1798.
Greco D, Kotronen A, Westerbacka J et al. Gene expression in human NAFLD. Am J Physiol Gastrointest Liver Physiol 2008;294:G1281–G1287.
McCullough AJ . Pathophysiology of nonalcoholic steatohepatitis. J Clin Gastroenterol 2006;40 (Suppl 1) S17-29.
Nakamuta M, Kohjima M, Morizono S et al. Evaluation of fatty acid metabolism-related gene expression in nonalcoholic fatty liver disease. Int J Mol Med 2005;16:631–635.
Rolo AP, Teodoro JS, Palmeira CM . Role of oxidative stress in the pathogenesis of nonalcoholic steatohepatitis. Free Radic Biol Med 2012;52:59–69.
Wouters K, van Gorp PJ, Bieghs V et al. Dietary cholesterol, rather than liver steatosis, leads to hepatic inflammation in hyperlipidemic mouse models of nonalcoholic steatohepatitis. Hepatology 2008;48:474–486.
Alkhouri N, Carter-Kent C, Feldstein AE . Apoptosis in nonalcoholic fatty liver disease: diagnostic and therapeutic implications. Expert Rev Gastroenterol Hepatol 2011;5:201–212.
Ramalho RM, Cortez-Pinto H, Castro RE et al. Apoptosis and Bcl-2 expression in the livers of patients with steatohepatitis. Eur J Gastroenterol Hepatol 2006;18:21–29.
Friedman SL . Mechanisms of hepatic fibrogenesis. Gastroenterology 2008;134:1655–1669.
Brenner DA . Molecular pathogenesis of liver fibrosis. Trans Am Clin Climatol Assoc 2009;120:361–368.
Nakajima T, Nakashima T, Okada Y et al. Nuclear size measurement is a simple method for the assessment of hepatocellular aging in non-alcoholic fatty liver disease: comparison with telomere-specific quantitative FISH and p21 immunohistochemistry. Pathol Int 2010;60:175–183.
Hardwick RN, Fisher CD, Canet MJ et al. Diversity in antioxidant response enzymes in progressive stages of human nonalcoholic fatty liver disease. Drug Metab Dispos 2010;38:2293–2301.
Angel P, Hattori K, Smeal T et al. The jun proto-oncogene is positively autoregulated by its product, Jun/AP-1. Cell 1988;55:875–885.
Joshi-Barve S, Barve SS, Amancherla K et al. Palmitic acid induces production of proinflammatory cytokine interleukin-8 from hepatocytes. Hepatology 2007;46:823–830.
Videla LA, Tapia G, Rodrigo R et al. Liver NF-kappaB and AP-1 DNA binding in obese patients. Obesity (Silver Spring) 2009;17:973–979.
Hess J, Angel P, Schorpp-Kistner M . AP-1 subunits: quarrel and harmony among siblings. J Cell Sci 2004;117:5965–5973.
Waldmann I, Walde S, Kehlenbach RH . Nuclear import of c-Jun is mediated by multiple transport receptors. J Biol Chem 2007;282:27685–27692.
Schreck I, Al-Rawi M, Mingot JM et al. c-Jun localizes to the nucleus independent of its phosphorylation by and interaction with JNK and vice versa promotes nuclear accumulation of JNK. Biochem Biophys Res Commun 2011;407:735–740.
Karin M, Gallagher E . From JNK to pay dirt: jun kinases, their biochemistry, physiology and clinical importance. IUBMB Life 2005;57:283–295.
Singh R, Wang Y, Xiang Y et al. Differential effects of JNK1 and JNK2 inhibition on murine steatohepatitis and insulin resistance. Hepatology 2009;49:87–96.
Hellerbrand C . Molecular targets for antifibrotic therapy in liver disease: using magic bullets for crossfire rather than a one-sided shotgun attack. Gut 2013, doi:10.1136/gutjnl-2013-305908 (e-pub ahead of print).
Brenner DA, Seki E, Taura K et al. Non-alcoholic steatohepatitis-induced fibrosis: Toll-like receptors, reactive oxygen species and Jun N-terminal kinase. Hepatol Res 2011;41:683–686.
Nateri AS, Riera-Sans L, Da CC et al. The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK signaling. Science 2004;303:1374–1378.
Gao M, Labuda T, Xia Y et al. Jun turnover is controlled through JNK-dependent phosphorylation of the E3 ligase Itch. Science 2004;306:271–275.
Spangler B, Vardimon L, Bosserhoff AK et al. Post-transcriptional regulation controlled by E-cadherin is important for c-Jun activity in melanoma. Pigment Cell Melanoma Res 2011;24:148–164.
Kappelmann M, Kuphal S, Meister G et al. MicroRNA miR-125b controls melanoma progression by direct regulation of c-Jun protein expression. Oncogene 2012;32:2984–2991.
Liang L, Wong CM, Ying Q et al. MicroRNA-125b suppressesed human liver cancer cell proliferation and metastasis by directly targeting oncogene LIN28B2. Hepatology 2010;52:1731–1740.
Acknowledgements
We want to thank Rudolf Jung, Monika Artinger, Ruth Schewior and Marina Fink for excellent technical assistance. This work was supported by grants from the German Research Association (DFG) to AB and CH, and the Medical Faculty of the University of Regensburg (ReForM) to CH and AB; MM was supported by the National Center for Tumor Diseases (DKFZ) and the Helmholtz Association; JCE and RS were supported by the Bavarian Genome Network (BayGene). We acknowledge the Human Tissue and Cell Research (HTCR) Foundation for supporting our research by making human liver tissue available.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
In order to determine the mechanisms of non-alcoholic steatohepatitis (NASH), the authors developed a murine model. Livers of mice fed a NASH-inducing diet (ND) exhibit the pathological changes found in human NASH. Transcriptome-wide gene expression analysis identified AP-1 as a key factor causing transcriptional changes seen in NASH mice. Additionally, analysis of differentially expressed genes and proteins identified c-Jun as the hub in the largest deregulated network in ND-livers.
Rights and permissions
About this article
Cite this article
Dorn, C., Engelmann, J., Saugspier, M. et al. Increased expression of c-Jun in nonalcoholic fatty liver disease. Lab Invest 94, 394–408 (2014). https://doi.org/10.1038/labinvest.2014.3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2014.3
Keywords
This article is cited by
-
RNA adenosine deaminase (ADAR1) alleviates high-fat diet-induced nonalcoholic fatty liver disease by inhibiting NLRP3 inflammasome
Laboratory Investigation (2022)
-
N-n-Butyl haloperidol iodide ameliorates liver fibrosis and hepatic stellate cell activation in mice
Acta Pharmacologica Sinica (2022)
-
Deficiency of WTAP in hepatocytes induces lipoatrophy and non-alcoholic steatohepatitis (NASH)
Nature Communications (2022)
-
Hepatocyte expressed chemerin-156 does not protect from experimental non-alcoholic steatohepatitis
Molecular and Cellular Biochemistry (2022)
-
Human hepatic in vitro models reveal distinct anti-NASH potencies of PPAR agonists
Cell Biology and Toxicology (2021)