- NEWS
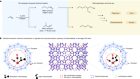
‘Why didn’t we think to do this earlier?’ Chemists thrilled by speedy atomic structures
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 563, 16-17 (2018)
doi: https://doi.org/10.1038/d41586-018-07213-3
References
Gruene, T. et al. Angew. Chem. Int. Ed. https://doi.org/10.1002/anie.201811318 (2018).
Jones, C. G. et al. Preprint at ChemRxiv https://doi.org/10.26434/chemrxiv.7215332.v1 (2018).

 The revolution will not be crystallized: a new method sweeps through structural biology
The revolution will not be crystallized: a new method sweeps through structural biology