Abstract
DNA methylation is one of several epigenetic changes observed in cells. Aberrant methylation of tumor suppressor genes, proto-oncogenes, and vital cell cycle genes has led many scientists to investigate the underlying cellular mechanisms of DNA methylation under normal and pathological conditions. Although DNA methylation is necessary for normal mammalian embryogenesis, both hypo- and hypermethylation of DNA are frequently observed in carcinogenesis and other pathological disorders. DNA hypermethylation silences the transcription of many tumor suppressor genes, resulting in immortalization of tumor cells. The reverse process, demethylation and restoration of normal functional expression of genes, is augmented by DNA methylation inhibitors. Recent studies suggest that DNA hypomethylation may also control gene expression and chromosomal stability. However, the roles of and relationship between hypomethylation and hypermethylation are not well understood. This review provides a brief overview of the mechanism of DNA methylation, its relationship to extrinsic stimulation including dietary intake and aging, and of abnormally methylated DNA in breast and colorectal cancers, which could be used as prognostic and diagnostic markers.
Similar content being viewed by others
Main
During translation and transcription, DNA exerts its effects in cells through regulatory mechanisms including mRNA stabilization, transcription, and epigenetic changes.1 The pattern of altered gene expression or epigenetic change is of major importance in common malignancies.2, 3 Methylation of the DNA, histone deacetylation, ubiquitination, and phosphorylation are examples of epigenetic change.4 DNA methylation, unlike the other epigenetic changes, does not alter the nucleotide sequence.
Most cytosine-phosphoguanine (CpG) dinucleotides are unevenly distributed throughout the genome and remain in short stretches or clusters (500–2000 bp), called CpG islands.5, 6, 7 These islands are located in the promoter region and are found in half of all human genes.8 In mammals, DNA methylation occurs after replication, when a methyl group (CH3) is added to the 5′ position of cytidyl residues in the dinucleotide sequence CpG9, 10 (Figure 1). Endonucleases, which normally degrade foreign DNA, regulate gene expression by silencing genes when the CpG is methylated.11 CpG islands remain unmethylated in housekeeping genes and methylated or silenced in other genes.12
DNA is methylated by DNA methyltransferases (DNMTs), which transfer the methyl group from S-adenosylmethionine (SAM) to generate patterns of genomic methylation that silence genes13, 14, 15 (Figure 1). The DNMTs known to date are DNMT1, DNMT2, DNMT3a, DNMT3b, and DNMT3L.15, 16 All methyltransferases have homology and different functions. DNMT1 maintains established methylation patterns in hemi-methylated genes by copying methylation patterns from the parent strand to the daughter and is expressed during the S-phase.17 DNMT2, a small protein of 391 amino acids, is known to have weak DNA methyltransferase activity.18 DNMT3a and DNTM3b, referred to as de novo methyltransferases, methylate unmethylated DNA. They initiate normal DNA methylation during embryonic development.19 DNMT3L does not bind to SAM, but increases the binding of DNMT3a to SAM. Since DNMT3L-deficient mice are sterile, DNMT3L is likely to be essential in the methylation process.19
Overexpression of DNMTs can be lethal in animals as well as in human cancers.20 DNMTs are involved in the downregulation of tumor suppressor genes and stimulation of proto-oncogenes.21, 22 Although DNMTs are important in DNA methylation, several findings indicate that DNMTs are not essential for the promotion of carcinogenesis.23, 24, 25
Hypo- and hypermethylation
When gene expression is altered due to DNA methylation, it is usually categorized as due to hypo-methylation or hypermethylation (Figure 2). DNA hypomethylation is associated with gene reactivation and chromosomal instabilities.26, 27 Functional outcomes of hypomethylation include the upregulation or overexpression of transcription of proto-oncogenes, increased recombination and mutation, X-chromosome inactivation, loss of imprinting, reactivation of transposable elements, and demethylation of xenobiotics10, 28, 29 (Figure 3). Activation of proto-oncogenes, reactivation of transposable elements, and loss of imprinting of genes are the results of hypomethylation and all promote cancer.30
When CpG islands are hypermethylated, the activity of the regulatory proteins that promote transcription is restricted due to the tightly packed nucleosomes.8 DNA hypermethylation is involved in gene repression and chromosomal instabilities.31 Results of hypermethylation include suppression of tumor suppressor genes, chromatin condensation, and suppression of DNA repair genes (Figure 4). Tumor suppressor genes contain unmethylated CpG islands in their promoter regions and are methylated in various malignancies.32, 33, 34 Both types of methylation occur simultaneously in various sporadic cancers and affect the function of human tumor suppressor genes and proto-oncogenes.35
Detection of DNA methylation
Currently, several methods are used to detect methylated DNA (Table 1).
Methylated DNA can be detected by converting unmethylated cytosine residues to uracil using sodium bisulfite modification, followed by polymerase chain reaction (PCR) to identify the unmethylated nucleotides.36 Methylation-specific PCR (MSP) detects the methylation of CpG islands, but with higher specificity and sensitivity.37 Quantitative real-time PCR (MethyLight or QM-MSP) is used to detect low levels of methylation that cannot be distinguished using MSP.38, 39
A modification of MSP, termed McrBC-methylation-sensitive-arbitrarily-primed-polymerase-chain-reaction (McrBC-msAP-PCR) requires methylation-specific GTP-dependent restriction endonuclease, McrBC, to detect differentially methylated sites within DNA, where hypermethylation and hypomethylation is observed by DNA fingerprint band intensity.40 The procedure for the methylated CpG island recovery assay (MIRA) requires isolated and sonicated genomic DNA, instead of sodium bisulfite.41 PCR reactions detect CpG island methylation after the DNA is incubated with a matrix containing methyl-CpG binding domain protein-2b (MBD2b) and methyl-CpG binding domain protein 3-like-1, which bind specifically to methylated DNA sequences.41, 42 MIRA can be used to study the methylation status of a wide array of genes in cancers, such as the lung.42 Currently, other array-based methods are being developed to screen the methylation pattern of several genes.39
Differential methylation hybridization (DMH) is an oligonucleotide array-based method, which determines the extent of methylation of CpG islands by comparison with a reference sample.44 Methylation-specific restriction enzymes (MseI) are used to obtain intact CpG islands. The CpG islands are fluorescently labeled and after subsequent PCR, they are hybridized to arrayed oligonucleotides that can discriminate between methylated and unmethylated alleles in regions of interest. Microarray methylation assessment of single samples (MMASS) has been shown to be more sensitive and is an optimized method for detecting the methylated and unmethylated sequences within the entire genome.45 MMASS uses the methylation-specific enzyme, McrBC, instead of MseI. It is able to detect unmethylated sequences more effectively than DMH because McrBC only cleaves methylated sequences and does not require a reference sample.45 Methylation target array (MTA) simultaneously determines whether or not genes and CpG islands in multiple tumors are hypermethylated and can be correlated to clinicopathological features of the patient.46 An advantage of MTA is that a single nylon filter can be used repeatedly to probe for various genes which are indicative of DNA methylation.47
DNA methylation and cancer: overview
Aberrant DNA methylation is one of the many potential causes for the abnormal growth of cancer cells, but it is also known to protect against intestinal cancer.4 Different types of cancers are associated with methylation of tumor suppressor genes and proto-oncogenes, causing alterations in functional gene expression. Cancer-specific DNA methylation patterns have been detected in free-floating DNA released from dead cancer cells.5, 48 A decrease in the expression of tumor suppressor genes correlates with an increase in methylation of DNA in the promoter region.49, 50 Aberrant methylation of tumor suppressor genes in many cancers, resulting in the downregulation of transcriptional activation, has been reported.
In some cancers, both hypermethylation and hypomethylation are observed. Hypomethylation increases progressively with increasing malignancy grade in breast, ovarian, cervical, and brain cancers.51 Breast and colorectal cancers are malignancies commonly caused by regional hypermutability or global hypomethylation.
DNA methylation and breast cancer
Many factors contribute to the pathogenesis of breast cancer, which is one of the most common malignancies among females. These factors include family history, nutrition, age, and epigenetic changes including DNA methylation. Methylation appears to be an early event in the etiology of breast carcinogenesis, resulting in the activation of many oncogenes and silencing of tumor suppressors to promote proliferation of abnormal cells.52, 53 It is debatable, however, whether global hypomethylation or regional hypermethylation occurs first during the development of breast cancer, since the phenomena are independent processes. It is not known whether or not it is possible to inhibit carcinogenesis by inhibiting one of these processes and not the others. Numerous studies have revealed various genes, which are either hypo- or hypermethylated in breast cancer (Table 2).
Metastatic breast cancer requires the expression of multiple genes. Regional hypermethylation and global hypomethylation are involved in different stages of breast cancer.76 Global hypomethylation could be a mechanism for late stages of breast cancer while local hypermethylation is plausible for early stages of breast cancer.22, 30 DNA methylation results in altered gene products including cell cycle regulators, steroid receptors, and cell adhesion molecules, which give rise to increased susceptibility to tumor development and decreased detoxification of carcinogens.58 Alterations in the breast cancer susceptibility gene product (BRCA) accounts for half of the inherited breast carcinomas.77 Its methylation is observed in breast and ovarian cancers, but not in colon and liver cancers, or leukemia indicating a tissue-specific process.58 The frequency of methylation of this gene product is 38.5% in sporadic breast cancer.60 Patients with a HER2/Neu-positive tumor indicate a highly aggressive breast cancer that requires special treatment, since it is amplified in 30% of invasive breast carcinomas.55 DNA methylation is prevalent in the highly aggressive HER2/Neu-positive breast cancers; this gene is amplified in 30% of the cancers.55 Increased aberrant methylation of steroid receptor genes and glycoproteins, such as progesterone receptor (PR) and E-cadherin, respectively, are associated with Her2/Neu-positive cancers. Hypermethylation of the GC-rich region and loss of expression in about 80% of invasive lobular carcinomas and lobular carcinoma in situ indicate the importance of the methylation of the CDH1 promoter in the pathogenesis of breast cancer.78 Although many mechanisms, including mutation and loss of heterozygosity (LOH), are attributed to the downregulation of CDH1 in breast cancer, CDH1 promoter methylation is the mostly likely cause.78, 79
The current criteria for detection and prognosis of breast cancer include an abnormal breast biopsy, tumor size, histological grade, estrogen and progesterone receptor status, and presence of the HER2/Neu oncogene.80, 81 Breast cancer can also be diagnosed by detecting the various aberrantly methylated genes. MSP is currently being used to detect the methylation status of various genes in breast biopsy tissues samples.59 Presence of methylated DNA in the nipple duct lavage fluids, needle aspirates of the breast, and molecular staging of sentinel lymph nodes are also used to predict breast cancer development.58 Ductal carcinoma in situ (DCIS), the most frequent breast cancer, can be detected early by observing the methylation of a panel of tumor suppressor or other cancer genes.82, 83 A 60-sample study with ductal lavage fluid from patients with a high risk of developing breast cancer and patients with breast cancer revealed that a nine-gene panel detection system using quantitative methylation-specific polymerase chain reaction (QM-MSP) can detect the rate of cancer cells more effectively than cytological and histological studies alone. Thus, earlier detection of breast cancer formation is possible.84 Aberrant methylation of four genes was detected by QM-MSP in the plasma DNA of patients with breast cancer and tumors were successfully detected in eight of 24 patients with early-stage breast cancer.85 Loss of MGMT (O(6)-methylguanine-DNA methyltransferase) was found to be associated with DNA methylation in a subset of breast cancers.86 Although DNMT3b, a de novo methyl transferase, is overexpressed in 30% of breast cancers, its expression alone was not considered to be a prognostic factor for breast cancer progression.87 Such studies highlight the importance of identifying specific methylated genes for diagnostic purposes, and for monitoring the efficacy of therapeutic modalities.
DNA methylation and colorectal cancer
Colorectal carcinoma is the third most common cancer in developed countries.88 Although age and other demographic and environmental features, including gender, weight, nutritional intake, and alcohol consumption, are prognostic of colorectal cancer, epigenetic alterations are also causal.89 Aberrant methylation is gradually acquired in the early stages of colorectal carcinoma.90 As in breast cancer, both hypomethylation and hypermethylation of genes occur in colorectal cancer91, 92 (Table 3). The genes for p53 and for retinoic acid receptor (RAR) are hypermethylated in colorectal cancer.108 A study with 65 colorectal carcinoma tissues demonstrated hypermethylation of the gene for the cell cycle regulatory protein, cyclin A1, in all cases, and for cadherin-13 in 65% cases.96 Methylation did not correlate significantly with any clinicopathological feature, and changes in methylation appeared in an early phase of colon carcinogenesis.96
Aberrant methylation of some genes, including those for estrogen receptor α (ERα) and myoblast determination 1-protein (MYOD), correlate with aging and the prognosis of colorectal cancer.109 UDP-glucoronosyltransferase (UGT1A1) gene expression is silenced and transcriptional activity is completely repressed in colon cancer cells due to direct methylation of its promoter region.97 Treatment with either the inhibitors of histone deacetylase or demethylating agents restores normal expression of UGT1A1 in hypermethylated cells but has no effect on hypomethylated cells.97
Rhee et al100 reported that DNA methyltransferases, DNMT1 and DNMT3b, are hypermethylated in colorectal cancer. Other studies suggest that methylation depends upon the type of cancer and that colorectal cancer can progress in the absence of DNMT1, as seen in the SW48 colorectal cancer cell line.108, 110 However, methylation can be reinitiated by introducing DNMT1 to the SW48 colon cancer cells lacking DNMT1 and DNMT3b.111 Therefore, DNMT1 is potentially important in the hypermethylation of CpG islands in the promoter region of many genes in human cancer cells.
Colorectal tumors are often identified by the level of their microsatellite instability (MSI), which is a defect in the ability of repairing mistakes during DNA replication. MSI, stratified as MSI high, MSI low, and MSI stable, is commonly correlated with the degree of the colorectal cancer methylator phenotype (CIMP) when diagnosing colorectal cancer at the molecular level.112 CIMP is distinguished as CIMP(+) or CIMP(−), although the existence of CIMP is still controversial. Supporters suggest an association with microsatellite instability (MSI) and proximal location of the colonic tumor.113 A study with 106 primary colorectal tumors, however, did not support the existence of CIMP in human colorectal cancer and, consequently, it was regarded as a statistical artifact.114 Colorectal cancer cannot be identified on the basis of tumor methylation status and CIMP alone.
Hereditary non-polyposis colorectal cancer (HNPCC, Lynch syndrome) accounts for 2–4% of all colorectal cancers and aberrant methylation of the mismatch repair genes, human mutL homolog 1 (hMLH1) or hMLH2, are the basis for the cancer.115 The combination of MSI-H and CIMP(−) is commonly observed in HNPCC.112 In a study with 97 colorectal adenoma cases, hMLH1 methylation was more frequently observed in overweight or obese patients.93 High-level MSI sporadic colon cancer and HNPCC share histological features, proximal tumor location, and presence of tumor-infiltrating lymphocytes. They differ, however, in having widespread promoter hypermethylation of specific genes such as hMLH1 and BRAF.112 Fewer methylated genes are found in HNPCC than in high-level MSI colorectal tumors.116
Inhibitors of DNA methylation and demethylators
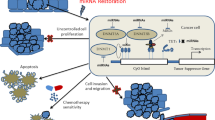
DNA methylation is a reversible process in which genes can be demethylated and restored to their original expression and function. DNA methylation inhibitors have been investigated as anticancer agents, since they block the activity of DNMTs and thus activate tumor suppressor genes54 (Figure 5). The antisense oligonucleotide to human DNMT1, MG98, acts as a DNA methylation inhibitor and downregulates the activity of DNMT1. It has had promising results in clinical trials in treating cancers of the head and neck.117 Recently, two novel inhibitors, NSC303530 and NSC401077, were shown to inhibit DNMTs in vitro and in vivo by blocking the active site of DNMT1.118 These inhibitors have also been proposed as potential antitumor drugs.118
Demethylation of aberrantly silenced genes can restore gene expression and function. Azacitidine (5-aza-C) and decitabine (5-aza-2′-deoxycytidine or 5-aza-2′-CdR) are two potent DNA demethylating agents. 5-aza-C was developed as a cancer chemotherapeutic agent and was thought to inhibit the enzymes that methylate the cytosine residues in DNA.10, 119 However, it is now considered that it demethylates methylated DNA and acts as an anti-tumor agent in many cancer cells.120 Azacitidine is more toxic than its analog, decitabine.22
Decitabine has a short in vivo half-life and is able to reactivate previously silenced genes.121, 122 In clinical investigations in treating leukemia, decitabine was found to be most effective in an intensive dose with a short treatment time.123 Decitabine is a potent cytotoxic agent and shows in vitro antitumor activity against breast cancer cells.124 Both azacitidine and decitabine are effective in treating leukemia, which is characterized by hypermethylation.35, 125
Zebularine, another potent demethylator, was found to be effective against cancer cells in many studies. In earlier studies, it was found to have toxic effects in cancer cell lines.73 It has been shown to remove 25–60% of the methyl groups from methylated genes in a panel of seven human tumor cell lines.94 The most promising features of the drug include its stability, low toxicity to normal cells, and that it can be taken orally.126
Administration of decitabine can reactivate silenced tumor suppressor genes and the histone deacetylase inhibitor, LAQ824 (LAQ), can activate genes related to cell cycle arrest.123, 127 These agents can synergistically produce greater antineoplastic effects on MDA-MB-231 breast cancer cells, thereby ensuring application of these agents for future clinical trials.127 Lower doses of azacitidine and trichostatin A (TSA) are required to re-express ER in MDA-MB-231 (ER−) cells, when the drugs are used in combination than alone.58 Thus, the combination of drugs, which directly affects DNA methylation with drugs, causing other epigenetic changes, has considerable potential in increasing therapeutic affects.
Cancer Prevention, Dietary Intake, and DNA Methylation
Nutrition influences susceptibility to cancer. Approximately 35–50% of all cancers have a dietary component in their etiology.128 Some food constituents can promote the onset of cancer. Deficiency of fiber, folic acid, methionine/choline, zinc, selenium, and chemicals found only in fruits and vegetables all can also cause cancer.129 Excess intake of alcohol, animal fats, and salt promote cancer.130 It was predicted that the incidence of cancer in vegetarian subjects would be lower than in those on a meat diet. Even though the vegetarian diet has a lower intake of vitamin B and lower content of methionine, both of which are essential for eventual methylation, this does not cause cancer.131
The role of folic acid in cancer is still controversial. The methyl groups of 5-methylenetetrahydrofolate is the precursor of the methyl group of methionine and thereby of SAM.132 Low intake of folate combined with high alcohol intake can result in global hypomethylation and cause colorectal cancer. An increased risk of breast cancer occurs if folic acid is not metabolized correctly and the resultant supply of methyl groups to DNA in pre-menopausal women is insufficient.113, 132
Low intake of folate is associated with an increased risk of colorectal cancer.113 In vitro studies have indicated that colon cancer cell lines, when deprived of folic acid, have decreased viability.132 Uracil is misincorporated into DNA as a result of folate deficiency. However, once a cancerous lesion is present, folate intake enhances tumor growth.133 Methionine is taken as a nutritional supplement in adulthood to correct some genetically based epigenetic defects.29 However, excess intake of methionine can also impair DNA methylation. Nevertheless, folate deficiency, leading to aberrant methylation of DNA, is not the sole cause of colon carcinogenesis.
Caffeic acid and chlorogenic acid, two catechol containing coffee polyphenols, inhibit DNA methylation.134 They increase the formation of S-adenosyl-L-homocysteine (SAH), an inhibitor of DNA methylation. Partial inhibition of methylation in the promoter region of the retinoic acid receptor beta (RAR-β) gene by both caffeic acid and chlorgenic acid was demonstrated in breast cancer cell lines MCF-7 and MDA-MB-231.135
DNA Methylation and Aging
The risk of cancer increases with age. Only 10% of children have a chance of getting cancer, whereas adults have a 35% chance, because methylation of CpG islands in non-malignant tissues increases but the total number of methylated cysteine residues decreases with age.81, 136, 137 Individual genes are progressively methylated during aging due to chromosomal instability.138 Genes that change methylation status with age are tissue specific. The c-myc gene is hypomethylated in the spleen and the c-fox is hypermethylated in the liver but not in the spleen.28
The normal colonic mucosa of older females has higher methylation levels, making these cells more susceptible to differentiate into malignant cells.103, 139 Hypermethylation does not always result in malignancy. For example, hypermethylation of the estrogen receptor gene was observed in both normal and cancer colon tissues, suggesting that the relationship between hypermethylation and age in cancer might not be simple, and requires a more careful analysis.8
Concluding remarks
DNA methylation is important in gene regulation and expression. It is imperative to learn more about the regulation of how this simple and basic process becomes aberrant. The complete biological mechanisms that initiate and maintain methylation of DNA need to be fully explained. Both hypomethylation and hypermethylation of proto-oncogenes and/or tumor suppressor genes occurs in various cancers. Many known genes are aberrantly methylated in breast and colorectal cancers.
In studies with established inhibitors of DNA methylation and demethylation, some genes were shown to be able to resume normal function. Zebularine was shown to be effective, without many toxic affects, in clinical trials. Despite constant efforts, the most effective and least toxic drugs are yet to be discovered. Indeed, even more questions arise from the plethora of recent advances in our understanding of the underlying mechanisms of methylation in cancer and other malignancies.
References
Nephew KP, Huang TH . Epigenetic gene silencing in cancer and progression. Cancer Lett 2003;190:125–133.
Baylin SB, Ohm JE . Epigenetic gene silencing in cancer—a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 2006;6:107–116.
Perez-Plasencia C, Duenas-Gonzalez A . Can the state of cancer chemotherapy resistance be reverted by epigenetic therapy? Mol Cancer 2006;5:27.
Kim YI . Nutritional epigenetics: impact of folate deficiency on DNA methylation and colon cancer susceptibility. J Nutr 2005;135:2703–2709.
Laird PW, Jaenisch R . The role of DNA methylation in cancer genetic and epigenetic. Annu Rev Genet 1996;30:441–464.
Feltus FA, Lee EK, Costello JF, et al. DNA motifs associated with aberrant CpG island methylation. Genomics 2006;5:572–579.
Vertino PM, Yen RW, Gao J, et al. De novo methylation of CpG island sequences in human fibroblast overexpressing DNA cytosine-5-methyltransferase. Mol Cell Biol 1996;16:4555–4565.
Baylin SB . DNA methylation and gene silencing in cancer. Nat Clin Pract 2005;2:S4–S11.
Bird A . DNA methylation patterns epigenetic memory. Genes Dev 2002;16:6–21.
Dunn BK . Hypomethylation: one side of a larger picture. Ann NY Acad Sci 2003;983:28–42.
Farrell WE . Epigenetic mechanisms of tumorigenesis. Horm Metab Res 2005;37:361–368.
Antequera F, Bird A . Number of CpG islands and genes in human and mouse. Proc Natl Acad Sci USA 1993;90:11995–11999.
Ross SA . Diet and DNA methylation interactions in cancer prevention. Ann NY Acad Sci 2003;983:197–207.
Brenner C, Fuks F . DNA methyltransferases: facts, clue, mysteries. Curr Top Microbiol Immunol 2006;301:45–66.
Gopisetty G, Ramachandran K, Singal R . DNA methylation and apoptosis. Mol Immunol 2006;43:1729–1740.
Singal R, Ginder GD . DNA methylation. Blood 1999;93:4059–4070.
Agoston AT, Argani P, Yegnasubramanian S, et al. Increased protein stability causes DNA methyltransferase 1 dysregulation in breast cancer. J Biol Chem 2005;280:18302–18310.
Chen T, Li E . Establishment and maintenance of DNA methylation patterns in mammals. Curr Top Microbiol Immunol 2006;301:179–201.
Jaenisch R . DNA methylation and imprinting: why bother? Trends Genet 1997;13:323–329.
Rodenhiser D, Mann M . Epigenetics and human disease: translating basic biology into clinical applications. CMAJ 2006;174:341–348.
Robertson KD, Ait-Si-Ali S, Yokochil T, et al. DNMT1 forms a complex with Rb, E2F1, and HDAC1 and represses transcription from E2F-responsive promoters. Nat Genet 2000;25:338–342.
Szyf M . The DNA methylation machinery as a therapeutic target. Curr Drug Targets 2000;1:101–118.
Eads CA, Danenberg KD, Kawakami K, et al. CpG island hypermethylation in human colorectal tumors is not associated with DNA methyltransferase overexpression. Cancer Res 1999;59:2302–2306.
Rhee I, Jair KW, Yen RW, et al. CpG methylation is maintained in human cancer cells lacking DNMT1. Nature 2000;404:1003–1007.
Li E, Beard C, Forster AC, et al. DNA methylation, genomic imprinting, and mammalian development. Cold Spring Harb Symp Quant Biol 1993;58:297–305.
De Smet C, Loriot A, Boon T . Promoter-dependent mechanism leading to selective hypomethylation within the 5′ region of gene MAGE-A1 in tumor cells. Mol Cell Biol 2004;24:4781–4790.
Eden A, Gaudet F, Waghmare A, et al. Chromosomal instability and tumors promoted by DNA hypomethylation. Science 2003;300:455.
Richardson BC . Role of DNA methylation in the regulation of cell function: autoimmunity, aging and cancer. J Nutr 2002;132:2401S–2405S.
Waterland RA . Assessing the effects of high methionine intake on DNA methylation. J Nutr 2006;136:1706S–1710S.
Esteller M, Herman JG . Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumors. J Pathol 2002;196:1–7.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002;3:415–428.
Baylin SB, Herman JG, Graff JR . Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res 1998;72:141–196.
Jones PA, Laird PW . Cancer epigenetics comes of age. Nat Genet 1999;21:163–167.
Esteller M, Corn PG, Baylin SB, et al. A gene hypermethylation profile of human cancer. Cancer Res 2001;61:3225–3229.
Balch C, Montgomery JS, Paik HI . New anti-cancer strategies: epigenetic therapies and biomarkers. Front Biosci 2005;10:1897–1931.
Herman JG, Graff JR, Myohanen S, et al. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 1996;93:9821–9826.
Sadri R, Hornsby PJ . Rapid analysis of DNA methylation using new restriction enzyme sites created by bisulfite modification. Nucleic Acids Res 1996;24:5058–5059.
Ogino S, Brahmandam M, Kawasaki T, et al. Combined analysis of COX-2and p53 expressions reveals synergistic inverse correlations with microsatellite instability and CpG island methylator phenotype in colorectal cancer. Neoplasia 2006;8:458–464.
Wojdacz TK, Hansen LL . Techniques used in studies of age-related DNA methylation changes. Ann NY Acad Sci 2006;1067:479–487.
Tryndyak V, Kovalchuk O, Pogribyny IP . Identification of differentially methylated sites within unmethylated DNA domains in normal and cancer cells. Anal Biochem 2006;356:202–207.
Rauch T, Pfeifer GP . Methylated-CpG island recovery assay: a new technique for the rapid detection of methylated-CpG islands in cancer. Lab Invest 2005;85:1172–1180.
Rauch T, Li H, Wu X, et al. MIRA-assisted microarray analysis, a new technology for the determination of DNA methylation patterns, identifies frequent methylation of homeodomain-containing genes in lung cancer cells. Cancer Res 2006;66:7939–7947.
Xiong Z, Laird PW . COBRA: a sensitive and quantitative DNA methylation assay. Nucleic Acids Res 1997;25:2532–2534.
Ho SM, Tang WY . Techniques used in studies of epigenetic dysregulation due to aberrant DNA methylation: an emphasis of fetal-based adult diseases. Reprod Toxicol 2007, Jan 19 [E-pub ahead of print].
Ibrahim AE, Thorne NP, Baird K, et al. MMASS: an optimized array-based method for assessing CpG island methylation. Nucleic Acids Res 2006;34:e136.
Chen C-M, Chen H-L, Hsiau TH-C, et al. Methylation target array for rapid analysis of CpG island hypermethylation in multiple tissue genomes. Am J Pathol 2003;163:37–45.
Ballestar E, Paz MF, Valle L, et al. Methyl-CpG binding proteins identify novel sites of epigenetic inactivation in human cancer. EMBO J 2003;22:6335–6345.
Bastian PJ, Yegnasubramanian S, Palapattu GS, et al. Molecular biomarker in prostate cancer: the role of Cpg island hypermethylation. Euro Urol 2004;46:698–708.
Schmutte C, Jones PA . Involvement of DNA methylation in human carcinogenesis. Biol Chem 1998;379:377–388.
Salozhin SV, Prokhorchuk EB, Geogiev GP . Methylation of DNA—one of the major epigenetic markers. Biochem 2005;70:525–532.
Costello JF, Plass C . Methylation matters. J Med Genet 2001;38:285–303.
Jackson K, Yu MC, Arakwa K, et al. DNA hypomethylation is prevalent even in low-grade breast cancers. Cancer Biol Ther 2004;12:1225–1231.
Chekhun VF, Kulik GI, Yurchenko OV, et al. Role of DNA hypomethylation in the development of the resistance to doxorubicin in human MCF-7 breast adenocarcinoma cells. Cancer Lett 2006;231:87–93.
Szyf M, Pakneshan P, Rabbani SA . DNA methylation and breast cancer. Biochem Pharmacol 2004;68:1187–1197.
Ross JS, Linette GP, Stec J, et al. Breast cancer biomarkers and molecular medicine. Expert Rev Mol Diagn 2003;3:573–585.
Wischnewski F, Pantel K, Schwarzenback H . Promoter demethylation and histone acetylation mediate gene expression of MAGE-A1, -A2, -A3, and -A12 in human cancer cells. Mol Cancer Res 2006;4:339–349.
Narayan A, Ji W, Zhang X-Y, et al. Hypomethylation of pericentromeric DNA in breast adenocarcinomas. Int J Cancer 1998;77:833–838.
Yang X, Yan L, Davidson NE . DNA methylation in breast cancer. Endocr Relat Cancer 2001;8:115–127.
Miyamoto K, Fukutomi T, Akashi-Tanaka S, et al. Identification of 20 genes aberrantly methylated in human breast cancers. Int J Cancer 2005;116:407–414.
Birgisdottir V, Stefansson OA, Bodvarsdottir SK, et al. Epigenetic silencing and deletion of the BRCA1 gene in sporadic breast cancer. Breast Cancer Res 2006;8:R38.
Caldeira JRF, Prando EC, Quevedo FC, et al. CDH1 promotor hypermethylation and E-cadherin protein expression in infiltrating breast cancer. BMC Cancer 2006;6:48.
Oshiro MM, Kim LJ, Wozhiak RJ, et al. Epigenetics silencing of DSC3 is a common event in human breast cancer. Breast Cancer Res 2005;7:R669–R680.
Fackler MJ, McVeigh M, Evron E, et al. DNA methylation of RASSF1A, HIN-1, RAR-beta, cyclin D2 and twist in in situ and invasive lobular breast carcinoma. Int J Cancer 2003;107:970–975.
Prasad GL, Valverius EM, McDuffie E, et al. Complementary DNA cloning of a novel epithelial cell marker protein, HME1, that may be downregulated in neoplastic mammary cells. Cell Growth Differ 1992;3:507–513.
Pampalakis G, Sotiropoulou G . Multiple mechanisms underlie the aberrant expression of the human kallikrein 6 gene in breast cancer. Biol Chem 2006;387:773–782.
Sidiropoulos M, Pampalakis G, Sotiropoulou G, et al. Downregulation of human kallikrein 10 (KLK10/NES1) by CpG island hypermethylation in breast, ovarian, and prostate cancers. Tumor Biol 2005;26:324–336.
Takahashi Y, Miyoshi Y, Takahata C, et al. Down-regulation of LATS1 and LATS2 mRNA expression by promoter hypermethylation and its association with biologically aggressive phenotype in human breast cancers. Clin Cancer Res 2005;11:1380–1385.
Fenton H, Carlile B, Montgomery EA, et al. LKB1 protein expression in human breast cancer. Appl Immunohistochem Mol Morphol 2006;14:146–153.
Esteller M . Epigenetic lesions causing genetic lesions in human cancer: promoter hypermethylation of DNA repair genes. Eur J Cancer 2000;36:2294–2300.
Hesson L, Dallol A, Minna JD, et al. NORE1A, a homologue of RASSF1A tumor suppressor gene is inactivated in human cancer. Oncogene 2003;22:947–954.
Silva J, Silva JM, Dominguez G, et al. Concomitant expression of p16INK4a and p14ARF in primary breast cancer and analysis of inactivation mechanisms. J Path 2003;199:289–297.
Kobatake T, Yano M, Toyooka S, et al. Aberrant methylation of p57KIP2 gene in lung and breast cancers and malignant mesotheliomas. Oncol Rep 2004;12:1087–1092.
Cheng CK, Chow LWC, Loo WTY, et al. The cell cycle checkpoint gene Rad9 is a novel oncogene activated by 11q13 amplification and DNA methylation in breast cancer. Cancer Res 2005;65:8646–8654.
Lau QC, Raja E, Salto-Tellez M, et al. RUNX3 is frequently inactivated by dual mechanisms of protein mislocalization and promoter hypermethylation in breast cancer. Cancer Res 2006;66:6512–6520.
Levine JJ, Stimson-Crider KM, Vertino PM . Effects of methylation on expression of TMS/ASC in human breast cancer cells. Oncogene 2003;22:3475–3488.
Van Zhee KJ, Calvano JE, Bisogna M . Hypomethylation and increased gene expression of p16INK4a in primary and metastatic breast carcinoma as compared to normal breast tissue. Oncogene 1998;16:2723–2727.
Elstrodt F, Hollestelle A, Nagel JHA, et al. BRCA1 mutation analysis of 41 human breast cancer cell lines reveals three new deleterious mutants. Cancer Res 2006;66:41–45.
Droufakou S, Deshmane V, Roylance R, et al. Multiple ways of silencing E-cadherin gene expression in lobular carcinoma of the breast. Int J Cancer 2001;92:404–408.
Saario D, Moreno-Bueno G, Hardisson D, et al. Epigenetic and genetic alterations of APC and CDH1 genes in lobular breast cancer: relationships with abnormal E-cadherin and catenin expression and microsatellite instability. Int J Cancer 2003;106:208–215.
Lostumbo A, Mehta D, Setty S, et al. Flow cytometry: a new approach for the molecular profiling of breast cancer. Exp Mol Pathol 2006;80:46–53.
Laird PW . The power and the promise of DNA methylation markers. Nature Rev Cancer 2003;3:253–266.
Evron E, Dooley WC, Umbricht CB, et al. Detection of breast cancer cells in ductal lavage fluid by methlyation-specific PCR. Lancet 2001;357:1335–1336.
Dua RS, Isacke CM, Gui GPH . The intraductal approach to breast cancer biomarker discovery. J Clin Oncol 2006;24:1209–1216.
Fackler MJ, Malone K, Zhang Z, et al. Quantitative multiplex methylation-specific PCR analysis doubles detection of tumor cells in breast ductal fluid. Clin Cancer Res 2006;12:3306–3310.
Hoque MO, Febg O, Toure P, et al. Detection of aberrant methylation of four genes in plasma DNA for the detection of breast cancer. J Clin Oncol 24;2006:4262–4269.
Munot K, Bell SM, Lane S, et al. Pattern of expression of genes linked to epigenetic silencing in human breast cancer. Hum Pathol 2006;37:989–999.
Girault I, Tozlu S, Lidereau R, et al. Expression analysis of DNA methyltransferases 1, 3A, and 3B in sporadic breast carcinomas. Clin Cancer Res 2003;9:4415–4422.
Jubb AM, Quirke P, Oates AJ . DNA methylation, a biomarker for colorectal cancer: implications for screening and pathological utility. Ann NY Acad Sci 2003;983:251–267.
Grady WM . Epigenetic events in the colorectum and in colon cancer. Biochem Soc Trans 2005;33:684–688.
Shames DS, Minna JD, Gazdar AF . DNA methylation in health, disease, and cancer. Curr Mol Med 2007;7:85–102.
Bariol C, Suter C, Cheong K, et al. The relationship between hypomethylation and CpG island methylation in colorectal neoplasia. Am J Pathol 2003;162:1361–1371.
Kennedy EP, Hamilton SR . Genetics of colorectal cancer. Semin Surg Oncol 1998;15:126–130.
Ye C, Shrubsole MJ, Cai Q, et al. Promoter hypermethylation status of the MGMT, hMLH1, and CDKN2A/p16 genes in non-neoplastic mucosa of patients with and without colorectal adenomas. Oncol Rep 2006;16:429–435.
Esteller M . CpG island methylation and histone modifications: biology and clinical significance. Ernst Schering Res Found Workshop 2006a;57:115–126.
Wynter CV, Kambara T, Walsh MD, et al. DNA methylation patterns in adenomas from FAP, multiple adenoma and sporadic colorectal carcinoma patients. Int J Cancer 2006;118:907–915.
Xu XL, Yu J, Zhang HY, et al. Methylation profile of the promoter CpG islands of 31 genes that may contribute to colorectal carcinogenesis. World J Gastroenterol 2004;10:341–354.
Gagnon JF, Bernard O, Villeneuve L, et al. Irinotecan inactivation is modulated by epigenetic silencing of UGT1A1 in colon cancer. Clin Cancer Res 2006;12:1850–1858.
Shu J, Jelinek J, Chang H, et al. Silencing of bidrectional promoters by DNA methylation in tumorigenesis. Cancer Res 2006;66:5077–5084.
Ogino S, Kawasaki T, Kirkner GJ, et al. Down-regulation of p21 (CDKNIA/CIPI) is inversely associated with microsatellite instability and CpG island methylator phenotype (CIMP) in colorectal cancer. J Path 2006;210:147–154.
Rhee I, Bachman KE, Park BH, et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 2002;416:552–556.
Akiyama Y, Watkins N, Suzuki H, et al. GATA-4 and GATA-5 transcription factor genes and potential downstream antitumor target genes are epigenetically silenced in colorectal and gastric cancer. Mol Cell Biochem 2003;23:8429–8439.
Liu L, Zhang J, Bates S, et al. A methylation profile of in vitro immortalized human cell lines. Int J Oncol 2005;26:275–285.
Kawakami K, Ruszkiewicz A, Bennett G, et al. DNA hypermethylation in the normal colonic mucosa of patients with colorectal cancer. Br J Cancer 2006;94:593–598.
Bachman KE, Park BH, Rajagopalan H, et al. Histone modifications and silencing prior to DNA Methylation of a tumor suppressor gene. Cancer Cell 2003;3:89–95.
Goebel B, Muller H, Reimer T, et al. Gene methylation data—a new challenge for bioinformaticians? Methods Inf Med 2005;44:516–519.
Dickinson RE, Dallol A, Bieche I, et al. Epigenetic inactivation of SLIT3 and SLIT1 genes in human cancers. Br J Cancer 2004;91:2071–2078.
Okochi-Takada E, Nakazawa K, Wakabayashi M, et al. Silencing of the UCHL1 gene in human colorectal and ovarian cancers. Int J Cancer 2006;119:1338–1344.
Vousden KH . p53: death star. Cell 2000;103:691–694.
Ahuja N, Li Q, Mohan AL, et al. Aging and DNA methylation in colorectal mucosa and cancer. Cancer Res 1998;58:5489–5494.
Ting AH, Jair K, Schuebel KE, et al. Differential requirement for DNA methyltransferase 1 in maintaining human cancer cell gene promoter hypermethylation. Cancer Res 2006;66:729–735.
Jair KW, Bachman KE, Suzuki H, et al. De novo CpG island methylation in human cancer cells. Cancer Res 2006;66:682–692.
Jass JR, Walsh MD, Barker M, et al. Distinction between familial and sporadic forms of colorectal cancer showing DNA microsatellite instability. Eur J Cancer 2002;38:858–866.
Ulrich CM . Nutrigenetics in cancer research—folate metabolism and colorectal cancer. J Nutr 2005;135:2698–2701.
Anacleto C, Leopoldino AM, Rossi B, et al. Colorectal cancer ‘methylator phenotype’: fact or artifact? Neoplasia 2005;7:331–335.
Gazzoli I, Loda M, Garber J, et al. A hereditary nonpolyposis colorectal carcinoma case associated with hypermethylation of the hMLH1 gene in normal tissue and loss of heterozygosity of the unmethylated allele in the resulting microsatellite instability-high tumor. Cancer Res 2002;62:3925–3928.
McGivern A, Wynter CVA, Whitehall VLJ, et al. Promoter hypermethylation frequency and BRAF mutations distinguish hereditary non-polyposis colon cancer from sporadic MSI-H colon cancer. Fam Cancer 2004;3:101–107.
Davis AJ, Gelmon KA, Siu LL . Phase I and pharmacologic study of the human DNA methyltransferase antisense oligodeoxynucleotide MG98 given as a 21-day continueous infusion every 4 weeks. Invest New Drugs 2003;21:85–97.
Siedlecki P, Boy RG, Musch T, et al. Discovery of two novel, small-molecule inhibitors of DNA methylation. J Med Chem 2006;49:678–683.
Jones PA, Taylor SM . Cellular differentiation, cytidine analogs and DNA methylation. Cell 1980;20:85–93.
Stresemann C, Brueckner B, Musch T, et al. Functional diversity of DNA methyltransferase inhibitors in human cancer cell lines. Cancer Res 2006;66:2794–2800.
Jones P . Effects of 5-azacytidine and its 2′-deoxyderivative on cell differentiation and DNA methylation. Pharmacol Ther 1985;28:17–27.
Fahrner JA, Eguchi S, Herman JG, et al. Dependence of histone modifications and gene expression on DNA hypermethylation in cancer. Cancer Res 2002;62:7213–7218.
Momparler RL, Bovenzi V . DNA methylation and cancer. J Cell Physiol 2000;183:145–154.
Bovenzi V, Le NL, Cote S, et al. DNA methylation of retinoic acid receptor beta in breast cancer and possible therapeutic role of 5-aza-2′-CdR. Anticancer Drugs 1999;10:471–476.
Esteller M . DNA methylation and cancer therapy: new developments and expectations. Curr Opin Oncol 2005;17:55–60.
Esteller M . Epigenetics provides a new generation of oncogenes and tumour-suppressor genes. Br J Cancer 2006b;94:176–183.
Hurtubise A, Momparler RL . Effect of histone deactylase inhibitor LAQ824 on antineoplastic action of 5-aza-2′-deoxycytidine (decitabine) on human breast carcinoma cells. Cancer Chemother Pharmacol 2006;58:618–625.
Jacobs MM . Diet, nutrition and cancer research: an overview. Nutr Today 1993;27:19–23.
Dreosti IE . Nutrition, cancer, and aging. Ann NY Acad Sci 1998;854:371–377.
Gerber B, Muller H, Reimer T, et al. Nutrition and lifestyle factors on the risk of developing breast cancer. Breast Cancer Res Treat 2003;79:265–276.
Geisel J, Schorr H, Bodis M, et al. The vegetarian lifestyle and DNA methylation. Clin Chem Lab Med 2005;43:1164–1169.
Duthie SJ, Narayanan S, Brand GM, et al. Impact of folate deficiency on DNA stability. J Nutr 2002;132:2444S–2449S.
Choi SW, Mason JB . Folate and carcinogenesis: an integrated scheme. J Nutr 2002;132:2350S–23505.
Lee WJ, Shim JY, Zhu BT . Mechanisms of inhibition of DNA methyltransferases by tea catechins and bio flavonoids. Mol Pharmacol 2005;68:1018–1030.
Lee WJ, Zhu BT . Inhibition of DNA methylation by caffeic acid and chlorogenic acid, two common catechol-containing coffee polyphenols. Carcinogenesis 2006;27:269–277.
Catania J, Fairweather DS . DNA methylation and cellular aging. Mutat Res 1991;256:283–293.
Richardson B, Yung R . Role of DNA methylation in the regulation of cell function. J Lab Clin Med 1999;134:333–340.
Issa JP . Aging, DNA methylation and cancer. Crit Rev Onco Hematol 1999;32:31–43.
Baylin SB, Herman JG . DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet 2000;16:168–174.
Acknowledgements
This work was supported by the Claire Booth Luce Fellowship (to AA) and by the National Institutes of Health Grants R01HL070885 (to DKA) and R01HL073349 (to DKA) and Carpenter Chair (to RFM) of Creighton University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Agrawal, A., Murphy, R. & Agrawal, D. DNA methylation in breast and colorectal cancers. Mod Pathol 20, 711–721 (2007). https://doi.org/10.1038/modpathol.3800822
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800822
Keywords
This article is cited by
-
Induced Tamoxifen Resistance is Mediated by Increased Methylation of E-Cadherin in Estrogen Receptor-Expressing Breast Cancer Cells
Scientific Reports (2019)
-
A study of the role of Notch1 and JAG1 gene methylation in development of breast cancer
Medical Oncology (2016)
-
DNA methylation as a promising landscape: A simple blood test for breast cancer prediction
Tumor Biology (2015)
-
Epigenetic regulation of the secreted frizzled-related protein family in human glioblastoma multiforme
Cancer Gene Therapy (2014)
-
Frequent epigenetic inactivation of RSK4 by promoter methylation in cancerous and non-cancerous tissues of breast cancer
Medical Oncology (2014)