Abstract
Despite the high incidence of prostate cancer, only limited data are available on genes or chromosomes specifically involved in its initiation and progression. We have applied comparative genomic hybridization to routinely processed, paraffin-embedded, tissues at different times in prostatic tumor progression to screen the tumor genome for gains and losses. Our panel included specimens derived from 56 different patients: 23 patients with primary, prostate-confined carcinomas; 18 patients with regional lymph node metastases; and 15 patients with distant metastases. Chromosome arms that most frequently showed losses, included 13q (55%), 8p (48%), 6q (43%), 5q (32%), 16q (25%), 18q (20%), 2q (18%), 4q (18%), 10q (18%), and Y (16%). Gains were often seen of chromosome arms 8q (36%), 17q (23%), Xq (23%), 7q (21%), 3q (18%), 9q (18%), 1q (16%), Xp (16%). Furthermore, specific high-level amplifications, eg, of 1q21, 1q25, and Xq12 to q13, were found in metastatic cancers. A significant accumulation of genetic changes in distant metastases was observed, eg, loss of 10q (p = 0.03) and gain of 7q (p = 0.03) sequences. In addition, investigation of a potential biomarker identified in previous studies by our group, ie, extra copies of #7 and/or #8, revealed a high prevalence of 7pq and/or 8q gain in the distant metastases (p = 0.02). Importantly, gains were observed more frequently in tumors derived from progressors after radical prostatectomy, than in nonprogressors (mean time of follow-up, 74 months). Specifically, gain of chromosome 7pq and/or 8q sequences appeared an accurate discriminator between the progressors and nonprogressors. Multivariate analysis showed a significant correlation between progressive disease and the number of chromosomes with gains. This correlation also held true when stage (p = 0.007) or grade (p = 0.002) were taken into account. Likewise, this applied for gain of chromosome 7pq and/or 8q sequences (p = 0.03 and p = 0.005 for stage or grade, respectively). Additionally, an increase in the number of chromosomes with gains per case was related to a decrease in biochemical progression-free survival (Ptrend <0.001). More specifically, the gain of 7pq and/or 8q sequences markedly reduced the biochemical progression-free survival (p < 0.001). In conclusion, this study has, firstly, documented the spectrum of chromosomal alterations in subsequent stages of prostate cancer, a number of which had not been described previously. It allowed us to identify chromosomal regions related to advanced tumor stage, ie, loss of 10q24 and gain of 7q11.2 and/or 7q31 sequences. Secondly, gain of 7pq and/or 8q was identified as a potential genetic discriminator between progressors and nonprogressors after radical surgery.
Similar content being viewed by others
Introduction
Prostate cancer is the most commonly diagnosed, male noncutaneous malignancy and the second leading cause of cancer-related death in men in Western industrialized countries. Its incidence is continuously rising, with over 200,000 new cases diagnosed each year, resulting in 40,000 deaths annually (Landis et al, 1998). From estimates based on i.a. autopsy studies, it is known that approximately 25% to 50% of men over the age of 50 years, who have no clinical evidence of prostate cancer, harbor microscopic foci of well-differentiated to moderately differentiated cancer within the prostate. When these numbers are extrapolated to the American male population, approximately 11,000,000 American men over 50 years of age have latent prostate cancer (Carter and Coffey, 1990; Gittes, 1991). These data suggest that a large portion of prostate cancer is a clinically indolent disease and that only a minority of men with a histological diagnosis of prostate carcinoma will develop clinically significant, life-threatening tumors (Scardino et al, 1992). Population-based, prostate-specific antigen (PSA) screening has resulted in a marked increase in the early detection of prostate cancer (Potosky et al, 1995; Schröder et al, 1998). However, at present it remains unclear what percentage of these early-detected cases represents sub-clinical disease, which might not require aggressive treatment. Therefore, the ability to discriminate indolent prostate cancers from tumors having the potential to progress to lethal metastatic disease has become of paramount importance. Current methods for assessing the prognosis of prostate cancer, such as clinical and pathological staging and histopathological grading, fail to provide consistent predictive information regarding clinical outcome and therapeutic strategy in individual cases (Gittes, 1991; Gleason, 1992). Hence, there is a need for the identification of biological markers for tumor aggressiveness. An understanding of the molecular genetic changes that accompany both the initiation and the progression of prostatic cancer may provide these markers.
As with most types of human cancer, the stepwise accumulation of genetic alterations in genes important for regulation of normal growth (tumor suppressor genes), cell death (involving apoptotic pathways), and/or activation of oncogenes, contribute to the process of carcinogenesis. Conventional cytogenetic studies of prostatic adenocarcinoma have revealed loss of the Y chromosome, trisomy of chromosome 7, del(7)(q22), del(8)(p21), del (10)(q24), and the appearance of double minutes (reviewed in Brothman et al, 1999). Loss of heterozygosity (LOH) analyses have shown frequent loss on chromosome arms 3p, 6q, 7q, 8p, 9p, 10pq, 13q, 16q, 17q, and 18q (Cooney et al, 1996a, 1996b; Cunningham et al, 1996; Dahiya et al, 1997; Gao et al, 1995; Gray et al, 1995; Latil et al, 1997; Li et al, 1999; Melamed et al, 1997; Perinchery et al, 1999; Saric et al, 1999; Takahashi et al, 1995; Ueda et al, 1997; Vocke et al, 1996). Furthermore, comparative genomic hybridization (CGH) analysis applied to a panel of both primary and recurrent tumors revealed losses of 8p and 13q in primary tumors in over 30% of cases, whereas recurrent tumors showed gains of 8q and of chromosomes X and 7, as well as loss of 8p, in over one-half of the cases (Visakorpi et al, 1995b). Further, the recurrent tumors exhibited 8q gain >10 times more often than the primary tumors (Visakorpi et al, 1995b). CGH study of a panel of mostly regional lymph node metastases showed loss of 8p, 10q, 13q, 16q, and 17p, as well as gain of 8q, 1q, 11p, and 3q sequences, in 50% or more of tumors (Cher et al, 1996). Fluorescent in situ hybridization (FISH) studies revealed numerical alterations of chromosomes 7, 8, 10, 16, 17, 18, X, and Y (reviewed in Alers and van Dekken, 1996; Brothman et al, 1999; van Dekken et al, 1997), as well as both deletions and amplifications of specific chromosomal regions, eg, on 8p22 (Huang et al, 1996; Macoska et al, 1994), and MYC on 8q24 (Bubendorf et al, 1999; Jenkins et al, 1997). Furthermore, we (Alers et al, 1997, 1998) and others (Alcaraz et al, 1994; Takahashi et al, 1994) have identified alterations of chromosomes 7 and/or 8 as potential tumor progression markers. Altogether, these molecular (cyto)genetic studies have identified multiple, nonrandom genetic alterations in prostate cancer. However, the precise genetic basis of the development and progression of this disease remains largely unknown.
In the present study, we have applied CGH to formalin-fixed, paraffin-embedded tumor material of patients in various stages of prostatic tumor development. Our goals were to (a) obtain an overview of chromosomal alterations, ie, chromosomal losses, gains, and high-level amplifications, involved in the different stages of prostatic tumor progression; (b) study whether there is an accumulation of genetic changes in prostatic tumor progression; (c) look in more detail for a specific involvement of certain chromosomal alterations during prostatic tumor progression; and (d) assign possible candidate genes underlying prostate cancer.
Results
Overview of Genetic Changes
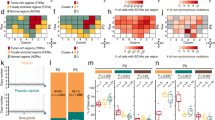
The results of CGH analysis of 56 patients in different stages of prostatic tumor progression (23 primary localized tumors, 18 regional lymph node metastases, and 15 distant metastases) are summarized in Table 1 and Figure 1. Chromosomal imbalances were detected in 52 cases (93%). Fifty cases (89%) showed chromosomal loss, whereas 42 cases (75%) displayed gain of one or more chromosome arms. Chromosomal loss occurred 1.5-fold more often than chromosomal gain (Table 1). Chromosomal losses were frequently (≥15%) encountered on chromosome arms 13q (55%), 8p (48%), 6q (43%), 5q (32%), 16q (25%), 18q (20%), 2q (18%), 4q (18%), 10q (18%), and Y (16%). Chromosomal gains occurring in ≥ 15% of cases were located at chromosomal arms 8q (36%), 17q (23%), Xq (23%), 7q (21%), 3q (18%), 9q (18%), 1q (16%), and Xp (16%). Examples of chromosomal alterations occurring in different stages of prostatic tumorigenesis are depicted in Figure 2. Minimal overlapping regions could be determined at multiple locations: minimal regions of overlap for losses were assigned to 2q21-q22, 4q26-q27, 5q15-q21, 6q15-q16, 8p12-p21, 8p22-pter, 10q24, 13q21, 16q22, and 18q21 and may be indicative for the presence of tumor suppressor genes. Minimal overlapping sites for gains could be found at 1q21, 3q21, 6p21, 7q11.2, 7q31, 8q24, 9q32-qter, 17q11.2, 17q24-qter, and Xq12-q13. These sites may point at oncogenes. Specific and/or high-level amplifications were seen in metastatic tumors only and were found at 1q21 (twice), 1q25, 3q21, 4q21, 4q28.3, 7q32, 10q21.3, and Xq12-q13 (twice) (Fig. 2A).
Chromosomal ideograms showing the summary of DNA copy number changes, detected by CGH, in tumors of 56 patients in different stages of prostatic tumor progression (23 primary, localized tumors, 18 regional lymph node metastases, 15 distant metastases). Losses are displayed on the left of the ideogram (red); gains are shown on the right (green). Minimal regions of overlap are also indicated. Frequent loss (≥ 15%) is detected of 2q, 4q, 5q, 6q, 8p, 10q, 13q, 16q, 18q, and Y sequences. Frequent gain is encountered of 1q, 3q, 7q, 8q, 9q, 17q, and Xpq sequences.
Examples of CGH analysis of archival, formalin-fixed, paraffin-embedded, tumor samples in different stages of prostatic tumor progression. The chromosomal ideograms are shown along with the ratio profiles and the digitized fluorescent images. A, High-level amplifications of several sites in prostatic cancer: 1q21 in a prostatic tumor metastasis to the bone, 1q21 and 1q25 in a regional lymph node metastasis of prostate cancer, 3q21 in a prostatic tumor metastasis to the lung, and Xq12-q13 (androgen receptor locus), in a prostatic bone metastasis of a patient with hormone-refractory disease. B, Most common chromosomal alterations described in this study. All these changes could be detected in early, localized stages. Loss was seen in primary localized tumors of 5q, 6q, 8p, and 13q, whereas gain was distinguished of 8q. The combination of 8p loss and 8q gain was a recurrent finding and might be suggestive for i(8q) formation. C, Possible biomarkers for tumor aggressiveness identified in this study. All the alterations shown here occurred in metastases of prostatic cancer. Gain of 7q, gain of 7pq and/or 8q, loss of 10q, and possibly gain of 17q are regarded as markers for metastatic potential.
Genetic Changes during Tumor Progression
The results of CGH analysis of the three subsequent tumor-node-metastasis (TNM) stages are presented in Table 1. Examples of CGH analysis are shown in Figure 2. The distribution pattern of chromosome arms involved in recurrent (≥ 15%) chromosomal imbalances in primary localized tumors, regional lymph node metastases, and distant metastases is depicted in Figure 3.
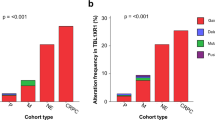
Schematic overview of the distribution of the various genetic alterations occurring in different stages of prostatic tumor progression, ie, primary localized tumors (PAC), regional lymph node metastases (LN META), and distant metastases (DIS META). Losses are depicted in panel A, gains in panel B. Chromosome arms, with alterations occurring in ≥ 15% of cases in one of the groups, are included in this graph. Clearly, some chromosomal alterations, such as loss of 8p and 13q, are seen in all tumor stages, whereas others, eg, gain of 7q, are detected more frequently in distant metastases.
The average number of aberrant chromosomes per case was significantly higher in the distant metastases than in either the primary tumors or the regional lymph node metastases (Fig. 4; p = 0.005 for distant metastases versus primary tumors and p < 0.001 for distant metastases versus regional lymph node metastases). The same was true for the number of chromosomes showing loss or gain per case (Fig. 4; p values, p = 0.007 and p = 0.001 for loss, and p = 0.01 and p = 0.05 for gain, respectively). The mean number of chromosomes showing an aberration, loss, or gain in the regional lymph node metastases was not significantly different from that in the primary tumors. Also, the type of chromosomal alterations did not differ significantly between these two groups. However, some chromosomal alterations, eg, loss of 6q (22% versus 73%; p = 0.005), or gain of 8q (17% versus 60%; p = 0.01), were less frequently encountered in the regional lymph node metastases than in the distant metastases (Fig. 3). This may indicate differential genetic changes for hematogenous versus lymphatic tumor spread.
Comparison of the genetic aberrations between primary localized prostate cancers (PAC), regional lymph node metastases (LN META), and distant metastases (DIS META), showing the average number (± sem) of altered chromosomes per patient. The number of aberrant chromosomes per patient was significantly higher in distant metastases compared with primary tumors (p = 0.005 for total number of aberrations per chromosome, p = 0.007 for chromosomes with loss, p = 0.01 for chromosomes with gain), and compared with the regional lymph nodes (p < 0.001, p = 0.001, p = 0.05, respectively).
To discriminate biological markers for tumor progression, we determined which chromosomal alterations were found more frequently in the distant metastases than in the primary tumors. Gain of chromosome 7q (p = 0.03) and loss of 10q (p = 0.03) sequences occurred significantly more often in the distant metastases than in the primary tumors (13% versus 47% for 7q gain, and 4% versus 33% for 10q loss), whereas gain of 17q showed a tendency (p = 0.09) to appear more frequent in the distant metastases (9% versus 33%) (Fig. 2C and 3). A potential biomarker, ie, gain of chromosome 7 and/or 8, identified in previous studies by our group (Alers et al, 1997, 1998) was investigated in more detail: Gain of chromosomes 7pq and/or 8q, occurred significantly more often in the distant metastases than in the primary tumors (39% versus 80%; p = 0.02).
In the primary tumors a statistically significant correlation was found between tumor stage (pT2-pT4) and the number of gained chromosomes per case (Rs = 0.4779, p = 0.02), but no correlation was found between stage and the number of aberrant or lost chromosomes per case. Specifically, a significant increase in the number of cases with gain of 7pq and/or 8q with increasing tumor stage was observed (Ptrend = 0.02), going from 17% in pT2, to 22% in T3, and to 75% of pT4 staged tumors. In contrast, no statistically significant correlation was found between the number of aberrant, lost, or gained chromosomes per case and tumor grade.
In our primary tumor panel, approximately one-half of the patients (n = 10) showed progression, ie, the occurrence of a biochemical relapse as measured by elevated serum PSA level (see “Materials and Methods” section). In all cases this was followed by local and/or distant recurrences. Importantly, the number of chromosomes with gains per case was significantly higher in progressing patients than in patients who did not display progression in follow-up (n = 11; mean 74 months, range 59 to 87 months; two patients excluded from analysis) (see “Materials and Methods” section). Especially, gains of chromosome 7pq and/or 8q occurred significantly more often in the progressor group than in the nonprogressor group. Multivariate analysis revealed that the number of gained chromosomes was significantly related to progression, even when tumor stage (p = 0.007) or grade (p = 0.002) were taken into account. For gain of 7pq and/or 8q, the corresponding P values were p = 0.03 and p = 0.005. When both stage and grade were taken into account, the additional prognostic value of the number of gained chromosomes was p = 0.002, and of gain of 7pq and/or 8q, p = 0.06. Figure 5A shows that an increase in the number of chromosomes with gains per case was related to a decrease in biochemical progression-free survival (Ptrend < 0.001). Moreover, Figure 5B depicts that the gain of 7pq and/or 8q sequences markedly reduced the biochemical progression-free survival (p < 0.001).
Discussion
This study represents the first detailed genome-wide survey of the locations and frequencies of DNA copy number changes occurring during prostatic tumor progression. The overall genomic profile and the distribution of the chromosomal alterations was in some aspects clearly different from other frequently occurring epithelial tumors, but also some concordant findings were found, such as gain of 8q sequences, which is reported in breast, liver, and esophageal cancer (reviewed in Alers and van Dekken, 1996; van Dekken et al, 1997). We were able to delineate minimal overlapping regions at several chromosomal sites. Furthermore, we identified some new high-level amplifications. The most prevalent and/or relevant chromosomal alterations, as well as some candidate tumor suppressor genes and oncogenes, will be discussed below.
Chromosomal Losses
The most frequent loss involved chromosome 13q with a minimal overlapping region at 13q21. Loss of 13q sequences is a recurrent finding in LOH and CGH studies of prostate cancer (Cher et al, 1996; Cooney et al, 1996b; Hyytinen et al, 1999; Melamed et al, 1997; Nupponen et al, 1998; Ueda et al, 1999; Visakorpi et al, 1995b). Chromosome 13q contains at least three putative tumor suppressor genes: RB1 (13q14), BRCA2 (13q12-q13), and DBM (13q14). Reduced expression of RB has been found in a subset of prostate carcinomas (Bookstein et al, 1990). However, the absence of mutations in both the RB1 and the BRCA2 gene indicated to several authors (Cooney et al, 1996b; Li et al, 1998) that tumor suppressor genes other than RB1 or BRCA1 may be the target of 13q deletions.
Loss of 8p sequences, as detected either by LOH, FISH, or CGH analyses (Cher et al, 1996; Huang et al, 1996; Macoska et al, 1994; Nupponen et al, 1998; Visakorpi et al, 1995b; Vocke et al, 1996), may well be the most common genetic alteration in prostate cancer. This loss also appears to be an early event in prostatic tumorigenesis, because LOH at 8p was described in prostatic intraepithelial neoplasia, the putative precursor lesion of prostate cancer (Emmert-Buck et al, 1995; Saric et al, 1999). In our study we distinguished two minimal overlapping deleted regions: 8p12-p21 and 8p22-pter. These regions coincide with those reported by LOH analysis, with homozygous deletions described in both the 8p12-p21 and the 8p22 region (Bova et al, 1993; Van Alewijk et al, 1999; Vocke et al, 1996). Several candidate genes have been studied at 8p, such as the NKX3.1 gene located at 8p12-p21 (Brothman et al, 1999; He et al, 1997). This homeobox gene was highly expressed in an androgen-dependent LNCaP cell line after androgen stimulation, but undetectable in two androgen-independent prostate cell lines. Another potential candidate gene at 8p22 is the recently identified FEZ1 gene, which codes for a leucine zipper protein the expression of which is altered in many tumors, including prostate, esophageal, and breast cancer (Ishii et al, 1999).
Loss of 6q sequences, with a minimal overlapping region at 6q15-q16, was the third most common chromosomal alteration in this panel. Loss of 6q sequences has been reported as both an early (Brothman et al, 1999) and a late event in prostatic tumorigenesis (Saric et al, 1999; Visakorpi et al, 1995b). Potential candidate genes in this region are the CCNC gene (6q21), which encodes the cell cycle regulatory protein cyclin C (Cooney et al, 1996a), and the transforming growth factor beta-activated kinase 1 gene (TAK1) gene (6q14-q21) (Weizmann, 2000).
Loss of 5q sequences, with a commonly deleted region of 5q15-q21, was more common in our panel than reported in literature (Cunningham et al, 1996); the highest frequency (53%) was seen in distant metastases. Loss on 5q has been associated with higher stage (Cunningham et al, 1996). Chromosome 5q21 contains the APC (5q21-q22)/MCC (5q21) region, of which abnormalities have been observed in prostate tumors (reviewed in Gao et al, 1997).
Loss of 16q, with the minimal overlapping region at 16q22, is one of the most consistent genetic alterations in prostatic cancer (Cher et al, 1996; Latil et al, 1997; Li et al, 1999; Nupponen et al, 1998; Visakorpi et al, 1995b). LOH at 16q23 and 16q24.3 was found to be associated with metastases of prostatic cancer (Latil et al, 1997; Li et al, 1999; Saric et al, 1999). One of the candidate genes is the CDH1 gene (16q22.1) encoding the E-cadherin protein. Immunostaining studies have shown that E-cadherin expression is decreased or absent in a significant proportion of prostate cancers and is linked to the metastatic potential of primary tumors (Umbas et al, 1994). However, sequencing analysis has not revealed mutations of the CDH1 gene (Li et al, 1999). Another candidate gene in this region might be the CTCF gene, which regulates MYC expression (Filippova et al, 1998).
Some of the less frequently detected deleted areas could be connected with known tumor suppressor genes, such as PTEN and MXI1, located at 10q23.3 and 10q24-q25, respectively. In our panel, loss of 10q sequences (minimal overlapping region 10q24) was associated with metastatic disease. This is in concordance with LOH analyses in which loss of 10q occurred more frequently in advanced stages (Komiya et al, 1996; Saric et al, 1999). Moreover, frequent PTEN mutations and deletions were predominantly detected in metastatic prostate cancers (Dong et al, 1998; Vlietstra et al, 1998). Mutation of the MXI1 gene was found in primary prostatic cancers by some authors (Eagle et al, 1995), but not by others (Gray et al, 1995). Finally, loss of 18q sequences preferentially occurred at 18q21 and was reported to be associated with metastasis in prostate cancer (Ueda et al, 1997). This region contains the DCC and DPC4 tumor suppressor genes, of which no abnormalities have been found in prostatic cancers (Gao et al, 1997; Ueda et al, 1997).
Chromosomal Gains
Gain of 8q, often involving the whole chromosome arm, but with a minimal overlapping region at 8q24, was the most common overrepresentation in this panel, occurring in up to 60% of distant metastases. Gain of 8q was often seen in metastases to the bone in association with 8p loss (Alers et al, 1997). This association suggests an isochromosome i(8q) formation (Alers et al, 1997; Bova et al, 1993; Macoska et al, 1994; Virgin et al, 1999). In addition, gain of chromosome 8 is reported to be associated with high tumor grade and stage (Alers et al, 1997; Takahashi et al, 1994). Chromosome 8q24 harbors the MYC oncogene. Amplification of this region was detected in a subset of metastatic and recurrent tumors (Bubendorf et al 1999; Jenkins et al, 1997; Nupponen et al, 1998) and was shown to correlate with the presence of regional lymph node metastases (Van den Berg et al, 1995), as well as with poor prognosis (Sato et al, 1999). Another candidate gene in this region may be the prostate stem cell antigen (PSCA), mapping to 8q24.2, which is overexpressed in the majority of prostate cancer specimens (Reiter et al, 1998).
Gain of chromosome 7, especially gain of 7q, was associated with advanced disease (occurring in 47% of distant metastases). We assigned 7q11.2 and 7q31 as minimal overlapping regions. Gain of chromosome 7 is correlated with high local tumor stage and grade (Alers et al, 1997; Takahashi et al, 1994), whereas aneusomy of chromosome 7 is a potential marker for poor prognosis (Alcaraz et al, 1994; Takahashi et al, 1995). Gain of chromosome 7p12-p21 and 7q11.3-q33 has been detected by CGH analysis in (lymph node) metastases (Cher et al, 1996). Chromosome arm 7p12 contains the EGFR proto-oncogene, a gene possibly involved in prostate cancer (Schwartz et al, 1999) and other human neoplasms. Furthermore, the MET proto-oncogene, which maps to 7q31.1, is expressed in the majority of both primary tumors and metastases (Pisters et al, 1995). However, no mutations of MET have been described in prostate cancer to date (Jenkins et al, 1998).
Gain of 17q sequences was seen in up to 33% of distant metastases, but less frequently in primary tumors. Two minimal overlapping regions for gain could be assigned, ie, 17q11.2 and 17q24-qter. Both loss and gain of 17q sequences have been reported (Cher et al, 1996; Gao et al, 1995). LOH analysis showed a clustering of loss of alleles around the BRCA1 locus (Gao et al, 1995). However, a recent study showed that BRCA1 mutations do not occur in sporadic prostate cancers (Uchida et al, 1999). The ERBB2 (her-2/neu) oncogene (17q21.1) is in the vicinity of one of our minimal overlapping regions. Whereas amplification of ERBB2 is frequently found in breast cancers (reviewed in Alers and van Dekken, 1996; van Dekken et al, 1997), the role of ERBB2 in prostate cancer remains somewhat controversial (Bubendorf et al, 1999; Ross et al, 1997; Schwartz et al, 1999). Recently, a cross-talk between the ERBB2 tyrosine kinase and the androgen receptor signal transduction pathways during prostate cancer progression towards androgen-independent disease was described (Craft et al, 1999). Interestingly, in our panel at least two of the five patients displaying 17q gain had hormone-refractory prostate cancer.
Amplifications
Amplifications, likely representing chromosomal amplicons, were seen as a discrete and/or high peak in the CGH profile in a number of metastatic cancers. The detection of amplifications other than Xq12 is a new finding, since several other CGH studies reported the absence of amplifications in prostate cancer (Cher et al, 1996; Visakorpi et al, 1995b). We detected amplification of 1q21 in two cases. In sarcomas, recurrent gains and amplifications have been demonstrated in 1q21-q23 and 1q21-q31 (reviewed in Knuutila et al, 1998). The 1q25 amplification coincides with the predisposition gene for familial forms of prostate cancer HPC1, linked to 1q24-q25, candidate oncogenes being SKI (1q22-q24) and ABL2 (1q24-q25) (Smith et al, 1996). We also detected a specific amplification of 3q21, a site also observed to be amplified in B cell neoplasms (Knuutila et al, 1998). Further, we distinguished high-level amplifications of the Xq12 region, harboring the androgen receptor locus, in two distant metastases of patients with hormone-refractory disease. Similar findings have been reported in literature (Bubendorf et al, 1999; Cher et al, 1996; Visakorpi et al, 1995a, 1995b). The observation that gain of the whole X chromosome was the most recurrent abnormality of this chromosomes suggests that also other genes on chromosome X, possibly the recently identified second cancer susceptibility locus on Xq27-q28 (Xu et al, 1998), might be of importance for the progression of prostate cancer.
In summary, this study has revealed a spectrum of chromosomal alterations, ie, losses, gains, and specific-high level amplifications, in different stages of prostatic tumor progression. Loss of 13q, 8p, 6q, 5q, and 16q, as well as gain of 8q sequences, were seen in ≥ 25% of primary tumors and may be regarded as (relatively) early genetic events. In general, we observed an accumulation of genetic changes, both losses and gains, in distant metastases compared with both primary tumors and regional lymph node metastases. This may reflect the increased genetic instability of metastasized tumors (Nowell, 1976). Loss of 10q, gain of 7q, and gain of chromosomes 7pq and/or 8q, were detected more frequently in distant metastases than in primary tumors and may therefore be considered as biomarkers for advanced disease. Candidate genes involved are PTEN and/or MXI1 at 10q23.3 and 10q24-q25, respectively, MET at 7q31, and MYC and/or PSCA at 8q24. Gains, especially of chromosome 7pq and/or 8q sequences, were potent genetic markers for adverse prognosis in primary tumors of patients after radical prostatectomy. Our results warrant further evaluation of these genetic markers in a large cohort of patients with localized prostate cancer. It is attractive to speculate that they might yield parameters for an optimal therapeutic strategy in the individual patient.
Materials and Methods
Patient Data
The panel of 60 routinely processed, formalin-fixed, paraffin-embedded, tissue blocks derived from 56 different patients in different stages of prostatic tumor progression, has been previously studied by interphase in situ hybridization (ISH) (Alers et al, 1997, 1998). This panel comprised 26 tumor blocks derived from 23 prostatic adenocarcinomas, from patients who showed no lymph node involvement at the time of radical prostatectomy (N0M0) and represented cases of clinically localized disease. The mean age of these patients was 60 years (range 47 to 70 years). The tumors were pathologically staged according to the pTNM classification (Hermanek et al, 1997). The cases included six pT2 (tumor confined within the prostate), nine pT3 (tumor extends through the prostate capsule), and eight pT4 tumors (tumor is fixed or invades adjacent structures other than seminal vesicles, including microscopical invasion of bladder wall). The tumors were graded according to the Gleason grading system (Gleason, 1992). The mean tumor Gleason score was G7 (range G5 to G10). None of the patients received endocrine or radiation therapy prior to operation.
Follow-up study revealed that 10 of these patients showed progression after radical prostatectomy. Progression was defined as a biochemical relapse, ie, an elevation in PSA level in the serum. In our panel, PSA level elevation was followed by clinical relapse, ie, distant metastases and/or local recurrence in all cases (in general, no histological material was available of the metastases). The usage of elevated (0.2 ng/ml or greater) PSA as a first indicator for imminent local or distant recurrent disease has been reported by several authors (Kupelian et al, 1996). In this study, a biochemical relapse was defined as (a) two consecutive PSA serum levels ≧ 0.2 ng/ml, or (b) a single observation of PSA >1 ng/ml. PSA levels ≧ 0.2 ng/ml occurring in the first six months after radical prostatectomy were not considered a biochemical relapse if followed by undetectable (<0.1 ng/ml) PSA values. The time of biochemical progression was measured from the date of surgery to the date of biochemical relapse detection according to our criteria. The mean time to biochemical progression was 17 months (range 2 to 59 months). The mean PSA level at time of biochemical relapse was 2.3 ng/ml (range 0.3 to 13.1 ng/ml). Thus far, 11 patients are still free from disease in follow-up, with mean time of follow-up being 74 months (range 59 to 87 months). The mean PSA level at the last clinical visit was <0.1 ng/ml (range 0 to <0.1 ng/ml). In two cases the periods of follow-up were too short (8 and 19 months, respectively) to be informative, and these cases were subsequently excluded from multivariate analysis. However, these patients were included in the progression-free survival curves (see below).
Our set of regional metastases comprised 18 pelvic lymph nodes with prostatic tumor metastasis, which were obtained prior to scheduled radical prostatectomy in most cases (all M0). The mean age of these patients was 63 years (range 46 to 75 years). The regional metastases were staged pN1 (18 cases). None of the patients had distant metastasis at time of surgery. One of the patients had received endocrine therapy. Our panel of 16 distant metastases derived from 15 patients comprised 4 distant peri-aortal lymph node metastases, 7 bone metastases, and 5 metastases in other sites like brain (2 cases), liver (1 case), skin (1 case), and lung (1 case). Three of the samples were obtained at time of autopsy. The mean age of the these patients was 65 years (range 41 to 78 years). Four of the patients with advanced disease had received endocrine treatment, four had received radiation therapy, and one patient had received a combination of both endocrine and radiation therapy.
Comparative Genomic Hybridization
Isolation of DNA from the formalin-fixed, paraffin-embedded tumor material was performed as described by (Alers et al 1997). Briefly, the tissue blocks were counterstained in DAPI (0.1 mg/ml in distilled water) for 5 minutes and placed under a fluorescence microscope, enabling a precise selection of the tumor area. Microdissection of the tumor areas was performed using a hollow bore coupled to the microscope. Lower boundaries were checked for the presence of tumor on 4 μm hematoxylin-eosin-stained tissue sections. Excised material was minced using a fine scalpel, deparaffinized in xylene and ethanol series and dried. Samples were digested in 1 ml of extraction buffer (10 mM Tris/HCl, pH 8.0, 100 mM NaCl, 25 mM EDTA, 0.5% SDS, and 300 μg/ml Proteinase K) and incubated at 55° C for 3 to 4 days (Isola et al, 1994). Fresh Proteinase K (300 μg/ml) was added every 24 hours. DNA was subsequently treated with RNAse (1:25 of 10 mg/ml) for 1 hour at 37° C. DNA was isolated according to standard protocols using phenol-chloroform extraction for at least four times. Concentration, purity, and molecular weight of the DNA were estimated using a fluorometer (DyNA Quant 200; Hoefer Biotech Inc., San Francisco, California), UV spectophotometry (Genequant; Pharmacia Biotech, Uppsala, Sweden), and ethidium bromide-stained agarose gels with control DNA series. Tumor DNA with a fragment size of <1 kb (90% of cases) was labeled with a platinum/biotin complex (bio-ULS), using the ULS biotin labeling kit (Kreatech Diagnostics, Amsterdam, The Netherlands), according to the manufacturer’s directions. This method has been described in more detail elsewhere (Alers et al, 1999). Tumor DNA with larger DNA fragment sizes was labeled with biotin by nick translation, according to the manufacturer’s directions (Nick Translation System; Gibco BRL, Gaithersburg, Maryland). Likewise, male reference DNA (Promega, Madison, Wisconsin) was labeled by nick translation with digoxigenin (Boehringer Mannheim, Indianapolis, Indiana). The reaction time and the amount of DNAse were adjusted to obtain a matching probe size for reference and tumor DNAs. Molecular weight of both tumor and reference DNA was checked by gel electrophoresis after nick translation and ranged between 200 and 1500 bp.
CGH was essentially performed according to the procedure described by Kallioniemi et al (1992). In brief, 400 ng of labeled archival tumor DNA, 200 ng of reference DNA, and 15 μg of unlabeled Cot-1 DNA were ethanol precipitated and dissolved in 10 μl of hybridization mixture (50% formamide, 0.1% Tween-20, and 10% dextran sulfate in 2 × SSC at pH 7.0). The probe mixture was denatured and hybridized to normal male metaphase chromosomes (Vysis Inc., Downers Grove, Illinois) for 3 days at 37° C. After washing of the slides, fluorescent detection of the biotin- and digoxigenin-labeled DNA probes was accomplished with avidin-fluorescein isothiocyanate and anti-digoxigenin rhodamine, respectively. Samples were counterstained with 4′, 6′-diamidino-2-phenyl indole in antifade solution.
Images were acquired with an epifluorescent microscope (Leica DM, Rijswijk, The Netherlands) equipped with a cooled CCD camera (Photometrics Inc., Tucson, Arizona), a triple-band pass beam-splitter and emission filters (P-1 filter set; Chroma Technology, Brattleboro, Vermont). For CGH analysis the Quips XL (Version 3.0.1) software (Vysis Inc.) was used. Loss of DNA sequences was defined as chromosomal regions where the mean green:red ratio was below 0.85, while gain was defined as chromosomal regions where the ratio was above 1.15. These threshold values were based on a series of (paraffin–embedded versus fresh) normal controls. Some chromosomal alterations, such as isolated gains of chromosome 1pter, 9q34, 11q13, #19, and 22q, were disregarded and excluded from analysis, because these areas are known to present variation in normal controls (Kirchhoff et al, 1998; Larramendy et al, 1998).
Statistical Analysis
Comparison of the average number of aberrant chromosomes per patient between groups was performed using the Mann-Whitney-U test. Percentages between groups were compared using Fisher’s exact test or the X2 test for trend, if indicated. Comparison of percentages of chromosomal alterations among groups was only performed if the alteration occurred in >10% of total cases. Correlation coefficients (Rs) given are Spearman’s. Multivariate analysis was performed using exact logistic regression. Progression-free survival curves were constructed by the Kaplan-Meier method. Curves were compared using the logrank test or the logrank test for trend if indicated. p = 0.05 (two-sided) was considered the limit of significance.
References
Alcaraz A, Takahashi S, Brown JA, Herath JF, Bergstralh EJ, Larson-Keller JJ, Lieber MM, and Jenkins RB (1994). Aneuploidy and aneusomy of chromosome 7 detected by fluorescence in situ hybridization are markers of poor prognosis in prostate cancer. Cancer Res 54:3998–4002.
Alers JC, Krijtenburg PJ, Hop WCJ, Bolle WA, Schröder FH, van der Kwast ThH, Bosman FT, and van Dekken H (1998). Longitudinal evaluation of cytogenetic aberrations in prostatic cancer: Tumours that recur in time display an intermediate genetic status between non-persistent and metastatic tumours. J Pathol 185:273–283.
Alers JC, Krijtenburg PJ, Rosenberg C, Hop WC, Verkerk AM, Schröder FH, van der Kwast TH, Bosman FT, and van Dekken H (1997). Interphase cytogenetics of prostatic tumor progression: Specific chromosomal abnormalities are involved in metastasis to the bone. Lab Invest 77:437–448.
Alers JC, Rochat J, Krijtenburg PJ, van Dekken H, Raap AK, and Rosenberg C (1999). Universal linkage system (ULS): An improved method for labeling archival DNA for comparative genomic hybridization. Genes Chromosomes Cancer 25:301–305.
Alers JC and van Dekken H (1996). Interphase cytogenetic analysis of solid tumors by non-isotopic DNA in situ hybridization (Review). Prog Histochem Cytochem 31:1–133.
Bookstein R, Rio P, Madreperla SA, Hong F, Allred C, Grizzle WE, and Lee WH (1990). Promotor deletion and loss of retinoblastoma expression in human prostate carcinoma. Proc Natl Acad Sci USA 87:7762–7766.
Bova GS, Carter B, Bussemakers MJ, Emi M, Fujiwara Y, Kyprianou N, Jacobs SC, Robinson JC, Epstein JI, Walsh PC, and Isaacs WB (1993). Homozygous deletion and frequent allelic loss of chromosome 8p22 loci in human prostate cancer. Cancer Res 53:3869–3873.
Brothman AR, Maxwell TM, Cui J, Deubler DA, and Zhu XL (1999). Chromosomal clues to the development of prostate tumors (Review). Prostate 38:303–312.
Bubendorf L, Kononen J, Koivisto P, Schraml P, Moch H, Gasser TC, Willi N, Mihatsch MJ, Sauter G, and Kallioniemi OP (1999). Survey of gene amplifications during prostate cancer progression by high-throughput fluorescence in situ hybridization on tissue microarrays. Cancer Res 59:803–806.
Carter HB and Coffey DS (1990). The prostate: An increasing medical problem. Prostate 16:39–48.
Cher ML, Bova GS, Moore DH, Small EJ, Carroll PR, Pin SS, Epstein JI, Isaacs WB, and Jensen RH (1996). Genetic alterations in untreated metastases and androgen-independent prostate cancer detected by comparative genomic hybridization. Cancer Res 56:3091–3102.
Cooney KA, Wetzel JC, Consolino CM, and Wojno KJ (1996a). Identification and characterization of proximal 6q deletions in prostate cancer. Cancer Res 56:4150–4153.
Cooney KA, Wetzel JC, Meravjer SD, Macoska JA, Singleton TP, and Wojno KJ (1996b). Distinct regions of allelic loss on 13q in prostate cancer. Cancer Res 56:1142–1145.
Craft N, Shostak Y, Carey M, and Sawyers CL (1999). A mechanism for hormone-independent prostate cancer through modulation of androgen receptor signaling by the HER-2/neu tyrosine kinase. Nat Med 5:280–285.
Cunningham JM, Shan A, Wick MJ, McDonnell SK, Schaid DJ, Tester DJ, Qian J, Takahashi S, Jenkins RB, Bostwick DG, and Thibodeau SN (1996). Allelic imbalance and microsatellite instability in prostatic adenocarcinoma. Cancer Res 56:4475–4482.
Dahiya R, McCarville J, Hu W, Lee C, Chui RM, Kaur G, and Deng G (1997). Chromosome 3p24–26 and 3p22–12 loss in human prostatic adenocarcinoma. Int J Cancer 71:20–25.
Dong JT, Sipe TW, Hyytinen ER, Li CL, Heise C, McClintock DE, Grant CD, Chung LW, and Frierson HF Jr (1998). PTEN/MMAC1 is infrequently mutated in pT2 and pT3 carcinomas of the prostate. Oncogene 17:1979–1982.
Eagle LR, Yin X, Brothman AR, Williams BJ, Atkin NB, and Prochownik EV (1995). Mutation of the MXI1 gene in prostate cancer. Nat Genet 9:249–255.
Emmert-Buck MR, Vocke CD, Pozzatti RO, Duray PH, Jennings SB, Florence CD, Zhuang Z, Bostwick DG, Liotta LA, and Linehan WM (1995). Allelic loss on chromosome 8p12–p21 in microdissected prostatic intraepithelial neoplasia. Cancer Res 55:2959–2962.
Filippova GN, Lindblom A, Meincke LJ, Klenova EM, Neiman PE, Collins SJ, Doggett NA, and Lobanenkov VV (1998). A widely expressed transcription factor with multiple DNA sequence specificity, CTCF, is localized at chromosome segment 16q22.1 within one of the smallest regions of overlap for common deletion in breast and prostate cancers. Genes Chromosomes Cancer 22:26–36.
Gao X, Porter AT, and Honn KV (1997). Involvement of the multiple tumor suppressor genes and 12-lipoxygenase in human prostate cancer. Therapeutic implications (Review). Adv Exp Med Biol 407:41–53.
Gao X, Zacharek A, Grignon DJ, Sakr W, Powell IJ, Porter AT, and Honn KV (1995). Localization of potential tumor suppressor loci to a < 2 Mb region on chromosome 17q in human prostate cancer. Oncogene 11:1241–1247.
Gittes RF (1991). Carcinoma of the prostate. N Eng J Med 324:236–245.
Gleason DF (1992). Histologic grading of prostate cancer. Hum Pathol 23:273–279.
Gray IC, Philips SMA, Lee SJ, Neoptolemos JP, Weissenbach J, and Spurr NK (1995). Loss of chromosomal region 10q23–q25 in prostate cancer. Cancer Res 55:4800–4803.
He WW, Sciavolino PJ, Wing J, Augustus M, Hudson P, Meissner PS, Curtis RT, Shell BK, Bostwick DG, Tindall DJ, Gelmann EP, Abate-Shen C, and Carter KC (1997). A novel human prostate-specific, androgen-regulated homeobox gene (NKX3.1) that maps to 8p21, a region frequently deleted in prostate cancer. Genomics 43:69–77.
Hermanek P, Hutter RVP, Sobin LH, Wagner G, and Wittekind CH, editors (1997). TNM atlas: Illustrated guide to the TNM/pTNM classification of malignant tumors, 4th ed. Berlin: Springer Verlag, 272–280.
Huang SF, Xiao S, Renshaw AA, Loughlin KR, Hudson TJ, and Fletcher JA (1996). Fluorescence in situ hybridization evaluation of chromosome deletion patterns in prostate cancer. Am J Pathol 149:1565–1573.
Hyytinen ER, Frierson HF Jr, Boyd JC, Chung LW, and Dong JT (1999). Three distinct regions of allelic loss at 13q14, 13q21–q22, and 13q33 in prostate cancer. Genes Chromosomes Cancer 25:108–114.
Ishii H, Baffa R, Numata SI, Murakumo Y, Rattan S, Inoue H, Mori M, Fidanza V, Alder H, and Croce CM (1999). The FEZ1 gene at chromosome 8p22 encodes a leucine-zipper protein, and its expression is altered in multiple human tumors. Proc Natl Acad Sci USA 96:3928–3933.
Isola J, DeVries S, Chu L, Ghazvini S, and Waldman F (1994). Analysis of changes in DNA sequence copy number by comparative genomic hybridization in archival paraffin-embedded tumor samples. Am J Pathol 145:1301–1308.
Jenkins RB, Qian J, Lee HK, Huang H, Hirasawa K, Bostwick DG, Proffitt J, Wilber K, Lieber MM, Liu W, and Smith DI (1998). A molecular cytogenetic analysis of 7q31 in prostate cancer. Cancer Res 58:759–766.
Jenkins RB, Qian J, Lieber MM, and Bostwick DG (1997). Detection of c-myc oncogene amplification and chromosomal anomalies in metastatic prostatic carcinoma by fluorescence in situ hybridization. Cancer Res 57:524–531.
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, and Pinkel D (1992). Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 258:818–821.
Kirchhoff M, Gerdes T, Rose H, Maahr J, Ottesen AM, and Lundsteen C (1998). Detection of chromosomal gains and losses in comparative genomic hybridization analysis based on standard reference intervals. Cytometry 31:163–173.
Knuutila S, Björkqvist AM, Autio K, Tarkkanen M, Wolf M, Monni O, Szymanska J, Larramendy ML, Tapper J, Pere H, El-Rifai W, Hemmer S, Wasenius VM, Vidgren V, and Zhu Y (1998). DNA copy number amplifications in human neoplasms: Review of comparative genomic hybridization studies (Review). Am J Pathol 152:1107–1123.
Komiya A, Suzuki H, Ueda T, Yatani R, Emi M, Ito H, and Shimazaki J (1996). Allelic losses at loci on chromosome 10 are associated with metastasis and progression of human prostate cancer. Genes Chromosomes Cancer 17:245–253.
Kupelian P, Katcher J, Levin H, Zippe C, and Klein E (1996). Correlation of clinical and pathologic factors with rising prostate-specific antigen profiles after radical prostatectomy alone for clinically localized prostate cancer. Urology 48:249–260.
Landis SH, Murray T, Bolden S, and Wingo PA (1998). Cancer statistics, 1998. CA Cancer J Clin 48:6–29.
Larramendy ML, El-Rifai W, and Knuutila S (1998). Comparison of fluorescein isothiocyanate- and Texas red-conjugated nucleotides for direct labeling in comparative genomic hybridization. Cytometry 31:174–179.
Latil A, Cussenot O, Fournier G, Driouch K, and Lidereau R (1997). Loss of heterozygosity at chromosome 16q in prostate adenocarcinoma: Identification of three independent regions. Cancer Res 57:1058–1062.
Li C, Berx G, Larsson C, Auer G, Aspenblad U, Pan Y, Sundelin B, Ekman P, Nordenskjöld M, van Roy F, and Bergerheim USR (1999). Distinct deleted regions on chromosome segment 16q23–24 associated with metastases in prostate cancer. Genes Chromosomes Cancer 24:175–182.
Li C, Larsson C, Futreal A, Lancaster J, Phelan C, Aspenblad U, Sundelin B, Liu Y, Ekman P, Auer G, and Bergerheim USR (1998). Identification of two distinct deleted regions on chromosome 13 in prostate cancer. Oncogene 16:481–487.
Macoska JA, Trybus TM, Sakr WA, Wolf MC, Benson PD, Powell IJ, and Pontes JE (1994). Fluorescence in situ hybridization analysis of 8p allelic loss and chromosome 8 instability in human prostate cancer. Cancer Res 54:3824–3830.
Melamed J, Einhorn JM, and Ittmann MM (1997). Allelic loss on chromosome 13q in human prostate carcinoma. Clin Cancer Res 3:1867–1872.
Nowell PC (1976). The clonal evolution of tumor cell populations. Science 194:23–28.
Nupponen NN, Kakkola L, Koivisto P, and Visakorpi T (1998). Genetic alterations in hormone-refractory recurrent prostate carcinomas. Am J Pathol 153:141–148.
Pisters LL, Troncoso P, Zhau HE, Li W, von Eschenbach AC, and Chung LW (1995). C-met proto-oncogene expression in benign and malignant human prostate tissues. J Urol 154:293–298.
Perinchery G, Bukurov N, Nakajima K, Chang J, Li LC, and Dahiya R (1999). High frequency of deletion on chromosome 9p21 may harbor several tumor-suppressor genes in human prostate cancer. Int J Cancer 83:610–614.
Potosky AL, Miller BA, Albertsen PC, and Kramer BS (1995). The role of increasing detection in the rising incidence of prostate cancer. JAMA 273:548–552.
Reiter RE, Gu Z, Watabe T, Thomas G, Szigeti K, Davis E, Wahl M, Nisitani S, Yamashiro J, Le Beau MM, Loda M, and Witte ON (1998). Prostate stem cell antigen: A cell surface marker overexpressed in prostate cancer. Proc Natl Acad Sci USA 95:1735–1740.
Ross JS, Sheehan C, Hayner-Buchan AM, Ambros RA, Kallakury BV, Kaufman R Jr, Fisher HAG, Rifkin MD, and Muraca PJ (1997). Prognostic significance of HER-2/neu gene amplification status by fluorescence in situ hybridization of prostate carcinoma. Cancer 79:2162–2170.
Saric T, Brkanac Z, Troyer DA, Padalecki SS, Sarosdy M, Williams K, Abadesco L, Leach RJ, and O’Connell P (1999). Genetic pattern of prostate cancer progression. Int J Cancer 81:219–224.
Sato K, Qian J, Slezak JM, Lieber MM, Bostwick DG, Bergstralh EJ, and Jenkins RB (1999). Clinical significance of alterations of chromosome 8 in high-grade, advanced, nonmetastatic prostate carcinoma. J Natl Cancer Inst 91:1574–1580.
Scardino PT, Weaver R, and Hudson MA (1992). Early detection of prostate cancer. Hum Pathol 23:211–222.
Schröder FH, van der Maas P, Beemsterboer P, Boeken Kruger A, Hoedemaeker R, Rietbergen J, and Kranse R (1998). Evaluation of the digital rectal examination as a screening test for prostate cancer. Rotterdam section of the European Randomized Study of Screening for Prostate Cancer. J Natl Cancer Inst 90:1817–1823.
Schwartz S Jr, Caceres C, Morote J, De Torres I, Rodriguez-Vallejo JM, Gonzalez J, and Reventos J (1999). Gain of the relative genomic content of erbB-1 and erbB-2 in prostate carcinoma and their association with metastasis. Int J Oncol 14:367–371.
Smith JR, Freije D, Carpten JD, Grönberg H, Xu J, Isaacs SD, Brownstein MJ, Bova GS, Guo H, Bujnovszky P, Nusskern DR, Damber JE, Bergh A, Emanuelsson M, Kallioniemi OP, Walker-Daniels J, Bailey-Wilson JE, Beaty TH, Meyers DA, Walsh PC, Collins FS, Trent JM, and Isaacs WB (1996). Major susceptibility locus for prostate cancer on chromosome 1 suggested by a genome-wide search. Science 274:1371–1374.
Takahashi S, Qian J, Brown JA, Alcaraz A, Bostwick DG, Lieber MM, and Jenkins RB (1994). Potential markers of prostate cancer aggressiveness detected by fluorescence in situ hybridization. Cancer Res 54:3574–3579.
Takahashi S, Shan AL, Ritland SR, Delacey KA, Bostwick DG, Lieber MM, Thibodeau SN, and Jenkins RB (1995). Frequent loss of heterozygosity at 7q31.1 in primary prostate cancer is associated with tumor aggressiveness and progression. Cancer Res 55:5115–5119.
Uchida T, Wang C, Sato T, Gao J, Takashima R, Irie A, Ohori M, and Koshiba K (1999). BRCA1 gene mutation and loss of heterozygosity on chromosome 17q21 in primary prostate cancer. Int J Cancer 84:19–23.
Ueda T, Emi M, Suzuki H, Komiya A, Akakura K, Ichikawa T, Watanabe M, Shiraishi T, Masai M, Igarashi T, and Ito H (1999). Identification of a 1-cM region of common deletion on 13q14 associated with human prostate cancer. Genes Chromosomes Cancer 24:183–190.
Ueda T, Komiya A, Emi M, Suzuki H, Shiraishi T, Yatani R, Masai M, Yasuda K, and Ito H (1997). Allelic losses on 18q21 are associated with progression and metastasis in human prostate cancer. Genes Chromosomes Cancer 20:140–147.
Umbas R, Isaacs WB, Bringuier PP, Schaafsma HE, Karthaus HFM, Oosterhof GON, DeBruyne FMJ, and Schalken JA (1994). Decreased E-cadherin expression is associated with poor prognosis in patients with prostate cancer. Cancer Res 54:3929–3933.
Van Alewijk DC, Van der Weiden MM, Eussen BJ, Van den Andel-Thijssen LD, Ehren-van Eekelen CC, Konig JJ, van Steenbrugge GJ, Dinjens WN, and Trapman J (1999). Identification of a homozygous deletion at 8p12–21 in a human prostate cancer xenograft. Genes Chromosomes Cancer 24:119–126.
van Dekken H, Rosenberg C, Krijtenburg PJ, and Alers JC (1997). Interphase cytogenetics and comparative genomic hybridization of human epithelial cancers and precursor lesions. Histochem Cell Biol 108:419–430.
Van den Berg C, Guan XY, von Hoff D, Jenkins R, Bittner M, Griffin C, Kallioniemi O, Visakorpi T, McGill J, Herath J, Epstein J, Sarosdy M, Meltzer P, and Trent J (1995). DNA sequence amplification in human prostate cancer identified by chromosome microdissection: Potential prognostic implications. Clin Cancer Res 1:11–18.
Virgin JB, Hurley PM, Nahhas FA, Bebchuk KG, Mohamed AN, Sakr WA, Bright RK, and Cher ML (1999). Isochromosome 8q formation is associated with 8p loss of heterozygosity in a prostate cancer cell line. Prostate 41:49–57.
Visakorpi T, Hyytinen E, Koivisto P, Tanner M, Keinanen R, Palmberg C, Palotie A, Tammela T, Isola J, and Kallioniemi OP (1995a). In vivo amplification of the androgen receptor gene and progression of human prostate cancer. Nat Genet 9:401–406.
Visakorpi T, Kallioniemi AH, Syvänen AC, Hyytinen ER, Karhu R, Tammela T, Isola JJ, and Kallioniemi OP (1995b). Genetic changes in primary and recurrent prostate cancer by comparative genomic hybridization. Cancer Res 55:342–347.
Vlietstra RJ, Van Alewijk DC, Hermans KG, van Steenbrugge GJ, and Trapman J (1998). Frequent inactivation of PTEN in prostate cancer cell lines and xenografts. Cancer Res 58:2720–2723.
Vocke CD, Pozzatti RO, Bostwick DG, Florence CD, Jennings SB, Strup SE, Duray PH, Liotta LA, Emmert-Buck MR, and Linehan WM (1996). Analysis of 99 microdissected prostate carcinomas reveals a high frequency of allelic loss on chromosome 8p12–p21. Cancer Res 56:2411–2416.
Weizmann Institute of Science Genome and Bioinformatics (2000). Weizmann Institute of Science, Israel. (http://bioinfo.weizmann.ac.il)
Xu J, Meyers D, Freije D, Isaacs S, Wiley K, Nusskern D, Ewing C, Wilkens E, Bujnovszky P, Bova GS, Walsh P, Isaacs W, Schleutker J, Matikainen M, Tammela T, Visakorpi T, Kallioniemi OP, Berry R, Schaid D, French A, McDonnell S, Schroeder J, Blute M, Thibodeau S, Grönberg H, Emanuelsson M, Damber JE, Bergh A, Jonsson BA, Smith J, Bailey-Wilson J, Carpten J, Stephan D, Gillanders E, Amundson I, Kainu T, Freas-Lutz D, Baffoe-Bonnie A, Van Aucken A, Sood R, Collins F, Brownstein M, and Trent J (1998). Evidence for a prostate cancer susceptibility locus on the X chromosome. Nat Genet 20:175–179.
Acknowledgements
This study was supported by grants from the Dutch Cancer Society, Grant No. EUR 97–1404, and the Sacha Swarttouw-Hijmans Foundation. CR was supported by the Postgraduate School Molecular Medicine-Pathophysiology of Growth and Differentiation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Alers, J., Rochat, J., Krijtenburg, PJ. et al. Identification of Genetic Markers for Prostatic Cancer Progression. Lab Invest 80, 931–942 (2000). https://doi.org/10.1038/labinvest.3780096
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780096
This article is cited by
-
Identification of FBXL4 as a Metastasis Associated Gene in Prostate Cancer
Scientific Reports (2017)
-
ASEQ: fast allele-specific studies from next-generation sequencing data
BMC Medical Genomics (2015)
-
EPMA position paper in cancer: current overview and future perspectives
EPMA Journal (2015)
-
Race-associated biological differences among Luminal A breast tumors
Breast Cancer Research and Treatment (2015)
-
Entwicklung eines dreidimensionalen Prostatakarzinomzellkulturmodells
Der Urologe (2008)