Abstract
The recent identification of a novel protein–protein interaction module, termed PB1, in critical signaling molecules such as p62 (also known as sequestosome1), the atypical PKCs, and Par-6, has unveiled the existence of a new set of signaling complexes, which can be central to several biological processes from development to cancer. In this review, we will discuss the most recent advances on the role that the different components of these complexes have in vivo and that are relevant to human disease. In particular, we will review what we are learning from new data from knockout mice, and the indications from human mutations on the real role of these proteins in the physiology and biology of human diseases. The role that PKCζ, PKCλ/ι, and Par-4 have in lung and prostate cancer in vivo and in humans will be extensively covered in this article, as will the multifunctional role of p62 as a novel hub in cell signaling during cancer and inflammation, and the mechanistic details and controversial data published on its potential role in aggregate formation and signaling. All this published information is shedding new light on the proposed pathological implications of these PB1-regulators in disease and shows their important role in cell physiology.
Similar content being viewed by others
Main
The protein kinase C (PKC) isozymes constitute a family of Ser/Thr kinases of the AGC group, which are subdivided into classical, novel, and atypical isoforms, based on structure and sequence homology and on their cofactor requirements. They all contain a C-terminal kinase domain linked through a variable ‘V3’ domain to an N-terminal regulatory domain, in which most of the structural differences reside (Figure 1a). The latter contains three functional elements: (i) an inhibitory region (pseudosubstrate), (ii) a C1 domain or zinc finger (one copy or tandem repeats), and (iii) a C2 domain. Distinct features of the regulatory domains contribute substantially to the particular roles of individual isoforms and to their respective mechanisms of action.
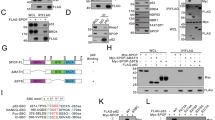
The atypical PKC network. (a) Domain organization of the different PKC subfamilies: aPKC, atypical PKC; cPKC, classical PKC; nPKC, novel PKC. PB1, PB1 dimerization domain; PS, pseudosubstrate region; ZF, zinc finger; C2, C2 domain. (b) The atypical PKC isoforms establish a network of protein interactions with adapter proteins (such as p62 and Par-6) binding through the PB1 domain and with regulators (such as Par-4) through their zinc finger domain. PB1–PB1 interactions confer specificity to the actions of the aPKCs. The interaction with p62 allocates the aPKCs in the NF-κB pathway, whereas through Par-6 the aPKCs regulate cell polarity. Domain architectures of the different proteins in the network are shown: Zn ZZ, ZZ-type zinc finger; TB, TRAF6 binding; LIR, LC3-interacting region; UBA, ubiquitin associated; LZ, leucine zipper, ZF, Zinc Finger; CRIB, Cdc42 Rac-interactive binding
The atypical protein kinase C (aPKC) subfamily is composed of two members, PKCζ and PKCλ/ι. PKCλ is the mouse homolog of the human PKCι. The two aPKC isoforms are highly related, sharing an overall amino acid identity of 72%.1 The conservation in their sequences is most striking in the catalytic domain, which is also conserved among other PKC isotypes that belong to the classical and novel subfamilies. In contrast, the regulatory domain of the aPKC subfamily diverts from other members of the PKC family; it has only one zinc finger, whereas the other PKCs have two1 (Figure 1a). Through the zinc-finger domain, the aPKCs bind Par-4, a negative regulator of their enzymatic activity. Similar to the novel PKCs, the aPKCs lack the characteristic C2 domain that is present in the classical isoforms (Figure 1a). These important structural differences may explain why the aPKCs are insensitive to Ca2+, diacylglycerol, and phorbol esters, which are potent activators of the other isoforms.1
The PB1 Domain: a Distinctive Characteristic of the aPKCs
The recent identification of the protein-interaction domain PB1, present at the N-terminus of the aPKCs, has opened new avenues to explore the roles of these kinase isoforms by looking for adapters and regulators that could shed light on their functions2 (Figure 1). It is well known that PKCs are kinases that display little selectivity in vitro and when overexpressed in cells. This invokes the need for cellular mechanisms to confer functional specificity while preserving the capacity for cross talk, which is necessary for the regulation of complex biological processes. As the aPKCs have been implicated in diverse cellular functions, different adapter proteins must exist to achieve the required specificity during cell signaling.3 In this regard, the PB1s are dimerization/oligomerization domains present in adapter and scaffold proteins, as well as in kinases, and serve to organize platforms to assemble the protein complexes necessary for such specificity. The PB1 domains are named after the prototypical domains found in Phox and Bem1p, which mediate polar–heterodimeric interactions. The PB1 domains comprise about 80 amino acid residues and are grouped into three types: type I (or type A), type II (or type B), and type I/II (or type AB). The type I domain group contains a conserved acidic DX(D/E)GD segment (called the OPCA motif) that interacts with a conserved lysine residue from a type II domain. Type I includes the PB1 domains of p40phox, MEK5, and NBR1, whereas type II occurs in p67phox, Par-6, MEKK2, and MEKK3. The type I/II PB1 domain, containing both the OPCA motif and the invariant lysine, is present in the aPKCs and in p62 (also known as sequestosome-1).2, 4 Type I and type II PB1 domains interact with each other in a front-to-back manner resulting in heterodimers in which acidic residues on the OPCA motif form salt bridges with basic residues of the type II PB1 domain.
Two-hybrid screenings in yeast identified p62 and Par-6 as selective adapters for the aPKCs.3, 5, 6, 7, 8 Par-6 has been shown to be central to the control of cell polarity and, through its PB1 domain, allocates the aPKCs specifically in polarity-related functions2 (Figure 1b). On the other hand, the p62/aPKC-signaling platform has a critical role in NF-κB activation9 (Figure 1b). p62 interacts with PKCζ and PKCλ/ι, but not with any of the other closely related PKC family members. It is not a substrate, and does not seem to significantly affect the intrinsic kinase activity of PKCζ or PKCλ/ι. Moreover, it harbors a number of domains that support its role as a scaffold in cell signaling (Figure 1b). Thus, the formation of aPKC complexes with different adapters, scaffold proteins, and regulators, such as Par-6, p62, and Par-4, serves to confer specificity and plasticity to the actions of these kinases and to establish signaling networks that control several key cellular functions. However, the factors that determine which complex is formed at a given time and within a specific cell context remain to be identified (Figure 1). In this regard, new phosphorylation events have been identified that take place in the PB1 domain of PKCι, which may offer an explanation of a novel mechanism that could account for switches between the different interacting partners.10
Specificity and Function of the aPKCs
Before the recent availability of loss-of-function animal models, the similarity between PKCλ/ι and PKCζ and the lack of rigorously specific genetic and biochemical tools have hampered the effort to assign unique functions to the individual isoforms. For example, many studies have used commercially available antibodies that do not discern between the two aPKC isoforms. Also, attempts to inhibit aPKC enzymatic activity in cell cultures have made extensive use of a peptide with the sequence of the pseudosubstrate, which is identical for both aPKCs and therefore non-specific. In addition, the pseudosubstrate region is highly conserved among all the PKCs, not only the aPKCs, which casts serious doubts on studies that rely solely on the use of such reagents to establish the role of the aPKCs in a given function.11, 12 Moreover, the overexpression of dominant-negative and active forms of the two aPKCs does not necessarily discriminate between specific functions for each isoform, as these manipulations may impinge on pathways other than those with physiological relevance to each aPKC isotype. Despite these problems, however, the genetic inactivation of these isoforms in vivo by using knock out (KO) mouse technology is starting to shed light on their specific roles.
The fact that PKCλ/ι KO in mice is embryonic lethal at early stages, probably because of defects in cell polarity,13 whereas the PKCζ KO mice are born in Mendelian ratios,14 was a first indication of the different and specific functions that each of these kinases might have in vivo. The phenotypic analysis of PKCζ KO mice revealed a role for this kinase in the control of the immune response.2 That is, PKCζ-deficient mice displayed alterations in the development of secondary lymphoid organs, showing morphological defects in the spleen's marginal zone and Peyer's Patches, and a reduced percentage of mature B cells.14, 15 In keeping with this, the loss of PKCζ impaired B-cell survival and proliferation in response to activation through the B-cell receptor with no major alterations in T-cell proliferation.15 In addition, the analysis of PKCζ-deficient mice unveiled an important role of PKCζ in the control of T-cell polarization programs, specifically during Th2 differentiation. In fact, PKCζ levels were increased during Th2 but not Th1 differentiation of CD4+ T cells, and the loss of PKCζ impaired the secretion of Th2 cytokines in vitro and in vivo, Jak1 activation, and the nuclear translocation and tyrosine phosphorylation of Stat6, essential downstream targets of IL-4 signaling. Moreover, PKCζ KO mice displayed dramatic inhibition of ovalbumin (OVA)-induced allergic airway disease, strongly suggesting that PKCζ might be a good candidate for a novel therapeutic target in asthma, through the control of the IL-4 pathway.16
The aPKCs have also been implicated as important mediators in the control of cell survival through the activation of NF-κB in multiple cell systems.3, 17, 18 Indeed, the genetic inactivation of PKCζ in mice supports a key role of this isoform in the activation of NF-κB in that PKCζ deficiency impairs NF-κB at two levels.14 In the lung, where PKCζ is especially abundant, this kinase is required for the activation of IKK in vivo, whereas in other systems, such as embryo fibroblasts, endothelial cells, and B cells,15, 19 PKCζ controls the phosphorylation of the RelA subunit of the NF-κB complex at Ser311, enabling its interaction with the transcriptional co-activator CBP and subsequent gene expression.20 Therefore, depending on the system, PKCζ could be considered an IKKβ kinase or may act downstream of IKKβ by controlling the transcriptional activity of the NF-κB complex.
On the other hand, unlike PKCζ, PKCλ/ι is required during development, as evidenced by the fact that PKCλ/ι-deficient mice die by embryonic day 9.5, likely because of abnormalities in development detected as early as day 6.5.13 This phenotype is in agreement with that found for the disrupted expression of the aPKC orthologs in C. elegans,21 Xenopus,11 and Drosophila.22, 23, 24, 25 Functional knockouts in these organisms result in early embryonic lethality because of defects in polarity and asymmetric cell division.21, 22, 23, 24, 25 The analysis of tissue-specific conditional PKCλ/ι-deficient mice is helping to elucidate the in vivo role of this atypical isoform. For example, the selective deficiency of PKCλ/ι in the liver resulted in increased insulin sensitivity,26 whereas β-cell deficiency impaired glucose-induced insulin secretion and glucose tolerance.27 Muscle-specific PKCλ/ι-deficient mice also provided evidence of its role in insulin action with a phenotype closely mimicking the human metabolic syndrome.28 Whether PKCλ/ι is dysfunctional in acquired insulin resistance has yet to be determined in humans. In relation to its function in the immune response, PKCλ/ι, similar to PKCζ, has an essential role in Th2 establishment and allergic airway disease in mice.29 Thus, the specific deletion of PKCλ/ι in activated T cells showed that this kinase is required for T-cell polarity and the activation of Th2-transcription factors, such as NF-κB, NFATc1, and GATA-3, linking defects in polarity to a functional impact on the Th2-mediated responses.29
The aPKC Regulators and Adapters Provide Clues to aPKC Function
Studying the in vivo function of the adapters and interacting partners of the aPKCs is an important tool to comprehensively address the physiological role of this kinase network. In fact, the analysis of the phenotype of Par-4-deficient mice confirmed the functional implication of the aPKCs in the immune response, mostly through their ability to regulate NF-κB. The available data support a model according to which the interaction of Par-4 with the zinc-finger region of the aPKC regulatory domain leads to the inhibition of aPKC enzymatic activity and the consequent reduction of NF-κB activity.30 In this regard, the loss of Par-4 in embryo fibroblasts leads to increased aPKC and NF-κB activation.31 Consistent with this, the NF-κB-dependent antiapoptotic protein XIAP is expressed at significantly elevated levels in Par-4–null cells, which correlates with reduced caspase-3 activation and apoptosis.31 In addition, Par-4 and PKCζ KO mice display opposite immune system phenotypes in vivo.15, 32 That is, although PKCζ–/– mice have impaired B-cell proliferation and function,15 Par-4–/– mice have increased B-cell and T-cell proliferation.32 Also, Par-4–/– T cells overproduce the Th2 cytokine IL-4,33 whereas PKCζ–/– T cells show impaired Th2 polarization and IL-4 secretion ex vivo and in vivo.16 As Par-4 binds and inhibits both aPKC isoforms, understanding the role of this negative regulator might help to predict the impact of inhibiting both isozymes in vivo, as could be the case with aPKC pharmacological inhibitors.
With regard to the adapter protein p62, it has been shown to be required for NF-κB signaling in several systems,34, 35, 36 including Drosophila, in which a functionally relevant homolog termed Ref(2)P has been identified.37 It is noted that p62 has been shown to be required for the sustained phase of NF-κB activation during T-cell differentiation, a process that is critical in asthma and other allergic diseases.38 Interestingly, p62 levels are induced on T-cell differentiation,38 suggesting that p62 is necessary to control biochemical events required for proper differentiation. The loss of p62 in T cells impairs their ability to produce Th2 cytokines ex vivo and is required for an optimal lung inflammatory response.38 Therefore, p62, similar to PKCζ and PKCλ/ι, emerges as an important component of the signaling cascades regulating Th2 function and asthma.
On the other hand, Par-6 has been shown by genetic manipulations to be critically implicated in the control of cell polarity in C. elegans and Drosophila.7, 8 Although genetic data have yet to be produced to prove the role of the aPKCs and Par-6 in different aspects of mammalian cell polarity, overexpression analyses have implicated the Par-6/aPKC complex in the control of the epithelial–mesenchymal transition,39 T-cell40 and neuronal polarity,41 and cell polarity in migrating astrocytes,42 among other functions. Our recent demonstration of a role for PKCλ/ι in the control of T-cell polarity in knockout cells and mice is the first genetic demonstration of such a role for an aPKC in mammals. Therefore, the formation of aPKC complexes with different adapters and scaffold proteins serves to confer specificity and plasticity to the actions of these kinases.
The Cancer Biology of the aPKCs
Cell transfection and overexpression experiments suggested a role for the aPKC pathways in tumorigenesis and led to different proposed mechanisms of action. However, the actual pathophysiological relevance of the different components of this network is now being established by studies in human cancer samples, and, importantly, in mouse genetic KO models as well.
In the case of PKCζ, a number of studies support its clinical relevance in tumorigenesis, including reports on altered expression in different types of cancer. Most of the studies, however, are restricted to microarray data with very limited reliable information on protein expression, as the antibodies used for these analyses recognize both aPKCs, thus precluding any conclusion regarding isoform specificity. It is noted that different studies in human tumor specimens have reported both the up- and downregulation of PKCζ expression, which may indicate tissue-specific roles for this aPKC isoform. That is, downregulated expression of PKCζ has been shown in glioblastoma,43, 44, 45, 46 lung cancer,47, 48, 49 kidney renal clear cell carcinoma,50 melanoma,51 and pancreatic cancer,52 whereas its upregulation has been shown in prostate cancer,53, 54 bladder cancer,55, 56 and lymphomas57, 58 All these studies point to an important role for PKCζ in human carcinogenesis. They have also unveiled a need for studies on the mechanism of PKCζ action to rigorously test whether it is an oncogene or, on the contrary, a tumor suppressor, and whether this depends on tissue-specific factors. A recent study from the Vogelstein laboratory, which could be of high relevance to this question, describes a genome-wide catalog of genetic changes in breast and colorectal tumors in which exons representing more than twenty-thousand transcripts from more than eighteen-thousand genes were analyzed.59 In this study, they identified a few commonly mutated genes. Interestingly, their analysis led to the identification of a mutated form of PKCζ (S514F), along with mutations in other genes in human colorectal cancer that can be grouped bioinformatically in a pathway including, in addition to PKCζ, relevant signal transducers such as IRS2, IRS4, PIK3R5, PTEN, RHEB, RPS6KB1, and PIK3CA.59 These results reinforce the earlier suggestion of a role for PKCζ in human tumorigenesis. Surprisingly, biochemical analysis of the consequence of the S514F mutation revealed that it significantly impaired PKCζ enzymatic activity.49 This could be interpreted to mean that tumorigenesis is associated with impaired PKCζ and that, in fact, this kinase should be considered as a tumor suppressor. Consistent with this hypothesis, the overexpression of catalytically active PKCζ, but not the S514F mutant, in Ras-expressing NIH-3T3 fibroblasts restrained tumor growth in a xenograft in vivo model.49 Furthermore, overexpression of the K281R kinase-dead mutant of PKCζ in the same in vivo model further enhanced tumorigenesis.49
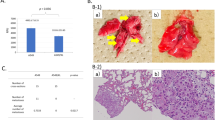
Collectively, the results of these studies strongly suggest that PKCζ activity is important for restraining tumorigenesis in vivo. In keeping with this notion, PKCζ levels were low or undetectable in a significant proportion of human lung carcinomas.49 A consideration of this finding, combined with the potential existence of inactivating mutations similar to the one identified in colon cancer patients, suggests that, in fact, the loss or inactivation of PKCζ will lead to tumorigenesis. A more physiological cancer model using PKCζ KO mice strongly supports these conclusions.49 That is, the analysis of a mouse model of pulmonary adenocarcinoma in which oncogenic Ras is inducibly expressed in type II alveolar epithelial cells in response to doxycycline60 showed that lung tumorigenesis is dramatically enhanced in a PKCζ KO background compared with that of WT mice.49
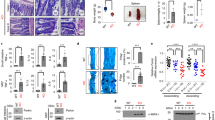
A negative role for PKCζ in carcinogenesis contrasts with the apparent role of PKCλ/ι. That is, xenograft experiments in Ras-transformed embryo fibroblasts showed that the loss of PKCλ/ι dramatically impaired tumorigenesis (Figure 2). This is in agreement with results from Fields and co-workers showing that the loss of PKCλ/ι severely impaired intestinal tumor formation in the ApcMin/+ mouse model.61 It is quite remarkable that KO mice lacking PKCλ/ι show phenotypes completely opposite to PKCζ KO mice because the aPKCs share a striking degree of homology. Consistent with this, an increasing number of studies in humans show aberrant expression of PKCλ/ι in several cancer types,62 but, contrary to PKCζ, PKCλ/ι is highly upregulated in all types of tumors, through gene amplification, increased mRNA expression, and protein overexpression. For example, PKCλ/ι protein levels are overexpressed in non-small cell lung adenocarcinomas (NSCLC), and the PKCλ/ι gene is frequently amplified in this tumor type.63 Furthermore, these studies propose that PKCλ/ι expression is a useful marker of poor prognosis in this type of malignancy.47, 63, 64 Moreover, PKCλ/ι is genomically amplified and overexpressed in ovarian cancers,65, 66, 67, 68 giving rise to the loss of apical–basal epithelial cell polarity.66 A similar mechanism has been proposed in a recent report on overexpression and altered localization of PKCλ/ι in breast cancer, suggesting that the normal apicobasal polarity is lost on the progression of a breast lesion to invasive ductal carcinoma.69 Gene locus amplification of PKCλ/ι has also been reported in esophageal squamous cell carcinoma.70 PKCλ/ι is also upregulated in bladder cancer,56 prostate cancer,71 sarcoma,72 lymphoma,58 and multiple myeloma.73 Taken together, these observations support a critical role for both aPKCs in cancer, but suggest that they are most likely playing opposite roles.
The Tumor Suppressor Role of Par-4
The fact that Par-4 binds both aPKCs, inhibiting their enzymatic activity, combined with the observation that its overexpression leads to increased apoptosis31, 74 is puzzling in light of the opposing roles of the two aPKCs in carcinogenesis. The generation of Par-4 KO mice allowed the establishment of its actual in vivo role in cancer. It is noted that 80% of the Par-4 KO females presented endometrial hyperplasia by 9 months of age, and at least 36% developed endometrial adenocarcinomas after 1 year of age.75 Par-4 KO males showed a high incidence of prostate hyperplasia and prostatic intraepithelial neoplasias.76 These data strongly suggest that Par-4 could be a novel tumor suppressor in these two types of tumorigenic process. The analysis of human tumors supports this notion. That is, a study using cDNA arrays, quantitative reverse transcription-PCR, and immunohistochemistry detected Par-4 downregulation in approximately 40% of human endometrial carcinomas.75 This study also showed that Par-4 promoter hypermethylation was detected in 32% of the tumors in association with low levels of Par-4 protein, and was more common in carcinomas positive for microsatellite instability,75 indicating that promoter hypermethylation is the mechanism whereby Par-4 levels are downregulated in tumor cells.
Interestingly, recent studies show that Par-4 expression is lost in a high percentage of human prostate carcinomas (about 60%), with a significant association between Par-4 promoter methylation and lack of Par-4 expression, and a clear association with PTEN loss (Diaz-Meco MT and Moscat J, unpublished observations). This is very interesting because PTEN loss has been associated with, and shown to be critical in, prostate cancer.77, 78 Interestingly, Par-4 KO mice, similar to PTEN-heterozygous mice, develop only benign prostate lesions, but, importantly, concomitant Par-4 ablation and PTEN heterozygosity lead to invasive prostate carcinoma in mice (Diaz-Meco MT and Moscat J, unpublished observations). These results establish a cooperation between the two tumor suppressors, Par-4 and PTEN, as relevant for the development of prostate cancer in mice and possible in humans as well.
Of potential interest to establishing the generality of Par-4 as a tumor suppressor in cancer are data showing that the loss of Par-4 clearly enhances lung carcinogenesis in a mouse lung cancer model in which oncogenic Ras is introduced following a knock-in strategy, and inducibly expressed in an endogenous manner.79 This and the doxycycline-inducible lung cancer model discussed above are physiologically relevant in vivo lung cancer models because the target cell, as in humans, is the type II pneumocyte that, in addition to the Clara cells, is the most likely precursor of human lung carcinoma.60, 80 Therefore, it appears that Par-4 is a tumor suppressor not only in the endometrium and prostate cancer but also in lung cancer. Consistent with this concept, a study of its expression in human NSCLC revealed that 47% of tumors were negative for Par-4 as determined by IHC, and that there was a clear correlation between the loss of Par-4 and tumor type. That is, 41% of the adenocarcinomas were negative for Par-4 expression, whereas only 6% of squamous cell carcinomas showed negative staining for Par-4. Also, when the adenocarcinomas were stratified by grade, it was clear that 74% of grade III tumors had lost Par-4 expression, whereas 59% of grade I-II tumors were negative for Par-4. Together, these data show that Par-4 is a relevant tumor suppressor gene in a significant number of human malignancies, strongly suggesting that this protein may have an important function in the prevention of, at least, endometrial, prostate, and lung cancer.
What Lies Downstream of Par-4?
As Par-4 manifests tumor suppressor activities76, 79 and is known to be involved in the binding and inhibition of PKCλ/ι and PKCζ,74 it follows that both aPKCs are downstream targets of Par-4 in carcinogenesis. However, the fact that the inhibition or deletion of PKCζ enhances tumorigenesis,49 similar to Par-4 deletion,75, 76, 79 whereas the inactivation of PKCλ/ι blocks tumorigenesis (Figure 2), could be interpreted to mean that PKCλ/ι, but not PKCζ, is the bona fide downstream target of Par-4 in cancer. Surprisingly, the simultaneous genetic in vivo inactivation of Par-4 and PKCζ in double KO mice leads to the ablation of Par-4 deficiency-induced prostate hyperplasia and PIN (Diaz-Meco MT and Moscat J, unpublished observations). This observation is counterintuitive with regard to the data supporting roles for Par-4 and PKCζ as tumor suppressors. An explanation for this apparently paradoxical observation is based on the different roles of PKCζ in benign and malignant tumorigenesis, and is clearly linked to its mechanism of action. That is, it is possible that under conditions of hyperplasia in which tumors do not undergo metabolic stress, PKCζ, likely through its role as an activator of Akt79 and NF-κB,14 is necessary for the tumor proliferation and survival state unleashed by the loss of Par-4. That is the case for the development of benign hyperplasia produced in the Par-4 KO mice, which is completely eliminated by the simultaneous inactivation of PKCζ (Diaz-Meco MT and Moscat J, unpublished observations). However, in the case of more aggressive tumors, similar to those triggered by the expression of oncogenic Ras, which are characterized by a high degree of metabolic and nutrient stress, the loss of PKCζ would trigger another pathway that results in enhanced proliferation of the tumor cells49 (Figure 2). The basis for this explanation was obtained in experiments where the growth of PKCζ-deficient Ras-transformed embryonic fibroblasts was investigated under conditions of plentiful nutrient availability, or under conditions of nutrient deficiency.49 The latter situation is more likely to occur in the context of large, aggressive tumors. Interestingly, under conditions of nutrient and mitogen availability, the loss of PKCζ clearly impaired the proliferation of Ras-transformed cells, whereas under conditions of nutrient and mitogen deficiency the opposite phenotype was observed.49 This indicates that PKCζ controls two antagonistic pathways for oncogenic cell proliferation (Figure 2): one that is required under normal conditions, and another that is inhibitory under nutrient-deficient stress situations.
The molecular mechanisms controlling either pathway need to be clarified, but it is interesting to note that the expression of Ras in lung tissues triggers the expression of a myriad of NF-κB-dependent genes in wild-type mice, which is completely inhibited in a PKCζ KO background.49 Despite the inhibition of NF-κB, IL-6 levels are enhanced in Ras-transformed KO tissues and cells, which have been shown to be important in allowing these mutant cells to proliferate under highly aggressive tumorigenic and nutrient-stressed conditions.49 The details of how PKCζ negatively influences IL-6 production still need to be worked out, but are likely to involve epigenetic changes in the IL-6 promoter.49
The aPKCs, Mammalian Cell Polarity, and Cancer
The analysis of Par-4 KO and Par-4/PKCζ DKO mice revealed that PKCζ, in addition to having an important role in NF-κB gene expression, also regulates Akt by direct phosphorylation.79 The ability of PKCζ to influence these two important signaling cascades is of great relevance for the mechanism of action of the Par-4/PKCζ cassette, at least in prostate and lung cancer.79 However, the mechanism whereby PKCλ/ι regulates tumorigenesis in vivo is still a mystery. It is possible that PKCλ/ι could be having a major role in a signaling cascade different from those controlled by PKCζ, involving, for example, the Par-6/Par-3 polarity complex. Recent in vitro experiments link components of the polarity complexes to cancer, suggesting that the overexpression of Par-6 leads to increased growth factor-independent cell proliferation.81 This, in turn, results in the hyperplastic development of polarized cells in three-dimensional acini, because of aPKC-dependent regulation of the MEK/ERK signaling cassette, but surprisingly without affecting cell polarity.81 These results would be consistent with the already known ability of overexpressed aPKCs to modulate MEK82, 83 but are difficult to reconcile with the genetically well-established role of Par-6 in cell polarity, at least in lower organisms.84 In the same vein, the manipulation of the levels of another polarity protein, named Scribble, also leads to changes in ERK activity, but in this case associated with alterations in the polarity phenotype.85 Scribble is particularly interesting as its polarization has been shown to be under the control of PKCλ/ι in KO T cells,29 and has also been shown in in vitro transfection experiments to channel apoptosis signals activated by the interaction of the oncogene ERB2 with the Par-6/aPKC module,86 or by expression of the Myc oncogene in vitro in cell-culture model systems.87 The interaction between Par-6 and PKCλ/ι in the human lung adenocarcinoma cell line A549 has also been suggested to be relevant for transformation, at least in overexpression and co-transfection experiments.88 Therefore, a number of in vitro overexpression experiments implicate different polarity proteins in growth signaling control. However, in vivo demonstrations of the actual role of the aPKCs in cell polarity control in relevant cancer models are, unfortunately, still sorely lacking.
That the only aPKC existing in lower organisms has an important role in polarity is widely accepted.2, 84 In this regard, genetic studies in Drosophila and C. elegans support an important role for their aPKC, which is more closely related to PKCλ/ι than to PKCζ in cell polarity.2, 84 There is also a consensus that the loss of cell polarity could be relevant to carcinogenesis in mammalian cells.84 Surprisingly, although the genetic inactivation of PKCλ/ι in mature T cells gives rise to polarity defects that translate into activation defects in vitro and in vivo,29 PKCλ/ι deficiency in the liver or intestine, or PKCζ deficiency in the lung, does not result in detectable alterations in cell polarity under basal, non-transformed conditions.26, 49, 89 It is believed that polarity loss should increase tumorigenesis and metastasis.90 However, the only available in vivo data implicating PKCλ/ι in cancer shows that its inactivation prevents, rather than enhances, the tumorigenic phenotype of ApcMin/+ mice.61 This important observation goes against the purported role of PKCλ/ι in cell polarity and its impact on promoting transformation when ablated. It is possible that the simple KO of one of the aPKCs is not sufficient to drive the polarity phenotype in certain cell types and tissues, and that the simultaneous inactivation of PKCλ/ι and PKCζ is required for cell polarity effects to become detectable. However, this type of experiment, in which both aPKCs are deleted, could lead to confounding results, as the complete ablation of both aPKCs might lead to restructuring of different PB1 protein complexes, which could artifactually create novel signaling units whose physiological and real pathological significance would be questionable. Probably a better strategy would be to create genetic knock-in models with mutations in polarity genes, which could potentially be identified in unbiased genome-wide genetic screens of human tumor samples. The identification of the PKCζ S514F mutation described above49, 59 combined with the fact that its expression, at least in Ras-expressing xenografts, gives a phenotype consistent with that of the Ras-expressing PKCζ KO lung, might lead the way to genetic experiments that are more physiologically relevant to cancer.
The Role of p62 as a Hub in Cell Signaling
The fact that p62 is a molecule rich in different modular structures suggested, from the time of its initial identification, that it would have a critical role in the organization of cell signaling events that mediate a variety of cell functions6, 91 (Figure 3). Similar to the analysis of the aPKCs and Par-4, the analysis of mutations in humans and the phenotype of the KO mice have enlightened us with regard to its importance and mechanism of action, and have also brought about a new set of questions.
The first indication of a physiological function for p62 came from the discovery that it harbors several mutations associated with Paget's disease of the bone (PDB).92, 93 This suggested that a major role for p62 in humans was likely to be linked to the control of bone homeostasis.94 In fact, p62 KO mice display defects in osteoclastogenesis in vitro as a consequence of inefficient activation of NF-κB in KO osteoclasts activated by RANK-L or TNFα, two critical cytokines in the activation of the osteoclastogenic response in vivo.95 In addition, RANK-L triggers the formation of a p62–aPKC–TRAF6 complex in the pre-osteoclast cell line RAW 264.7 and in primary bone marrow-derived macrophages (BMDMs). However, given that the PKCζ−/− BMDMs, in contrast to BMDMs with PKCλ/ι knocked down, do not have osteoclastogenic defects, this suggests that PKCλ/ι may be the aPKC that acts in a signaling complex with p62 in this pathway95 (Diaz-Meco MT and Moscat J, unpublished observations). Interestingly, the expression of p62 with a PDB mutation resulted in hyperactivated NF-κB and gain-of-function osteoclastogenesis, a phenotype consistent with that of the human disease.96 Therefore, these studies, in combination with others showing a role for p62 in NF-κB activation at the level of the E3 ubiquitin ligase TRAF6 in other systems,9, 91, 97 solidly established p62 as a critical player in the sustained activation of NF-κB. This link between p62 and NF-κB can also explain, at least in part, its role in cancer. Levels of p62 were dramatically increased in several tumor types, especially human lung cancers where more than 60% of lung adenocarcinomas and more than 90% of squamous cell carcinomas displayed elevated p62 protein levels, as determined by the immunohistochemical analysis of tissue microarrays.9 Therefore, the facts that p62 is an activator of NF-κB, that this transcription factor has been shown to have a critical role in cancer,98 and that human tumors have elevated p62 levels, strongly suggested that p62 could be an important player in tumorigenesis. In fact, experiments using the Ras-inducible lung cancer mouse model, discussed above, clearly showed that the p62 KOs show an outstanding resistance to carcinogenesis in this system,9 very likely because of impaired Ras-induced NF-κB activation,9 a critical pathway for Ras-induced survival.99 In this regard, the genetic inactivation of PKCζ also inhibits Ras-induced NF-κB but, in contrast to p62, PKCζ KO mice display enhanced tumorigenesis.49 This is mediated by an overproduction of IL-6 through a κB-independent mechanism.49 Thus, although PKCζ and p62 both control the expression of NF-κB-dependent genes, suggesting that the p62-aPKC module is relevant for Ras-induced NF-κB, the mechanisms whereby they participate in the NF-κB pathway in response to Ras and the final outcome in carcinogenesis are different. That is, unlike PKCζ, p62 is required to activate the IKK complex through the activation of K63-mediated polyubiquitination of TRAF6 and to regulate the nuclear translocation of RelA/p65 in response to Ras.9, 35, 95, 97 PKCζ, on the other hand, can both positively regulate NF-κB at the transcriptional level and, at the same time, exert a negative effect on IL-6 production through a κB-independent pathway, which is responsible for the observed increase in carcinogenesis.14, 20, 49 In this scenario, PKCζ could emerge as a critical step in the generation of inflammatory cytokines that might decide the final outcome of the carcinogenic process. The role of a p62–PKCλ/ι complex in Ras-induced carcinogenesis has not yet been addressed in in vivo mouse models.
In keeping with a role for p62 in cancer and the importance of NF-κB in this process, a recent study showed that knocking down p62 in stroma cells from multiple myeloma (MM) patients significantly inactivated the support of myeloma cell growth, likely because of the reduced production of IL-6, TNFα, and RANK-L by the p62-deficient stroma cells.100 It is noted that aPKC activity was increased in marrow stromal cells from MM patients and that its activity was inhibited on knockdown of p62 in these cells.100 This indicates that the p62–aPKC complex is an important step regulating NF-κB to modulate the MM stromal environment. This is particularly relevant in light of new information implicating NF-κB in multiple myeloma. That is, two laboratories, by using multiple-pronged genomic and gene-expression profiling approaches, have identified NF-κB-activating mutations in one-fifth of several hundred myeloma cell lines and patient samples.101, 102 These include gain-of-function mutations in positive regulators of NF-κB, such as NIK, NFκB1, NFκB2, and receptors of the TNF receptor superfamily, and loss-of-function mutations in genes encoding negative regulators such as TRAF3, TRAF2, and CylD.101, 102 Interestingly, there is also evidence that p62 interacts with CylD, which suggests a potential dual role of p62 in regulating not only the ubiquitination and subsequent activation of NF-κB signaling intermediaries but also its inactivation by deubiquitination through CylD.103, 104
Consistent with this, the analysis of p62 KO mice also suggested a role for this molecule in controlling the accumulation of polyubiquitinated proteins, whose role in cell toxicity and tissue damage has been inferred from observations that they accumulate, along with p62, in several degenerative diseases.105 For example, it has been shown that p62-deficient brains display increased accumulation of polyubiquitinated proteins, which can be because of the impaired proteasome activity103 or decreased deubiquitination associated with p62 deficiency.103 Interestingly, defects in autophagy also result in the accumulation of polyubiquitinated protein aggregates.106, 107 On the other hand, in vivo data in genetically modified mice support a role for autophagy in the control of p62 levels. That is, mice deficient in one of the critical autophagy genes, Atg7, display dramatically increased levels of p62 and an accumulation of polyubiquitinated aggregates that colocalize with p62.108 Surprisingly, Atg7/p62 double KO hepatocytes, but not neurons, lack these aggregates, suggesting that p62 could, under some circumstances, have a still-to-be clarified role in their formation, at least in the liver.108 These results have been interpreted to mean that p62, and possibly another scaffold protein termed NBR1,109 might have structural roles in the formation of these aggregates. The precise role of these aggregates in cell pathophysiology still needs to be clarified. However, an elegant work recently published by Rubinsztein and co-workers110 adds another angle to this problem that complicates the proposed role of p62 as a structural element in the formation of polyubiquitinated aggregates.108, 111, 112 This group showed that the increased accumulation of p62 in autophagy-deficient cells led to the inhibition of the proteasome, which caused the accumulation of polyubiquitinated aggregates.110 Obviously, in the absence of p62, the proteasome is not inhibited and the accumulated polyubiquitinated proteins are efficiently degraded in autophagy-impaired cells,110 potentially explaining the lack of aggregates in the Atg7/p62 double KO mice without the need to invoke a hypothetical role for p62 in the building up of the aggregates.108 This proposed hyperactivation of the proteasome in Atg7/p62-deficient livers would be in agreement with the observation that the amount of polyubiquitinated proteins accumulated in that organ in these doubly mutant mice, as determined by SDS-PAGE, is lower than that in the single Atg7-deficient mice.108
One possibility is that p62, by binding the autophagy adapter LC3 and ubiquitinated proteins, brings aggregated or misfolded proteins to the autophagosome for their degradation.113, 114, 115 Recent RNA interference and overexpression experiments in several cell culture systems suggest that possibility.113, 114, 115 If this were the case, it would be very interesting because it would mean that cells have evolved a p62-dependent mechanism to dispose of excess proteins in the absence of a functional proteasome. However, it still needs to be clarified whether this putative p62-mediated autophagy mechanism is functional under physiological conditions, whether it is specific to pathological situations in which the proteasome is inhibited, or whether it is just an artifact of cell culture conditions. Again, this is complicated because of the connection of p62 to the proteasome, wherein p62 has been shown to have a critical role in the delivery of K63-polyubiquitinated proteins for degradation through a p62–S5a interaction.116
It appears that the proteasome cannot degrade insoluble misfolded proteins, which suggests that cellular surveillance pathways must exist to modulate protein quality control independent of the proteasome. There is the potential for p62 to be involved in such a mechanism; however, solid genetic evidence for this still needs to be produced. In this regard, recent findings reveal that the co-chaperones BAG1 and BAG3 are both involved in regulating the proteasomal and autophagic pathways,117 where the BAG3/BAG1 ratio maintains appropriate protein homeostasis. Clearly then, the state of protein aggregation, which is a critical determinant of its solubility, and the degree of polyubiquitination are key regulatory factors that enable the sorting and clearance of protein aggregates. Thus, as the cell ages and experiences a decline in proteasome activity, it might have to rely increasingly on autophagy. In this scenario, the levels of p62 would confer a protective role by reducing the toxicity of these misfolded proteins. In vitro studies reveal that K63-linked ubiquitin may be preserved as a signal to selectively facilitate the clearance of tau and SOD-1 inclusions by autophagy.118 With the development of antibodies selective for K63 modification, recent studies have revealed that the accumulation of inclusions that are immunopositive for K63-ubiquitin chains characterizes a genetic mouse model of 26S proteasome depletion, as well as human diseases involving proteasome function deficiencies, such as Alzheimer's, Parkinson's, and Huntington's disease.119 The p62-mediated recruitment of misfolded proteins to autophagosomes could also involve the cooperation of other signals such as acetylation,120 although further studies will be needed to define the exact means whereby this occurs. In situations such as cancer where p62 is upregulated, 9, 121, 122, 123 cells could continue to survive and escape death thanks to the ability of p62 to activate NF-κB and to dispose of toxic aggregates, whereas, in a neurodegenerative disease such as Alzheimer's, where p62 levels are downregulated,124 cells would die because of the accumulation of misfolded toxic proteins and a lower level of pro-survival NF-κB. Therefore, p62 could be having a decisive role in a hypothetical decision step that determines whether polyubiquitinated proteins reach a critical threshold to form aggregates or whether they are degraded through the autophagy or proteasome routes. The importance of this point of intersection in vivo, as well as its relationship to health or disease, needs to be clarified with further genetic studies in physiologically relevant organismal systems. In any case, all the literature surrounding the ‘aggregate question’ needs urgent clarification. That is, we need to understand which are the ‘good aggregates,’ and which are the ‘toxic assets.’ For example, the recent observation that Atg7 deficiency leads to hepatotoxicity, which is reduced in the double Atg7/p62 KO mice, is at odds with the idea that p62 is necessary to package polyubiquitinated aggregates to make them harmless, as the double-mutant livers do not develop the aggregates that are observed in Atg7-deficient mice.108 In this regard, recent attempts to classify and understand the nature of these aggregates in yeast constitute one of the first serious efforts to better understand their actual roles and modes of action.125 This, along with the clarification of the role played by the proteasome and DUBs in these processes, is absolutely necessary if we are to firmly establish a role for p62 in aggregate homeostasis.
Another potential function for p62 was suggested by several studies that position p62 as a key component in receptor turnover. This idea links back to the first report of p62 localization to the endosome–lysosome pathway.126 That is, p62, through its UBA domain, associates with polyubiquitinated receptors, such as TrkA, and remains co-associated with the stage of late endosomes, although the receptor itself is degraded by lysosomes.126 Early in the sorting pathway for TrkA, the receptor can be deubiquitinated by CylD through recruitment to the p62 scaffold,127 resulting in receptor recycling. On the other hand, if a K63-polyubiquitinated receptor escapes this checkpoint, the chains are removed by proteasomal DUBs that associate with multivesicular bodies.126 Recent observations further substantiate the finding that proteasomal DUBs are capable of degrading K63 chains.128, 129 In keeping with these data, mice deficient in p62 hyperaccumulate K63 chains and TrkA itself.103 In this trafficking network, p62 remains co-associated and is required for presentation of the K63-polyubiquitinated receptor to the proteasomal DUBs. Thus, p62 appears to be necessary for the trafficking of cargo through the endosomal–lysosomal network. Interestingly, mutation in CHMP2B, a protein involved in endosome sorting, results in the accumulation of ubiquitinated proteins and p62, similar to what is seen in human disease.130 Together, these findings, along with recent data from the Tooze laboratory,131 lend further support to the temporal co-association of p62 with cargo early in the endosomal–lysosomal trafficking pathway and could therefore position p62 at the crossroads during the formation of autophagosomes in certain pathophysiological situations.
Taken together, all these published observations suggest that, indeed, p62 could have a more generalized role in trafficking of a wide array of substrates targeted for autophagy or lysosomal degradation. Although autophagy was once thought to be a relatively nonselective process, it is possible that p62 could provide selectivity through capture of specific substrates. Interestingly, p62 has recently been implicated in the degradation of peroxisomes132 and has been observed in the midbody along with the ubiquitin-related protein Atg8 during abscission, suggesting that autophagy may be coupled to cytokinesis.133 Given the conserved nature of the p62 function in flies and humans,112 as well as the diversity in the physiological processes connected to the function of p62, it is clear that this protein serves as a rheostat to fine-tune multiple pathways that impinge on survival or death. If confirmed, p62's role would redefine autophagy as a selective process. The task at hand will be to identify, in physiologically relevant models, specific substrates that interact with p62 to be degraded by autophagy and the regulatory mechanisms that contribute to those interactions. In this regard, the other modules in p62, and its partners, the aPKCs, NBR1, ERK, MEK5, TRAF6, and CylD, could also have roles in this newly proposed p62 function, although the genetic evidence available so far does not support their involvement in this process (Figure 4).
On the other hand, it is possible that, rather than having a role in autophagy, p62 signaling might be modulated by this cellular process, which, because of p62's role as a pro-survival protein, could serve to fine-tune the decision of the stressed cells to live or die. Thus, although it is clear that the proposed role for p62 in the formation of the autophagic vesicles113 is not supported by the genetic in vivo evidence from the p62 KO mice,108 nevertheless, autophagy could be a mechanism to regulate p62-mediated signaling, as has recently been proposed for the transcriptional regulator MEF2D.134 In this regard, the interaction of p62 with the E3 ubiquitin ligase TRAF6, which serves to promote the K-63 polyubiquitination of important intermediaries, such as TRAF6 itself,97 IKKγ,38 or TrkA,127 could be inactivated if p62 is degraded during autophagy, which could result in the shutting down of the survival signaling pathways regulated by p62 through NF-κB, accelerating autophagic cell death. This could be relevant under conditions in which apoptosis is partially inhibited, for example in certain tumors. In addition, hypoxia-induced autophagy has been shown to deplete cellular levels of p62,135, 136 which triggers ERK activation that might affect cell growth and survival during hypoxic conditions.135 Interestingly, these studies are in good agreement with previous data showing the negative regulation of ERK by p62 and its role in obesity in p62-deficient mice.137
Consistent with the notion that p62 is a ‘signaling organizer’ rather than ‘garbage disposal’ are the very recent data from the Ashkenazi laboratory.138 Interestingly, they show that p62 cytosolic speckles constitute receiving entities of a signaling complex made up of the TRAIL receptor and a ubiquitinated form of Caspase 8.138 The interaction of the Caspase 8 group with the p62 complex is necessary for the generation of higher-order Caspase 8 entities, which is required for TRAIL-induced Caspase 3 activation and apoptosis.138 These results are reminiscent of previously published observations showing the formation of aggregate complexes including p62, TRAF6, and the kinase-adapter IRAK, which is necessary for a full activation of NF-κB.35 According to this model, p62's role would be to organize different signaling complexes involving ubiquitinated signaling-relevant proteins that will determine the decision of cells to survive or undergo apoptosis. The fact that p62 can be degraded by autophagy suggests a mechanism whereby the cell could terminate these cascades under different situations, which may have pathophysiological implications in, for example, cancer. Whether the other components of the p62 hub such as the aPKCs are also involved in these ‘signaling aggregates,’ need to be clarified with further experimentation and will be essential for potential therapeutic intervention that would target the p62 ‘signaling organizer’ complex.
Concluding Remarks
The aPKCs, PKCζ and PKCλ/ι, assemble in different signaling complexes through their interactions with scaffold proteins and regulators such as Par-6, p62, and Par-4. This generates two kinds of complexes depending on the domains of the aPKCs involved. That is, the aPKCs interact through their PB1 domains to form complexes with Par-6 and p62, whereas the complex formed with Par-4 is mediated by the aPKC's zinc-finger domain. These interactions set in place a signaling network that serves to confer specificity and plasticity to the actions of these kinases in the control of different key cellular processes. Recent data from knockout mice deficient in different components of this network are helping to shed light on the roles of these signaling molecules in physiologically relevant processes, such as cancer initiation and progression. However, there are still many open questions that need to be addressed in the future. For example, what are the factors that determine which complex is formed at a given time and within a specific cell context and is there cross talk between the different complexes? Similarly, the in vivo roles of the different components of the complexes need to be addressed more systematically in the same cellular contexts to allow for a direct comparison of the different biological outcomes and phenotypes. Once the in vivo functions of each component have been identified, such as the recent advances in understanding the role of p62, the task at hand will be to discern the makeup of the relevant signaling complexes in each physiological process and the contributions of the different binding partners to these functions.
Abbreviations
- APC:
-
adenomatous polyposis coli gene
- aPKC:
-
atypical protein kinase C
- BAG1/3:
-
Bcl-2-associated athanogene-1/3
- CHMP2B:
-
charged multivesicular body protein 2B
- CYLD:
-
cylindromatosis gene
- DKO:
-
double knockout
- DUB:
-
deubiquitinating enzyme
- ERK:
-
extracellular responsive kinase
- IHC:
-
immunohistochemistry
- IκB:
-
inhibitor of κB
- IKK:
-
IκB kinase
- IL:
-
interleukin
- IRAK:
-
interleukin receptor-associated kinase
- IRS:
-
insulin receptor substrate
- Jak1:
-
Janus kinase-1
- KO:
-
knockout
- LC3:
-
microtubule-associated protein 1 light chain 3/MAP1LC3
- MEF2:
-
myocyte enhancer factor 2
- MEK:
-
MAPK/ERK kinase
- MEKK:
-
MEK kinase
- MM:
-
multiple myeloma
- NBR1:
-
next to BRCA1
- NF-AT:
-
nuclear factor of activated T cells
- NF-κB:
-
nuclear factor-B
- NIK:
-
NFκB-inducing kinase
- NSCLC:
-
non-small cell lung cancer OPCA-OPR-PC and AID
- p62/SQSTM1:
-
sequestosome-1
- OVA:
-
ovalbumin
- Par-4:
-
prostate androgen responsive-4
- Par-6:
-
partitioning defective-6
- PB1:
-
phox and Bem1p
- PDB:
-
Paget's disease of bone
- PIK3CA, phosphoinositide-3-kinase catalytic alpha polypeptidePI3KR5:
-
phosphoinositide-3-kinase regulatory subunit-5
- PKC:
-
protein kinase C
- PTEN:
-
tensin homolog deleted in chromosome ten
- RANK-L:
-
receptor activator of NF-κB-ligand
- RHEB:
-
Ras homolog enriched in brain
- RPS6KB1:
-
p70s6k ribosomal protein S6 kinase
- SOD:
-
superoxide dismutase
- Th:
-
T helper
- TNF:
-
tumor necrosis factor
- TRAF:
-
TNF receptor-associated factor
- TRAIL:
-
tumor necrosis factor-related apoptosis-inducing ligand
- TrkA:
-
tropomyosin-receptor- kinase
- UBA:
-
ubiquitin associated
- XIAP:
-
X-linked mammalian inhibitor of apoptosis protein
References
Nishizuka Y . Protein kinase C and lipid signaling for sustained cellular responses. FASEB J 1995; 9: 484–496.
Moscat J, Diaz-Meco MT, Albert A, Campuzano S . Cell Signaling and Function Organized by PB1 Domain Interactions. Mol Cell 2006; 23: 631–640.
Moscat J, Diaz-Meco MT . The atypical protein kinase Cs. Functional specificity mediated by specific protein adapters. EMBO Rep 2000; 1: 399–403.
Sumimoto H, Kamakura S, Ito T . Structure and function of the PB1 domain, a protein interaction module conserved in animals, fungi, amoebas, and plants. Sci STKE 2007; re6.
Puls A, Schmidt S, Grawe F, Stabel S . Interaction of protein kinase C zeta with ZIP, a novel protein kinase C- binding protein. Proc Natl Acad Sci USA 1997; 94: 6191–6196.
Sanchez P, De Carcer G, Sandoval IV, Moscat J, Diaz-Meco MT . Localization of atypical protein kinase C isoforms into lysosome- targeted endosomes through interaction with p62. Mol Cell Biol 1998; 18: 3069–3080.
Macara IG . Par proteins: partners in polarization. Curr Biol 2004; 14: R160–R162.
Ohno S . Intercellular junctions and cellular polarity: the PAR-aPKC complex, a conserved core cassette playing fundamental roles in cell polarity. Curr Opin Cell Biol 2001; 13: 641–648.
Duran A, Linares JF, Galvez AS, Wikenheiser K, Flores JM, Diaz-Meco MT et al. The signaling adaptor p62 is an important NF-kappaB mediator in tumorigenesis. Cancer Cell 2008; 13: 343–354.
Macek B, Benda C, Jestel A, Maskos K, Mann M, Messerschmidt A . Phosphorylation of the human full-length protein kinase Ciota. J Proteome Res 2008; 7: 2928–2935.
Dominguez I, Diaz-Meco MT, Municio MM, Berra E, Garcia de Herreros A, Cornet ME et al. Evidence for a role of protein kinase C zeta subspecies in maturation of Xenopus laevis oocytes. Mol Cell Biol 1992; 12: 3776–3783.
Dominguez I, Sanz L, Arenzana-Seisdedos F, Diaz-Meco MT, Virelizier JL, Moscat J . Inhibition of protein kinase C zeta subspecies blocks the activation of an NF-kappa B-like activity in Xenopus laevis oocytes. Mol Cell Biol 1993; 13: 1290–1295.
Soloff RS, Katayama C, Lin MY, Feramisco JR, Hedrick SM . Targeted deletion of protein kinase C lambda reveals a distribution of functions between the two atypical protein kinase C isoforms. J Immunol 2004; 173: 3250–3260.
Leitges M, Sanz L, Martin P, Duran A, Braun U, Garcia JF et al. Targeted disruption of the zetaPKC gene results in the impairment of the NF-kappaB pathway. Mol Cell 2001; 8: 771–780.
Martin P, Duran A, Minguet S, Gaspar ML, Diaz-Meco MT, Rennert P et al. Role of zeta PKC in B-cell signaling and function. EMBO J 2002; 21: 4049–4057.
Martin P, Villares R, Rodriguez-Mascarenhas S, Zaballos A, Leitges M, Kovac J et al. Control of T helper 2 cell function and allergic airway inflammation by PKC{zeta}. Proc Natl Acad Sci USA 2005; 102: 9866–9871.
Moscat J, Diaz-Meco MT, Rennert P . NF-kappaB activation by protein kinase C isoforms and B-cell function. EMBO Rep 2003; 4: 31–36.
Diaz-Meco MT, Berra E, Municio MM, Sanz L, Lozano J, Dominguez I et al. A dominant negative protein kinase C zeta subspecies blocks NF-kappa B activation. Mol Cell Biol 1993; 13: 4770–4775.
Anrather J, Csizmadia V, Soares MP, Winkler H . Regulation of NF-kappaB RelA phosphorylation and transcriptional activity by p21(ras) and protein kinase Czeta in primary endothelial cells. J Biol Chem 1999; 274: 13594–13603.
Duran A, Diaz-Meco MT, Moscat J . Essential role of RelA Ser311 phosphorylation by zetaPKC in NF-kappaB transcriptional activation. EMBO J 2003; 22: 3910–3918.
Tabuse Y, Izumi Y, Piano F, Kemphues KJ, Miwa J, Ohno S . Atypical protein kinase C cooperates with PAR-3 to establish embryonic polarity in Caenorhabditis elegans. Development 1998; 125: 3607–3614.
Wodarz A, Ramrath A, Kuchinke U, Knust E . Bazooka provides an apical cue for Inscuteable localization in Drosophila neuroblasts. Nature 1999; 402: 544–547.
Cox DN, Seyfried SA, Jan LY, Jan YN . Bazooka and atypical protein kinase C are required to regulate oocyte differentiation in the Drosophila ovary. Proc Natl Acad Sci USA 2001; 98: 14475–14480.
Cai Y, Yu F, Lin S, Chia W, Yang X . Apical complex genes control mitotic spindle geometry and relative size of daughter cells in Drosophila neuroblast and pI asymmetric divisions. Cell 2003; 112: 51–62.
Betschinger J, Mechtler K, Knoblich JA . The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature 2003; 422: 326–330.
Matsumoto M, Ogawa W, Akimoto K, Inoue H, Miyake K, Furukawa K et al. PKClambda in liver mediates insulin-induced SREBP-1c expression and determines both hepatic lipid content and overall insulin sensitivity. J Clin Invest 2003; 112: 935–944.
Hashimoto N, Kido Y, Uchida T, Matsuda T, Suzuki K, Inoue H et al. PKClambda regulates glucose-induced insulin secretion through modulation of gene expression in pancreatic beta cells. J Clin Invest 2005; 115: 138–145.
Farese RV, Sajan MP, Yang H, Li P, Mastorides S, Gower Jr WR et al. Muscle-specific knockout of PKC-lambda impairs glucose transport and induces metabolic and diabetic syndromes. J Clin Invest 2007; 117: 2289–2301.
Yang JQ, Leitges M, Duran A, Diaz-Meco MT, Moscat J . Loss of PKC lambda/iota impairs Th2 establishment and allergic airway inflammation in vivo. Proc Natl Acad Sci USA 2009; 106: 1099–1104.
Moscat J, Rennert P, Diaz-Meco MT . PKCzeta at the crossroad of NF-kappaB and Jak1/Stat6 signaling pathways. Cell Death Differ 2006; 13: 702–711.
Garcia-Cao I, Lafuente M, Criado L, Diaz-Meco M, Serrano M, Moscat J . Genetic inactivation of Par4 results in hyperactivation of NF-κB and impairment of JNK and p38. EMBO Rep 2003; 4: 307–312.
Lafuente MJ, Martin P, Garcia-Cao I, Diaz-Meco MT, Serrano M, Moscat J . Regulation of mature T lymphocyte proliferation and differentiation by Par-4. EMBO J 2003; 22: 4689–4698.
Duran A, Rodriguez A, Martin P, Serrano M, Flores JM, Leitges M et al. Crosstalk between PKCzeta and the IL4/Stat6 pathway during T-cell-mediated hepatitis. EMBO J 2004; 23: 4595–4605.
Sanz L, Sanchez P, Lallena MJ, Diaz-Meco MT, Moscat J . The interaction of p62 with RIP links the atypical PKCs to NF-kappaB activation. EMBO J 1999; 18: 3044–3053.
Sanz L, Diaz-Meco MT, Nakano H, Moscat J . The atypical PKC-interacting protein p62 channels NF-kappaB activation by the IL-1-TRAF6 pathway. EMBO J 2000; 19: 1576–1586.
Wooten MW, Seibenhener ML, Mamidipudi V, Diaz-Meco MT, Barker PA, Moscat J . The atypical protein kinase C-interacting protein p62 is a scaffold for NF-kappaB activation by nerve growth factor. J Biol Chem 2001; 276: 7709–7712.
Avila A, Silverman N, Diaz-Meco MT, Moscat J . The Drosophila atypical protein kinase C-ref(2)p complex constitutes a conserved module for signaling in the toll pathway. Mol Cell Biol 2002; 22: 8787–8795.
Martin P, Diaz-Meco MT, Moscat J . The signaling adapter p62 is an important mediator of T helper 2 cell function and allergic airway inflammation. EMBO J 2006; 25: 3524–3533.
Ozdamar B, Bose R, Barrios-Rodiles M, Wang HR, Zhang Y, Wrana JL . Regulation of the polarity protein Par6 by TGFbeta receptors controls epithelial cell plasticity. Science 2005; 307: 1603–1609.
Ludford-Menting MJ, Oliaro J, Sacirbegovic F, Cheah ET, Pedersen N, Thomas SJ et al. A network of PDZ-containing proteins regulates T cell polarity and morphology during migration and immunological synapse formation. Immunity 2005; 22: 737–748.
Shi SH, Jan LY, Jan YN . Hippocampal neuronal polarity specified by spatially localized mPar3/mPar6 and PI 3-kinase activity. Cell 2003; 112: 63–75.
Etienne-Manneville S, Hall A . Cell polarity: Par6, aPKC and cytoskeletal crosstalk. Curr Opin Cell Biol 2003; 15: 67–72.
Sun L, Hui AM, Su Q, Vortmeyer A, Kotliarov Y, Pastorino S et al. Neuronal and glioma-derived stem cell factor induces angiogenesis within the brain. Cancer Cell 2006; 9: 287–300.
French PJ, Swagemakers SM, Nagel JH, Kouwenhoven MC, Brouwer E, van der Spek P et al. Gene expression profiles associated with treatment response in oligodendrogliomas. Cancer Res 2005; 65: 11335–11344.
Rickman DS, Bobek MP, Misek DE, Kuick R, Blaivas M, Kurnit DM et al. Distinctive molecular profiles of high-grade and low-grade gliomas based on oligonucleotide microarray analysis. Cancer Res 2001; 61: 6885–6891.
Bredel M, Bredel C, Juric D, Harsh GR, Vogel H, Recht LD et al. Functional network analysis reveals extended gliomagenesis pathway maps and three novel MYC-interacting genes in human gliomas. Cancer Res 2005; 65: 8679–8689.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci USA 2001; 98: 13790–13795.
Stearman RS, Dwyer-Nield L, Zerbe L, Blaine SA, Chan Z, Bunn Jr PA et al. Analysis of orthologous gene expression between human pulmonary adenocarcinoma and a carcinogen-induced murine model. Am J Pathol 2005; 167: 1763–1775.
Galvez AS, Duran A, Linares JF, Pathrose P, Castilla EA, Abu-Baker S et al. Protein kinase Czeta represses the interleukin-6 promoter and impairs tumorigenesis in vivo. Mol Cell Biol 2009; 29: 104–115.
Lenburg ME, Liou LS, Gerry NP, Frampton GM, Cohen HT, Christman MF . Previously unidentified changes in renal cell carcinoma gene expression identified by parametric analysis of microarray data. BMC Cancer 2003; 3: 31.
Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J et al. Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res 2005; 11: 7234–7242.
Buchholz M, Braun M, Heidenblut A, Kestler HA, Kloppel G, Schmiegel W et al. Transcriptome analysis of microdissected pancreatic intraepithelial neoplastic lesions. Oncogene 2005; 24: 6626–6636.
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 2007; 9: 166–180.
Dhanasekaran SM, Dash A, Yu J, Maine IP, Laxman B, Tomlins SA et al. Molecular profiling of human prostate tissues: insights into gene expression patterns of prostate development during puberty. FASEB J 2005; 19: 243–245.
Sanchez-Carbayo M, Socci ND, Lozano J, Saint F, Cordon-Cardo C . Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J Clin Oncol 2006; 24: 778–789.
Dyrskjot L, Kruhoffer M, Thykjaer T, Marcussen N, Jensen JL, Moller K et al. Gene expression in the urinary bladder: a common carcinoma in situ gene expression signature exists disregarding histopathological classification. Cancer Res 2004; 64: 4040–4048.
Basso K, Margolin AA, Stolovitzky G, Klein U, Dalla-Favera R, Califano A . Reverse engineering of regulatory networks in human B cells. Nat Genet 2005; 37: 382–390.
Dave SS, Fu K, Wright GW, Lam LT, Kluin P, Boerma EJ et al. Molecular diagnosis of Burkitt's lymphoma. N Engl J Med 2006; 354: 2431–2442.
Wood LD, Parsons DW, Jones S, Lin J, Sjoblom T, Leary RJ et al. The genomic landscapes of human breast and colorectal cancers. Science 2007; 318: 1108–1113.
Fisher GH, Wellen SL, Klimstra D, Lenczowski JM, Tichelaar JW, Lizak MJ et al. Induction and apoptotic regression of lung adenocarcinomas by regulation of a K-Ras transgene in the presence and absence of tumor suppressor genes. Genes Dev 2001; 15: 3249–3262.
Murray NR, Weems J, Braun U, Leitges M, Fields AP . Protein kinase C betaII and PKCiota/lambda: collaborating partners in colon cancer promotion and progression. Cancer Res 2009; 69: 656–662.
Roffey J, Rosse C, Linch M, Hibbert A, McDonald NQ, Parker PJ . Protein kinase C intervention-the state of play. Curr Opin Cell Biol 2009; 21: 268–279.
Regala RP, Weems C, Jamieson L, Khoor A, Edell ES, Lohse CM et al. Atypical protein kinase C iota is an oncogene in human non-small cell lung cancer. Cancer Res 2005; 65: 8905–8911.
Talbot SG, Estilo C, Maghami E, Sarkaria IS, Pham DK, O-charoenrat P et al. Gene expression profiling allows distinction between primary and metastatic squamous cell carcinomas in the lung. Cancer Res 2005; 65: 3063–3071.
Zhang L, Huang J, Yang N, Liang S, Barchetti A, Giannakakis A et al. Integrative genomic analysis of protein kinase C (PKC) family identifies PKCiota as a biomarker and potential oncogene in ovarian carcinoma. Cancer Res 2006; 66: 4627–4635.
Eder AM, Sui X, Rosen DG, Nolden LK, Cheng KW, Lahad JP et al. Atypical PKC{iota} contributes to poor prognosis through loss of apical-basal polarity and Cyclin E overexpression in ovarian cancer. Proc Natl Acad Sci USA 2005; 102: 12519–12524.
Welsh JB, Zarrinkar PP, Sapinoso LM, Kern SG, Behling CA, Monk BJ et al. Analysis of gene expression profiles in normal and neoplastic ovarian tissue samples identifies candidate molecular markers of epithelial ovarian cancer. Proc Natl Acad Sci USA 2001; 98: 1176–1181.
Lu KH, Patterson AP, Wang L, Marquez RT, Atkinson EN, Baggerly KA et al. Selection of potential markers for epithelial ovarian cancer with gene expression arrays and recursive descent partition analysis. Clin Cancer Res 2004; 10: 3291–3300.
Kojima Y, Akimoto K, Nagashima Y, Ishiguro H, Shirai S, Chishima T et al. The overexpression and altered localization of the atypical protein kinase C lambda/iota in breast cancer correlates with the pathologic type of these tumors. Hum Pathol 2008; 39: 824–831.
Yang YL, Chu JY, Luo ML, Wu YP, Zhang Y, Feng YB et al. Amplification of PRKCI, located in 3q26, is associated with lymph node metastasis in esophageal squamous cell carcinoma. Genes Chromosomes Cancer 2008; 47: 127–136.
Yu YP, Landsittel D, Jing L, Nelson J, Ren B, Liu L et al. Gene expression alterations in prostate cancer predicting tumor aggression and preceding development of malignancy. J Clin Oncol 2004; 22: 2790–2799.
Segal NH, Pavlidis P, Antonescu CR, Maki RG, Noble WS, DeSantis D et al. Classification and subtype prediction of adult soft tissue sarcoma by functional genomics. Am J Pathol 2003; 163: 691–700.
Zhan F, Hardin J, Kordsmeier B, Bumm K, Zheng M, Tian E et al. Global gene expression profiling of multiple myeloma, monoclonal gammopathy of undetermined significance, and normal bone marrow plasma cells. Blood 2002; 99: 1745–1757.
Diaz-Meco MT, Municio MM, Frutos S, Sanchez P, Lozano J, Sanz L et al. The product of par-4, a gene induced during apoptosis, interacts selectively with the atypical isoforms of protein kinase C. Cell 1996; 86: 777–786.
Moreno-Bueno G, Fernandez-Marcos PJ, Collado M, Tendero MJ, Rodriguez-Pinilla SM, Garcia-Cao I et al. Inactivation of the candidate tumor suppressor par-4 in endometrial cancer. Cancer Res 2007; 67: 1927–1934.
Garcia-Cao I, Duran A, Collado M, Carrascosa MJ, Martin-Caballero J, Flores JM et al. Tumour-suppression activity of the proapoptotic regulator Par4. EMBO Rep 2005; 6: 577–583.
McMenamin ME, Soung P, Perera S, Kaplan I, Loda M, Sellers WR . Loss of PTEN expression in paraffin-embedded primary prostate cancer correlates with high Gleason score and advanced stage. Cancer Res 1999; 59: 4291–4296.
Suzuki H, Freije D, Nusskern DR, Okami K, Cairns P, Sidransky D et al. Interfocal heterogeneity of PTEN/MMAC1 gene alterations in multiple metastatic prostate cancer tissues. Cancer Res 1998; 58: 204–209.
Joshi J, Fernandez-Marcos PJ, Galvez A, Amanchy R, Linares JF, Duran A et al. Par-4 inhibits Akt and suppresses Ras-induced lung tumorigenesis. EMBO J 2008; 27: 2181–2193.
Tuveson DA, Shaw AT, Willis NA, Silver DP, Jackson EL, Chang S et al. Endogenous oncogenic K-ras(G12D) stimulates proliferation and widespread neoplastic and developmental defects. Cancer Cell 2004; 5: 375–387.
Nolan ME, Aranda V, Lee S, Lakshmi B, Basu S, Allred DC et al. The polarity protein Par6 induces cell proliferation and is overexpressed in breast cancer. Cancer Res 2008; 68: 8201–8209.
Berra E, Diaz-Meco MT, Lozano J, Frutos S, Municio MM, Sanchez P et al. Evidence for a role of MEK and MAPK during signal transduction by protein kinase C zeta. EMBO J 1995; 14: 6157–6163.
Wooten MW, Seibenhener ML, Neidigh KB, Vandenplas ML . Mapping of atypical protein kinase C within the nerve growth factor signaling cascade: relationship to differentiation and survival of PC12 cells. Mol Cell Biol 2000; 20: 4494–4504.
Suzuki A, Ohno S . The PAR-aPKC system: lessons in polarity. J Cell Sci 2006; 119 (Part 6): 979–987.
Dow LE, Elsum IA, King CL, Kinross KM, Richardson HE, Humbert PO . Loss of human Scribble cooperates with H-Ras to promote cell invasion through deregulation of MAPK signalling. Oncogene 2008; 27: 5988–6001.
Aranda V, Haire T, Nolan ME, Calarco JP, Rosenberg AZ, Fawcett JP et al. Par6-aPKC uncouples ErbB2 induced disruption of polarized epithelial organization from proliferation control. Nat Cell Biol 2006; 8: 1235–1245.
Zhan L, Rosenberg A, Bergami KC, Yu M, Xuan Z, Jaffe AB et al. Deregulation of scribble promotes mammary tumorigenesis and reveals a role for cell polarity in carcinoma. Cell 2008; 135: 865–878.
Frederick LA, Matthews JA, Jamieson L, Justilien V, Thompson EA, Radisky DC et al. Matrix metalloproteinase-10 is a critical effector of protein kinase Ciota-Par6alpha-mediated lung cancer. Oncogene 2008; 27: 4841–4853.
Murray NR, Jamieson L, Yu W, Zhang J, Gokmen-Polar Y, Sier D et al. Protein kinase Ciota is required for Ras transformation and colon carcinogenesis in vivo. J Cell Biol 2004; 164: 797–802.
Etienne-Manneville S . Polarity proteins in migration and invasion. Oncogene 2008; 27: 6970–6980.
Moscat J, Diaz-Meco MT, Wooten MW . Signal integration and diversification through the p62 scaffold protein. Trends Biochem Sci 2007; 32: 95–100.
Laurin N, Brown JP, Morissette J, Raymond V . Recurrent mutation of the gene encoding sequestosome 1 (SQSTM1/p62) in Paget disease of bone. Am J Hum Genet 2002; 70: 1582–1588.
Morissette J, Laurin N, Brown JP . Sequestosome 1: mutation frequencies, haplotypes, and phenotypes in familial Paget's disease of bone. J Bone Miner Res 2006; 21 (Suppl 2): P38–P44.
Leach RJ, Singer FR, Ench Y, Wisdom JH, Pina DS, Johnson-Pais TL . Clinical and cellular phenotypes associated with sequestosome 1 (SQSTM1) mutations. J Bone Miner Res 2006; 21 (Suppl 2): P45–P50.
Duran A, Serrano M, Leitges M, Flores JM, Picard S, Brown JP et al. The atypical PKC-interacting protein p62 is an important mediator of RANK-activated osteoclastogenesis. Dev Cell 2004; 6: 303–309.
Kurihara N, Hiruma Y, Zhou H, Subler MA, Dempster DW, Singer FR et al. Mutation of the sequestosome 1 (p62) gene increases osteoclastogenesis but does not induce Paget disease. J Clin Invest 2007; 117: 133–142.
Wooten MW, Geetha T, Seibenhener ML, Babu JR, Diaz-Meco MT, Moscat J . The p62 scaffold regulates nerve growth factor-induced NF-kappaB activation by influencing TRAF6 polyubiquitination. J Biol Chem 2005; 280: 35625–35629.
Baud V, Karin M . Is NF-kappaB a good target for cancer therapy? Hopes and pitfalls. Nat Rev Drug Discov 2009; 8: 33–40.
Mayo MW, Wang CY, Cogswell PC, Rogers-Graham KS, Lowe SW, Der CJ et al. Requirement of NF-kappaB activation to suppress p53-independent apoptosis induced by oncogenic Ras. Science 1997; 278 (5344): 1812–1815.
Hiruma Y, Honjo T, Jelinek DF, Windle JJ, Shin J, Roodman GD et al. Increased signaling through p62 in the marrow microenvironment increases myeloma cell growth and osteoclast formation. Blood 2009; 113: 4894–4902.
Annunziata CM, Davis RE, Demchenko Y, Bellamy W, Gabrea A, Zhan F et al. Frequent engagement of the classical and alternative NF-kappaB pathways by diverse genetic abnormalities in multiple myeloma. Cancer Cell 2007; 12: 115–130.
Keats JJ, Fonseca R, Chesi M, Schop R, Baker A, Chng WJ et al. Promiscuous mutations activate the noncanonical NF-kappaB pathway in multiple myeloma. Cancer Cell 2007; 12: 131–144.
Wooten MW, Geetha T, Babu JR, Seibenhener ML, Peng J, Cox N et al. Essential role of sequestosome 1/p62 in regulating accumulation of Lys63-ubiquitinated proteins. J Biol Chem 2008; 283: 6783–6789.
Jin W, Chang M, Paul EM, Babu G, Lee AJ, Reiley W et al. Deubiquitinating enzyme CYLD negatively regulates RANK signaling and osteoclastogenesis in mice. J Clin Invest 2008; 118: 1858–1866.
Zatloukal K, Stumptner C, Fuchsbichler A, Heid H, Schnoelzer M, Kenner L et al. p62 Is a common component of cytoplasmic inclusions in protein aggregation diseases. Am J Pathol 2002; 160: 255–263.
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 2006; 441: 880–884.
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006; 441: 885–889.
Komatsu M, Waguri S, Koike M, Sou YS, Ueno T, Hara T et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 2007; 131: 1149–1163.
Kirkin V, Lamark T, Sou YS, Bjorkoy G, Nunn JL, Bruun JA et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell 2009; 33: 505–516.
Korolchuk VI, Mansilla A, Menzies FM, Rubinsztein DC . Autophagy inhibition compromises degradation of ubiquitin-proteasome pathway substrates. Mol Cell 2009; 33: 517–527.
Bjorkoy G, Lamark T, Johansen T . p62/SQSTM1: a missing link between protein aggregates and the autophagy machinery. Autophagy 2006; 2: 138–139.
Nezis IP, Simonsen A, Sagona AP, Finley K, Gaumer S, Contamine D et al. Ref(2)P, the Drosophila melanogaster homologue of mammalian p62, is required for the formation of protein aggregates in adult brain. J Cell Biol 2008; 180: 1065–1071.
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 2005; 171: 603–614.
Pankiv S, Clausen TH, Lamark T, Brech A, Bruun JA, Outzen H et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 2007; 282: 24131–24145.
Shvets E, Fass E, Scherz-Shouval R, Elazar Z . The N-terminus and Phe52 residue of LC3 recruit p62/SQSTM1 into autophagosomes. J Cell Sci 2008; 121 (Part 16): 2685–2695.
Seibenhener ML, Babu JR, Geetha T, Wong HC, Krishna NR, Wooten MW . Sequestosome 1/p62 is a polyubiquitin chain binding protein involved in ubiquitin proteasome degradation. Mol Cell Biol 2004; 24: 8055–8068.
Gamerdinger M, Hajieva P, Kaya AM, Wolfrum U, Hartl FU, Behl C . Protein quality control during aging involves recruitment of the macroautophagy pathway by BAG3. EMBO J 2009; 28: 889–901.
Tan JM, Wong ES, Kirkpatrick DS, Pletnikova O, Ko HS, Tay SP et al. Lysine 63-linked ubiquitination promotes the formation and autophagic clearance of protein inclusions associated with neurodegenerative diseases. Hum Mol Genet 2008; 17: 431–439.
Paine S, Bedford L, Thorpe J, Mayer R, Cavey J, Bajaj N et al. Lys63-linked polyubiquitin pathology is a feature of neurodegeneration. Neurosci Lett 2009; 460: 205–208.
Jeong H, Then F, Melia Jr TJ, Mazzulli JR, Cui L, Savas JN et al. Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 2009; 137: 60–72.
Kitamura H, Torigoe T, Asanuma H, Hisasue SI, Suzuki K, Tsukamoto T et al. Cytosolic overexpression of p62 sequestosome 1 in neoplastic prostate tissue. Histopathology 2006; 48: 157–161.
Rolland P, Madjd Z, Durrant L, Ellis IO, Layfield R, Spendlove I . The ubiquitin-binding protein p62 is expressed in breast cancers showing features of aggressive disease. Endocr Relat Cancer 2007; 14: 73–80.
Qian HL, Peng XX, Chen SH, Ye HM, Qiu JH . p62 Expression in primary carcinomas of the digestive system. World J Gastroenterol 2005; 11: 1788–1792.
Du Y, Wooten MC, Gearing M, Wooten MW . Age-associated oxidative damage to the p62 promoter: implications for Alzheimer disease. Free Radic Biol Med 2009; 46: 492–501.
Kaganovich D, Kopito R, Frydman J . Misfolded proteins partition between two distinct quality control compartments. Nature 2008; 454: 1088–1095.
Geetha T, Seibenhener ML, Chen L, Madura K, Wooten MW . p62 serves as a shuttling factor for TrkA interaction with the proteasome. Biochem Biophys Res Commun 2008; 374: 33–37.
Geetha T, Jiang J, Wooten MW . Lysine 63 polyubiquitination of the nerve growth factor receptor TrkA directs internalization and signaling. Mol Cell 2005; 20: 301–312.
Saeki Y, Kudo T, Sone T, Kikuchi Y, Yokosawa H, Toh-e A et al. Lysine 63-linked polyubiquitin chain may serve as a targeting signal for the 26S proteasome. EMBO J 2009; 28: 359–371.
Cooper EM, Cutcliffe C, Kristiansen TZ, Pandey A, Pickart CM, Cohen RE . K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1. EMBO J 2009; 28: 621–631.
Filimonenko M, Stuffers S, Raiborg C, Yamamoto A, Malerod L, Fisher EM et al. Functional multivesicular bodies are required for autophagic clearance of protein aggregates associated with neurodegenerative disease. J Cell Biol 2007; 179: 485–500.
Razi M, Chan EY, Tooze SA . Early endosomes and endosomal coatomer are required for autophagy. J Cell Biol 2009.
Kim PK, Hailey DW, Mullen RT, Lippincott-Schwartz J . Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc Natl Acad Sci USA 2008; 105: 20567–20574.
Pohl C, Jentsch S . Midbody ring disposal by autophagy is a post abscission event of cytokinesis. Nat Cell Biol 2009; 11: 65–70.
Yang Q, She H, Gearing M, Colla E, Lee M, Shacka JJ et al. Regulation of neuronal survival factor MEF2D by chaperone-mediated autophagy. Science 2009; 323: 124–127.
Pursiheimo JP, Rantanen K, Heikkinen PT, Johansen T, Jaakkola PM . Hypoxia-activated autophagy accelerates degradation of SQSTM1/p62. Oncogene 2009; 28: 334–344.
Jaakkola PM, Pursiheimo JP . p62 degradation by autophagy: another way for cancer cells to survive under hypoxia. Autophagy 2009; 5: 410–412.
Rodriguez A, Duran A, Selloum M, Champy MF, Diez-Guerra FJ, Flores JM et al. Mature-onset obesity and insulin resistance in mice deficient in the signaling adapter p62. Cell Metab 2006; 3: 211–222.
Jin Z, Li Y, Pitti R, Lawrence D, Pham V, Lill J et al. Cullin3-based polyubiquitination and p62-dependent aggregation of caspase-8 mediate cell-extrinsic apoptosis signaling. Cell 2009; 137: 721–735.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Moscat, J., Diaz-Meco, M. & Wooten, M. Of the atypical PKCs, Par-4 and p62: recent understandings of the biology and pathology of a PB1-dominated complex. Cell Death Differ 16, 1426–1437 (2009). https://doi.org/10.1038/cdd.2009.119
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.119
Keywords
This article is cited by
-
Possible prognostic impact of PKCι genetic variants in prostate cancer
Cancer Cell International (2024)
-
PKCι induces differential phosphorylation of STAT3 to modify STAT3-related signaling pathways in pancreatic cancer cells
Journal of Cell Communication and Signaling (2023)
-
Plasticity of Paneth cells and their ability to regulate intestinal stem cells
Stem Cell Research & Therapy (2020)
-
Addiction to protein kinase Cɩ due to PRKCI gene amplification can be exploited for an aptamer-based targeted therapy in ovarian cancer
Signal Transduction and Targeted Therapy (2020)
-
The signaling axis atypical protein kinase C λ/ι-Satb2 mediates leukemic transformation of B-cell progenitors
Nature Communications (2019)