Abstract
Three new metabolites, chaetomugilin S, dechloro-chaetomugilin A and dechloro-chaetomugilin D, were isolated from a strain of Chaetomium globosum originally obtained from the marine fish Mugil cephalus, and their absolute stereostructures were elucidated based on the basis of spectroscopic analyses, including 1D and 2D NMR techniques and some chemical transformations. Particularly, chaetomugilins T and U are the first compouds without a chlorine atom in azaphilones isolated from this fungal strain, to date. In addition, these compounds moderately inhibited the growth of cultured P388, HL-60, L1210 and KB cell lines.
Similar content being viewed by others
Introduction
We have focused on potential new antitumor materials from marine-derived microorganisms that produce a number of compounds with unique structures. As a part of this study, we have examined metabolites from the fungus Chaetomium globosum OUPS-T106B-6 originally obtained from the marine fish Mugil cephalus, and isolated several azaphilones. We already reported their absolute stereostructures and cytotoxic activities.1, 2, 3, 4, 5, 6, 7 Our continuing search for cytotoxic metabolites from this fungal strain led to the isolation of three new azaphilones designated as chaetomugilins S–U (1–3). Azaphilones have various bioactivities such as antimicrobial activity, nitric oxide inhibition (cohaerins),8 gp120-CD4-binding inhibition (isochromophiliones, ochrephilone, screotiorin and rubrorotiorin),9 monoamine oxidase inhibition (luteusins, TL-1, TL-2 and chaetoviridins),10, 11 platelet-derived growth factor-binding inhibition (RP-1551s)12 and antimalarial activity (cochliodones).13 These metabolites including chaetomugilins A–R have a chlorine atom at C-5; however, dechloro-chaetomugilin A (2) and dechloro-chaetomugilin D (3) have no substituent at C-5. Chaetomugiline S (1) exhibited moderate cytotoxic activity against the murine P388 leukemia cell line, the human HL-60 leukemia cell line, the murine L1210 leukemia cell line, and the human KB epidermoid carcinoma cell line. We describe herein the absolute configuration of the stereogenic centers and biological activities of these compounds.
Results and discussion
Identification and structure determination
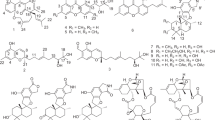
The microorganism from M. cephalus fish was cultured at 27 °C for 6 weeks in a medium (50 l) containing 1% soluble starch and 0.1% casein in 50% artificial sea water adjusted to pH 7.4, as reported previously.1, 2, 3, 4, 5, 6, 7 The EtOAc extract of the culture filtrate was purified, employing Sephadex LH-20, silica gel column chromatography and the reverse phased HPLC, to afford chaetomugilins S (1), dechloro-chaetomugilin A (2) and dechloro-chaetomugilin D (3) (Figure 1).
Chaetomugilin S (1) was assigned the molecular formula C23H27ClO6 based on the [M+H]+ peak in HRFABMS and the ratio of the intensity of isotope peaks (MH+/[MH+2]+). Its IR spectrum exhibited bands at 3423, 1716 and 1642 cm–1, characteristic of hydroxyl group, ester and α, β-unsaturated carbonyl groups. A close inspection of the 1H and 13C NMR spectra (Table 1) of 1 in DEPT and HMQC experiments revealed the presence of one primary methyl (C-13), 3 secondary methyls (11-CH3, 4′-methyl and C-6′), 1 tertiary methyl (7-CH3), 1 sp3-hybridized methylene (C-12), 5 sp3-methines (C-8, C-11, C-2′, C-4′ and C-5′) including an oxygen-bearing carbon (C-5′), 4 sp2-methines (C-1, C-4, C-9 and C-10) including an oxygen-bearing carbon (C-1), 2 quarternary oxygen-bearing sp3-carbons (C-7 and C-3′), 4 quarternary sp2-carbons (C-3, C-4a, C-5 and C-8a) including an oxygen-bearing carbon (C-3), and 2 carbonyls (C-6 and C-1′). The 1H–1H COSY analysis of 1 led to 3 partial structural units, as shown by bold-faced lines in Figure 2. The geometrical configuration of the double bond moiety (C-9—C-10) was deduced as the E configuration from the coupling constant of the olefinic protons (J9, 10=15.5 Hz). The connection of these units and the remaining functional groups was determined on the basis of the key HMBC correlations summarized in Figure 2. The connection of a chlorine atom to C-5 was reasonable from the molecular formula and the 13C NMR chemical shift of C-5 (δC 109.9). Thus, the planar structure of 1 was elucidated as shown in Figure 2.
The NOE correlations between H-8 and 7-CH3 in NOESY experiments, together with the coupling constant of the vicinal protons (J8, 2′=9.0 Hz) implied that H-8 was oriented cis to 7′-CH3 and trans to H-2′. The NOE correlations (H-2′/H-5′ and H-4′/7-CH3) together with the coupling constant (J4′, 5′=10.0 Hz) demonstrated that H-4′ and H-5′ were arranged in trans diaxial, and that 3′-OH was oriented cis to H-2′ (Figure 3). In CD spectrum of 1, it showed the same CD curve as that of 4 (Figure 4), and a negative Cotton effect (Δɛ353 –1.1) demonstrated clearly the absolute configuration at C-7 was S,4 allowing for the assignment of the absolute configuration of the asymmetric centers except C-11 (7S, 8S, 2′R, 3′R, 4′S and 5′S). To determine the absolute configurations at C-11 in 1, the alkaline degradation of 1 was carried out.4 The degradation of 1 with 5% potassium hydroxide afforded a carboxylic acid (yield 19.7%), which was identified with (2E,4S)-4-methylhex-2-enoic acid obtained from chaetomugilin D (4), whose absolute stereostructure had been already determined,4 by a similar manner in IR, UV, NMR spectra and the specific rotations (Scheme 1). Thus, the absolute configurations for C-11 of 1 were established as S.
Dechloro-Chaetomugilin A (2), which contained a chlorine atom less than chaetomugilin A (5), was assigned the molecular formula C23H28O7. Its general spectral features closely resembled those of 5 except for 1H NMR signal for H-4 (δH 6.10), and 13C NMR signals for C-4a (δC 144.3) and C-6 (δC 196.9) (Table 2). In addition, an olefinic methine (δH5.51,δC 107.3) newly appeared in 2. Analysis of 1H–1H COSY and HMBC correlations (from H-5 to C-4, C-7 and C-8a) elucidated the planar structure of 2, and implied that a chlorine atom at C-5 as other chaetomugilins was not present. In NOESY experiment of 2, the observed NOE correlations (H-8/H-5′, H-2′/H-4′ and H-8/7-CH3), together with the coupling constant between H-4′ and H-5′ (J4′, 5′=10.3 Hz), implied the relative configuration at C-7, C-8 and C-2′ –C-5′, i. e., H-8 is oriented cis to 7′-CH3 and H-5′, and trans to H-2′, 3′-OH and H-4′. In addition, a negative Cotton effect (Δɛ344 –2.1) in CD spectrum of 2 showed that the absolute configuration at C-7 was S,4 allowing for the assignment of the absolute configuration of the chiral centers except C-11 (7S, 8S, 2′R, 3′R, 4′R and 5′R) (Figure 5). For the absolute configurations at C-11 and C-12 in 2, a carboxylic acid produced by the alkaline degradation with 5% potassium hydroxide of 1 was identified with (2E,4R,5R)-5-hydroxy-4-methylhex- 2-enoic acid obtained from chaetomugilin A (5), in all spectral data and the optical rotation. Thus, the absolute configurations for C-11 and C-12 of 2 were established as both R.
Dechloro-Chaetomugilin D (3) that contained a chlorine atom less than chaetomugilin D (4), was assigned the molecular formula C23H28O6. Its general spectral features closely resembled those of 4 except for 1H NMR signal for H-4 (δH 6.09), and 13C NMR signals for C-4a (δC 145.0) and C-6 (δC 196.8) (Table 2), conversely, these chemical shifts were almost correspondent to those of chaetomugilin T (2). For the stereochemistry of 3, the NOEs characterizing the relative configuration at chiral centers and CD spectrum of 3 demonstrated that 3 was identical with dechlorine derivative of chaetomugilin D. The absolute configuration at C-11 in 3 was confirmed as S by the same procedure as the alkaline degradation of 1.
Cytotoxic activities
As a primary screen for antitumor activity, cancer cell growth inhibitory properties of chaetomugilin S (1), dechloro-chaetomugilin A (2) and dechloro-chaetomugilin D (3) were examined using the murine P388 leukemia cell line, the human HL-60 leukemia cell line, the murine L1210 leukemia cell line, and the human KB epidermoid carcinoma cell line. As shown in Tables 3, 1 exhibited moderate cytotoxic activity against the cancer cell lines. Henceforth, these compounds will be examined using a disease-oriented panel of 39 human cell lines14, 15 to reveal their selective cytotoxic activity and mode of action.
Experimental procedure
General procedures
Mps were determined on a Yanagimoto micro-melting point apparatus and are uncorrected. UV spectra were recorded on a Hitachi U-2000 spectrophotometer (Hitachi, Osaka, Japan) and IR spectra on a JASCO FT/IR-680 plus (JASCO, Tokyo, Japan). NMR spectra were recorded at 27 °C on Varian UNITY INOVA-500 and MERCURY spectrometers with tetramethylsilane (TMS) as an internal reference. FABMS was determined using a JOEL JMS-700 (Ver. 2) mass spectrometer (JOEL, Tokyo, Japan). Optical rotations were recorded on a JASCO J-820 polarimeter. Liquid chromatography over silica gel (mesh 230–400) was performed at medium pressure. HPLC was run on a Waters ALC-200 instrument equipped with a differential refractometer (R 401) and Shim-pack PREP-ODS (25 cm x 20 mm i.d.). Analytical TLC was performed on precoated Merck aluminum sheets (DC-Alufolien Kieselgel 60 F254, 0.2 mm) with the solvent system CH2Cl2-MeOH (19:1), and compounds were viewed under a UV lamp, and sprayed with 10% H2SO4, and then heated.
Culturing and isolation of metabolites
A strain of Chaetomium globosum was initially isolated from the marine fish Mugil cephalus, collected in Katsuura bay, Japan in October 2000. The marine fish was wiped with EtOH and its gastrointestinal tract applied to the surface of nutrient agar layered in a Petri dish. Serial transfers of one of the resulting colonies provided a pure strain of C. globosum. The fungal strain was cultured at 27 °C for 6 weeks in a liquid medium (70 l) containing 1% soluble starch and 0.1% casein in 50% artificial sea water adjusted to pH 7.4. The culture filtrate was extracted thrice with EtOAc. The combined extracts were evaporated in vacuo to afford a mixture of crude metabolites (22.1 g). The EtOAc extract was passed through Sephadex LH-20, using CHCl3–MeOH (1: 1) as the eluent. The second fraction (10.2 g) was chromatographed on a silica gel column with a CHCl3–MeOH gradient as the eluent. The 100% CHCl3 eluate (1.4 g) was purified by HPLC using MeCN–H2O (80: 20) as the eluent to afford 1 (4.3 mg). The MeOH–CHCl3 (1: 99) eluate (396.5 mg) was purified by HPLC using MeCN–H2O (55: 45) as the eluent to afford 3 (19.4 mg). The MeOH–CHCl3 (2: 98) eluate (1.7 g) was purified by HPLC using MeCN–H2O (35: 65) as the eluent to afford 2 (5.2 mg).
Chaetomugilin S (1): Yellow powder; mp 98–100 °C (CHCl3–MeOH); [α]D22 –22.5 (c 0.05, EtOH); UV λmax (EtOH) nm (log ɛ) 284 (3.70), 387 (3.80), 402 (3.78); IR νmax (KBr) cm–1 3423, 1716, 1642, 1560, 1522; HRFABMS m/z found 435.1572 [M+H]+ (clad for C23H2835ClO6, 435.1574). 1H and 13C NMR data are listed in Table 1.
Dechloro-Chaetomugilin A (2): Yellow powder; mp 110–112 °C (CHCl3–MeOH); [α]D22 –13.6 (c 0.07, EtOH); UV λmax (EtOH) nm (log ɛ) 282 (3.94), 368 (4.07), 384 (4.06); IR νmax (KBr) cm–1 3435, 1718, 1618, 1542, 1509; HRFABMS m/z found 417.1913 [M+H]+ (calculated for C23H29O7, 417.1913). 1H and 13C NMR data are listed in Table 2.
Dechloro-Chaetomugilin D (3): Yellow powder; mp 118–120 ° (CHCl3–MeOH); [α]D22 15.2 (c 0.22, EtOH); UV λmax (EtOH) nm (log ɛ) 281 (3.65), 367 (3.81), 381 (3.78); IR νmax (KBr) cm–1 3435, 1719, 1610, 1561, 1542; HRFABMS m/z found 401.1968 [M+H]+ (calculated for C23H29O6, 401.1966). 1H and 13C NMR data are listed in Table 3.
Degradation by potassium hydroxide of 1, 2 and 3
Chaetomugilin S (1) (8.6 mg) was dissolved in 3 ml of 5% aq. potassium hydroxide and stirred for 3 h at 100 °C. The reaction mixture was extracted with 10 ml of CHCl3. The water layer was adjusted to pH 3.0 with 9% sulfuric acid and re-extracted with 10 ml of AcOEt. The organic extract was concentrated to dryness in vacuo. The residue was purified by HPLC using a MeCN–H2O gradient (0: 100)—(60: 40) as the eluent to afford (2E,4S)-4-methylhex-2-enoic acid (0.5 mg, yield 19.7%)4 as a colorless oil.
Using the same procedure, dechloro-chaetomugilin A (2) (6.8 mg), whose absolute stereostructure has been established, was treated with 5% aq. potassium hydroxide (3 ml), and purified by HPLC to afford (2E, 4R, 5R)-5-hydroxy-4-methylhex-2-enoic acid (0.6 mg, yield 25.5%)4 as a colorless oil.
Using the same procedure, dechloro-chaetomugilin D (3) (3) (4.8 mg), whose absolute stereostructure has been established, was treated with 5% aq. potassium hydroxide (3 ml), and purified by HPLC to afford (2E,4S)-4-methylhex-2-enoic acid (0.3 mg, yield 19.5%)4 as a colorless oil.
Cytotoxic assay
Cytotoxic activities of chaetomugilin S (1), dechloro-chaetomugilin A (2) and dechloro-chaetomugilin D (3) were examined by the 3-(4,5-dimethyl-2-thiazolyl)- 2,5-diphenyl-2H-tetrazolium bromide (MTT) method. P388, HL-60, L1210 and KB cells were cultured in the Eagle’s Minimum Essential Medium (10% fetal carf serum) at 37 °C in 5% CO2. The test material was dissolved in dimethyl sulfoxide to give a concentration of 10 mM, and the solution was diluted with the Essential Medium to give concentrations of 200, 20 and 2 μM, respectively. Each solution was combined with each cell suspension (1 × 105 cells ml−1) in the medium, respectively. After incubation at 37 °C for 72 h in 5% CO2, the grown cells were labeled with 5 mg ml−1 MTT in phosphate-buffered saline, and the absorbance of formazan dissolved with 20% SDS in 0.1 N HCl was measured at 540 nm, using microplate reader (Model 450, Bio-rad, Osaka, Japan). Each value was expressed as a percentage, relative to a control cell suspension prepared without the test substance. All assays were performed three times, semilogarithmic plots were constructed from the averaged data, and the effective dose of the substance required to inhibit cell growth by 50% (IC50) was determined.

The alkaline degradation of 1.
References
Yamada, T. et alAbsolute stereostructures of cytotoxic metabolites, chaetomugilins A-C, produced by a Chaetomium species separated from a marine fish. Tetrahedron Lett. 49, 4192–4195 (2008).
Muroga, Y., Yamada, T., Numata, A. & Tanaka, R. Chaetomugilins, new selectively cytotoxic metabolites, produced by a marine fish-derived Chaetomium species. J. Antibiot. 61, 615–622 (2008).
Yamada, T., Muroga, Y. & Tanaka, R. New azaphilones, seco-chaetomugilins A and D, produced by a marine-derived Chaetomium globosum. Marine drugs 7, 249–257 (2009).
Muroga, Y., Yamada, T., Numata, A. & Tanaka, R. Chaetomugilins I–O new potent cytotoxic metabolites from a marine-fish-derived Chaetomium species. Stereochemistry and biological activities. Tetrahedron 65, 7580–7586 (2009).
Yamada, T., Muroga, Y., Shigeta, H., Numata, A. & Tanaka, R. Absolute stereostructures of chaetomugolins G and H produced by a marine-derived Chaetomium species. J. Antibiotic. 62, 353–357 (2009).
Muroga, Y., Yamada, T., Numata, A. & Tanaka, R. 11- and 4′-epimers of chaetomugilin A, novelcytostatic metabolites from marine fish-derived fungus Chatomium globosum. Helv. Chim. Acta 93, 542–549 (2010).
Yamada, T. et alNew class azaphilone produced by a marine fish-derived Chaetomium globosum. The stereochemistry and biological activities. Bioorg. Med. Chem. 19, 4106–4113 (2011).
Quang, D. N., Stadler, M., Fournier, J., Tomita, A. & Hashimoto, T. Cohaerins C-F four azaphilones from the xylariaceous fungus Annulohypoxylon cohaerens. Tetrahedron 62, 6349–6354 (2006).
Matsuzaki, K., Tahara, H., Inokoshi, J. & Tanaka, H. New brominated and halogen-less derivatives and structure-activity relationship of azaphilaones inhibiting gp120-CD4 binding. J. Antibiot. 51, 1004–1011 (1998).
Yoshida, E. et alisolation of three new azaphilones, luteusins C, D, and E, from an Ascomycete. Talaromyces luteus. Chem. Pharm. Bull. 44, 284–287 (1996).
Takahashi, M., Koyama, K. & Natori, S. Four new azaphilones from Chaetomium globosum var. flavo-viridae. Chem. Pharm. Bull. 38, 625–628 (1990).
Toki, S. et alRP-1551s, a family of azaphilones produced by Penicillium sp., inhibit the binding of PDGF to the extracellular domain of its receptor. J. Antibiot. 52, 235–244 (1999).
Phonkerd, N. et alBis-spiro-azaphilones and azaphilones from the fungi Chaetomium cochliodes VTh01 and C. cochliodes CTh05. Tetrahedron 64, 9636–9645 (2008).
Yamori, T. et alPotent antitumor activity of MS-247, a novel DNA minor groove binder, evaluated by an in vitro and in vivo human cancer cell line panel. Cancer Res. 59, 4042–4049 (1999).
Yamori, T. Panel of human cancer cell lines provides valuable database for drug discovery and bioinformatics. Cancer Chemother. Pharmacol 52 ((Suppl 1)), S74–S79 (2003).
Acknowledgements
We thank Dr T Ito (National Institute of Technology and Evaluation, Biological Resource Center) and Dr Endo (Kanazawa University) for identification of the fungal strain, and for supply of the cancer cells, respectively. We are grateful to Ms M Fujitake and Dr K Minoura of this university for MS and NMR measurements, respectively. This study was supported by a Grant-in-aid for High Technology from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yamada, T., Jinno, M., Kikuchi, T. et al. Three new azaphilones produced by a marine fish-derived chaetomium globosum. J Antibiot 65, 413–417 (2012). https://doi.org/10.1038/ja.2012.40
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2012.40
Keywords
This article is cited by
-
Genetic and Histopathological Alterations in Caco-2 and HuH-7 Cells Treated with Secondary Metabolites of Marine fungi
Journal of Gastrointestinal Cancer (2022)
-
Cytochalasans and azaphilones: suitable chemotaxonomic markers for the Chaetomium species
Applied Microbiology and Biotechnology (2021)
-
One Strain-Many Compounds Method for Production of Diverse Metabolic Profiles Using Chaetomium sp. from Astragalus membranaceus
Chemistry of Natural Compounds (2021)
-
New azaphilones from Penicillium variabile, a fungal endophyte from roots of Aconitum vilmorinianum
The Journal of Antibiotics (2020)