Abstract
The canonical Notch pathway that has been well characterized over the past 25 years is relatively simple compared to the plethora of recently published data suggesting non-canonical signaling mechanisms and cross talk with other pathways. The manner in which other pathways cross talk with Notch signaling appears to be extraordinarily complex and, not surprisingly, context-dependent. While the physiological relevance of many of these interactions remains to be established, there is little doubt that Notch signaling is integrated with numerous other pathways in ways that appear increasingly complex. Among the most intricate cross talks described for Notch is its interaction with the NF-κB pathway, another major cell fate regulatory network involved in development, immunity, and cancer. Numerous reports over the last 11 years have described multiple cross talk mechanisms between Notch and NF-κB in diverse experimental models. This article will provide a brief overview of the published evidence for Notch–NF-κB cross talk, focusing on vertebrate systems.
Similar content being viewed by others
CANONICAL NOTCH SIGNALING
Canonical Notch signaling has been recently reviewed by several authors,1, 2, 3, 4, 5, 6 and the reader is referred to these reviews for detailed information and additional references. Briefly, there are four mammalian Notch receptors (Notch-1, -2, -3, and -4) and five ligands (Jagged-1/-2, Delta-like 1, 3, and 4).7 Notch receptors and ligands are heterodimeric type I membrane proteins that require cell–cell contact for activation to regulate cell fate decisions. Notch signaling plays context-dependent roles in differentiation, proliferation, and apoptosis, and has been implicated in cancer progression both as an oncogene and a tumor suppressor. Notch receptors heterodimers include an extracellular subunit (NEC) containing multiple EGF-like repeats and three LIN-12-like repeats, and a transmembrane subunit (NTM).1 The 103-amino-acid C-terminal hydrophobic region of NEC, together with the 65 amino acids at the N terminus of the NTM subunit, forms the ‘heterodimerization domain’ (HD), which mediates the interaction between NEC and NTM.8 The LNR repeats prevent ligand-independent dissociation of the two subunits by ‘folding over’ the HD.9 Upon ligand binding to NEC, NTM and NEC dissociate and the ligand-bound NEC is endocytosed into the ligand-expressing cell.10 This unmasks the HD and triggers an extracellular cleavage in it by ADAM (a disintegrin and metalloproteinase) 10 or 17,1, 2 followed by an intramembranous cleavage by the γ-secretase complex.11 The cleavage of NTM by γ-secretase releases an intracellular domain of Notch of approximately 97 kDa (NIC) that translocates to the nucleus to act as a switch for transcription factor CSL (CBF-1/suppressor of hairless/Lag-1, also known as RBP-Jκ in mammals). CSL binds to responsive elements whose canonical sequence is CGTGGGAA and represses target genes by recruiting a multi-protein corepressor complex.1, 2 Upon binding of NIC to CSL, the corepressor complex is replaced with a coactivator complex which includes MAML1 (Mastermind in Drosophila), SKIP, and histone acetyltransferases PCAF, GCN5, or p300. This converts CSL into a transcriptional activator, driving the transcription of CSL target genes. These include among others the HES and HEY families, homologs of Drosophila ‘Enhancer of Split’ bHLH transcription factors.12 Other Notch target genes have been shown to play critical roles in carcinogenesis and cancer progression. These include the following: p21Cip/Waf,13 cyclin D1,14 cyclin A,15 several subunits of nuclear factor-κB (NF-κB),16 PPAR family nuclear receptors,17, 18 E3 ubiquitin ligase SKP2, which mediates degradation of p27kip1,19 c-Myc,20, 21 and many others.
NF-κB SIGNALING
A comprehensive review of NF-κB biology is beyond the scope of this manuscript, and the reader is referred to recent articles for further information.22, 23, 24, 25, 26, 27, 28, 29, 30 NF-κB/Rel transcription factors control the expression of a multitude of genes with multiple functions in immunity, inflammation, and cancer.29, 30 The NF-κB family include homo- or heterodimers formed by p50, p65 (RelA), c-Rel, p52, and RelB.29, 30 All five proteins contain a Rel homology domain (RHD), which participates in dimerization and DNA binding. The RHD contains a nuclear localization sequence (NLS), which in non-stimulated cells is masked through binding of IκB family inhibitory proteins. These include IκB-α, IκB-β, IκB-ɛ, Bcl-3, as well as NF-κB precursor proteins p100 (p52 precursor) and p105 (p50 precursor).30 IκB-α, IκB-β, and IκB-ɛ physically interact with and sequester NF-κB in the cytosol.31, 32 IκBα also contains a nuclear export signal, which allows it to sequester NF-κB in the nucleus and export it back to the cytoplasm.30 Both p100 and p105 have IκB activity. Bcl-3, conversely, is a nuclear protein that functions primarily as a coactivator. A common feature of these proteins is the presence of five to seven ankyrin repeats, which are required for the interaction with NF-κB.33 The ankyrin region binds the RHD, masking the NLS and preventing nuclear transport of NF-κB.
There are at least three mechanisms for activation of NF-κB:24, 30 (1) The canonical IKK pathway involves phosphorylation of IκB proteins by a specific kinase (IKK) ‘signalosome’ multiprotein complex, consisting of at least three subunits, IKKα, IKKβ, and multiple copies of regulatory subunit IKKγ/NEMO. The IKK complex is activated by stimuli such as T-cell receptor (TCR) ligation, TNF-α, IL-1, or LPS. The IKK signalosome, acting primarily through the catalytic activity of IKK-β, phosphorylates IκBα Ser32 and Ser36 or equivalent residues in IκBβ and ɛ.30 This triggers polyubiquitination at Lys21 and Lys22 of IκBα (or equivalent residues in IκBβ and ɛ), and proteasomal degradation of IκBs, allowing nuclear transport of NF-κB dimers. Activation of the IKK signalosome is mediated, at least in some cases, by monoubiquitination of IKKγ/NEMO, which helps recruit ubiquitin-binding proteins to the complex, such as kinase TAK1,34 which ultimately activate IKKβ. In the case of genotoxic effects, IKK activation can involve nuclear SUMOylation of IKKγ/NEMO, its phosphorylation by atangia–telangiectasia (ATM) kinase, replacement of SUMO with ubiquitin, nuclear export of an ATM/NEMO complex and recruitment of this complex to the IKK signalosome.30 (2) The non-canonical pathway is activated by ligands such as CD40L and lymphotoxin. These activate NIK kinase, which in turn activates and phosphorylates IKKα homodimers. IKKα phosphorylates the p100 precursor of p52, causing its partial degradation by the proteasome to generate p52/RelB heterodimers, or ‘NF-κB2’. These have affinity for a subset of κB responsive elements and generate a distinctive gene expression pattern.30 (3) Atypical or IKK-independent pathways, activated by stimuli such as hypoxia, oxidizing radicals, or UV radiation, which involve either Tyr phosphorylation of IκBα on Tyr 42 or Ser/Thr phosphorylation in its C-terminal PEST domain, leading to IκBα ubiquitination and degradation.30
ROLES OF IKK KINASES
IKKα and β, although structurally very similar, have different roles in the NF-κB pathway, and different NF-κB-independent functions. IKKβ, as a component of the IKK signalosome, appears predominant in the canonical pathway activation cascade and IκBα phosphorylation, while IKKα is predominant in the activation of the alternative, p52/RelB pathway (see above). Additionally, IKKα has multiple nuclear functions within and outside the NF-κB pathway. These include chromatin remodeling at NF-κB-dependent promoters by phosphorylation of Histone H3 (Ser10),35 phosphorylation of p65/RelA,36 and phosphorylation of co-repressor SMRT, leading to NF-κB-dependent transcriptional activation.37 IKKα can increase cyclin D1 expression by phosphorylating and activating the estrogen receptor-α (ERα) and it coactivator SRC-3 at the cyclin D1 promoter,38, 39 as well as by phosphorylating β-catenin at the same promoter40 and stabilizing cytoplasmic β-catenin.41 Both these activities may contribute to oncogenic effects, especially in breast cancer. In a possible feedback loop, IKKα can also directly phosphorylate cyclin D1, leading to its degradation.42
IKKβ, for its part, participates in regulating the TCR signal in a complex feedback loop. First, it functions downstream of the TCR in a T-cell specific complex including CARMA, BCL-10, and MALT1 (CBM), which leads to canonical NF-κB activation.30 Subsequently, it phosphorylates BCL-10, leading to its dissociation from the CBM complex and attenuating the signal.43 IKKβ phosphorylates p65/RelA at Ser536, stimulating its nuclear migration.44 IKKβ also phosphorylates and inactivates proapoptotic transcription factor FOXO3a.45 By phosphorylating a 14-3-3 protein that binds AU-rich binding tristetraprolin (TPP), IKKβ inhibits the binding of TPP to multiple growth factor and cytokine mRNAs, stabilizing these transcripts.46 IKKβ can also activate, and in some cases attenuate, the MEK–ERK pathway through phosphorylation of p105, which releases ERK pathway kinase TPL2/COT, and conversely, through degradation of scaffold protein DOK1.47, 48
This list of functions far from exhaustive, but it illustrates the complexity of effects through which the IKK kinases can mediate cross talk between NF-κB-activating stimuli and multiple other pathways regulating proliferation, apoptosis, and inflammation.
CROSS TALK BETWEEN NOTCH AND NF-κB
Over the past 11 years, numerous reports have described regulation of NF-κB by Notch and vice versa through different, context-dependent mechanisms. We will briefly summarize the mechanisms of cross talk that have been described so far (Table 1).
Transcriptional Regulation of NF-κB Pathway Members by Notch
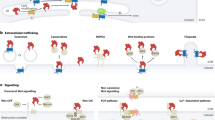
Oswald et al49 originally described transcriptional activation of the p100 promoter by CSL-responsive element that overlaps with a κB site. Through this effect, Notch would increase expression of the p52 precursor (Figure 1a). The overall effect would then depend on cellular context. Since p100 has IκB activity, the end result could be inhibition of the non-canonical pathway (see above). However, in the presence of active IKKα, this could also result in the rapid generation of p52/RelB heterodimers. Subsequently, Cheng et al16 showed that Notch-1 upregulates the expression of p50, p65, RelB, and c-Rel NF-κB subunits in murine bone marrow hematopoietic precursors (Figure 1a). Through this effect, Notch-1 controlled the LPS-induced proliferation of B cells and GM-CSF-induced differentiation of dendritic cells. Oakley et al50 reported that in stellate liver cells, CSL in the absence of Notch activation acts as a repressor of IκBα expression. Notch-1IC caused de-repression of IκBα expression, and thus lower NF-κB activity.
(a) Reciprocal transcriptional regulation between Notch and NF-κB. Notch has been reported to increase transcription of p52, p50, p65/RelA, RelB, and c-Rel, as well as IκBα, thus potentially stimulating or inhibiting NF-κB activity. Conversely, NF-κB has been reported to increase transcription of Notch ligand Jagged-1, as well as Notch targets Deltex-1 and HES-5. (b) Non-transcriptional cross talk mechanisms. Notch-1IC binds to p50, causing either inhibition or stimulation of NF-κB p50/p65 depending on relative amounts. Also, Notch-1IC by binding p50 blocks the binding of IκBα to p50/c-Rel complexes, thus preventing IκBα from causing nuclear export of p50/c-Rel. This results in increased permanence of p50/c-Rel in the nucleus and increased transcriptional activity. Notch-3IC binds and activates IKKα homodimers, resulting in activation of the alternate ‘NF-κB2’ pathway. Notch-1IC binds and activates the IKK signalosome, resulting in activation of the canonical NF-κB pathway. The distinct nuclear consequences of the canonical NF-κB and alternate ‘NF-κB2’ pathway are not depicted for clarity. In the nucleus, IKKα can de-repress Notch transcriptional activity by phosphorylating SMRT and/or by phosphorylating nuclear IκBα, which appears to function as a nuclear repressor of Notch activity.
Transcriptional Regulation of Notch Pathway Members by NF-κB
This cross talk mechanism has been described primarily in the context of B cells (Figure 1a). Bash et al51 reported transcriptional upregulation of Notch ligand Jagged-1 by NF-κB in B cells. More recently, Notch-2 and NF-κB have been reported to cooperate in marginal zone (MZ) B-cell development. In this model, NF-κB stimulates the expression of two known Notch targets, HES-5 and Deltex-1.52
Physical Interaction between Notch and NF-κB
In 1996, Guan et al53 reported that ectopically overexpressed Notch-1IC has an IκB-like activity in Jurkat cells, specifically associating with the p50 subunit of p50–p65 heterodimers and preventing κB-dependent transactivation. Interestingly, in that report, the authors showed the effect to be dose-dependent in co-transfection experiments (Figure 1b). Low amounts of Notch-1 stimulated NF-κB transcriptional activity, while higher amounts inhibited it. Subsequently, the same group mapped the interaction between Notch-1IC and p50 to a 109 amino-acid stretch including residues 1773–1881.54 This is at the very N terminus of Notch-1IC and overlaps with the RAM23 domain, which participates in Notch interaction with CSL. This would imply that these two interactions of Notch-1 may compete with one another. Under the conditions studied in that report, where Notch-1IC was overexpressed by transfection, the interaction was predominantly nuclear and had inhibitory effects on NF-κB.54 More recent data from the Osborne group, while confirming the physical interaction between Notch-1 and p50, have the opposite functional implications. First, Palaga et al55 showed that Notch-1 activates NF-κB downstream of TCR activation in murine T cells. The mechanism of this effect was later clarified by the same group. Shin et al56 recently showed that Notch-1 is responsible for the late but not the early wave of NF-κB activation that follows TCR activation. This effect is mediated by p50-mediated physical interaction of Notch-1IC with p50/c-Rel complexes. Through this interaction, Notch-1 competes with IκBα and prevents it from exporting NF-κB from the nucleus (Figure 1b). This increased nuclear retention results in activation of NF-κB-dependent transcription. So far, only Notch-1 has been reported to interact with NF-κB. Whether other Notch homologs have similar activities remains unclear.
Notch and the IKKs
Nickoloff et al12 showed that in normal human keratinocytes, Notch activation by a soluble ligand could trigger terminal differentiation and cornification. In that study, the authors showed that Notch activation was associated with rapid activation of NF-κB. This lasted approximately 2 h and was suppressed by a late increase in PPARα. The authors also showed that the soluble Notch ligand triggered rapid phosphorylation of IκBα, suggesting that the canonical, IKK signalosome-mediated pathway was somehow activated by the Notch ligand. Similar effects were observed by the same group in murine erythroleukemia (MEL) cells undergoing erythroid differentiation.57 In this model, Notch-1 expression is required to maintain survival during differentiation, but must be downregulated for terminal differentiation to occur. Antisense-mediated knockdown of Notch-1 caused differentiating cells to undergo apoptosis, while constitutive overexpression of Notch-1IC prevented differentiation. In an effort to dissect some of the pathways responsible for the survival effects of Notch-1 in this context, the authors showed that exposure of MEL cells to stromal cells expressing Notch ligand Jagged-2 caused rapid and short-lived activation of NF-κB, which was reversed by 4 h.57
Recent observations have shed some light on the possible mechanisms for rapid activation of NF-κB by Notch. Bellavia et al58 showed that Notch-3 activates multiple NF-κB pathways during T-cell differentiation. Notch-3IC overexpression causes a T-cell leukemia/lymphoma phenotype in transgenic mice. Transgenic premalignant thymocytes and T lymphoma cells overexpress pTα/pre-TCR and have constitutive NF-κB activation providing survival signals for immature thymocytes. Notch-3 can activate the canonical p50/p65 pathway in a way that is dependent on an IKKβ/IKKα/NIK. NF-κB p50/p65 complexes are recruited onto the cyclin D1, Bcl2-A1, and IL7-receptor-α promoters, resulting in T-cell survival and proliferation.58, 59 In the same cells but in the absence of pTα, Notch-3IC activates the non-canonical NF-κB pathway by physically binding to IKKα homodimers in an NIK-independent manner (Figure 1b). The resulting NF-κB2/p100 processing allows p52/RelB nuclear translocation triggering transcription of Bcl2-A1 and IL7-receptor-α genes.60 These observations raise the question of whether the canonical and/or the alternative NF-κB pathways contribute to the transforming effect of Notch-3 in this model. It is still unclear, however, whether endogenously expressed Notch-3 participates in activation of NF-κB through either of these mechanisms.
Vilimas et al,61 examining a murine model of T-cell acute lymphoblastic leukemia induced by overexpression of Notch-1IC, determined that NF-κB target genes are upregulated in expression profiles of Notch-1IC-transformed cells. In these cells, and in human cell lines derived from spontaneous T-cell acute lymphoblastic leukemia, Notch-1IC interacts with the IKK signalosome, increasing its IκBα kinase activity (Figure 1b). Thus, it appears that at least two Notch homologs have the ability to bind IKK kinases, increasing their activity. It remains unclear whether IKK regulation is a physiological function of Notch-1 and/or -3 or a consequence of high-level expression, either in transgenic models or as a consequence of Notch mutations in human T-ALL. Another side of the cross talk between Notch and IKKs has been uncovered by the Bigas group. This group has recently discovered that in colorectal carcinoma cells, nuclear IKKα phosphorylates SMRT not only in association with NF-κB but also in association with CSL.62 This leads to uncontrolled activation of Notch-mediated signaling (Figure 1b). Interestingly, the same group had described earlier a nuclear function of IκBα as an inhibitor of Notch-dependent transcription.63 In this case as well, IKKα could de-repress Notch-dependent transcription, presumably by phosphorylating IκBα (Figure 1b). Thus, it appears that at least in some contexts, stimuli that activate NF-κB also lead to Notch activation and inhibitors of NF-κB also inhibit Notch-dependent transcription.
Undefined Mechanisms
Knockdown of Notch-1 in Notch-1 antisense transgenic mice leads to impairment in long-term potentiation in hippocampal neurons, the neurobiological mechanism for long-term memory. NF-κB DNA-binding activity was found to be defective in Notch-1-deficient neurons.64 The mechanism(s) through which Notch-1 stimulates NF-κB activity in hippocampal neurons remain undetermined. Similarly, in pancreatic carcinoma cells, inhibition of Notch signaling is accompanied by decreased NF-κB activity, through mechanisms that are still to be clarified.65, 66
CONCLUSIONS AND FUTURE DIRECTIONS
The observations reported in the literature so far offer a complex and incomplete picture of the interactions between these two key cell fate determining pathways. As it is becoming increasingly clear in the case of other pathways, these interactions can be cooperative or antagonistic, and multiple levels of feedback are possible depending on the context. The physiological relevance of these interactions will need to be thoroughly investigated. However, it can be safely stated that those planning to manipulate the Notch-signaling pathway for experimental or therapeutic purposes would do well to examine the possible effects on NF-κB and vice versa. This could have implications in several fields.
In cancer biology, both Notch and NF-κB are prominent therapeutic targets. If murine and in vitro data are confirmed in human disease, the treatment of cancers dependent on Notch activity may benefit from combinations of agents targeting both pathways, for example, inhibiting Notch and IKK activities, or Notch and the proteasome. Evidence in support of this notion is offered by the fact that some cancers in which Notch plays a clearly oncogenic role, such as breast and pancreatic carcinomas, are also often characterized by high NF-κB activity. Ramdass et al67 showed that in a large series of cervical cancer specimens, the NF-κB and Notch pathways were commonly coactivated, as judged by expression of Notch target and NF-κB target genes, and nuclear localization of NF-κB immunoreactivity.
In immunology and inflammation, where the role of NF-κB is paramount, Notch-1 and Notch-3 have been suggested to regulate NF-κB activity during mature T-cell activation (Notch-1) and during thymocyte development (Notch-3). It is conceivable that combined manipulations of the two pathways may be useful in a number of indications, from graft versus host disease to transplant rejection.
In neurology, the possible role of Notch upstream of NF-κB in hippocampal neurons64 needs to be further investigated as a possible basis for neuronal plasticity in long-term memory. These studies may suggest possible toxic effects of long-term Notch inhibition in the brain, and contribute to our understanding of one of the most fundamental processes in the human mind.
The cross talk between Notch and NF-κB is a biological Pandora's box that has just been opened. Sorting out the mechanisms, biological implications, physiological significance and possible therapeutic applications of the multiple interactions between these two pathways will require considerable effort over the next several years, but is likely to generate invaluable information in a number of biological and medical fields.
References
Miele L . Notch signaling. Clin Cancer Res 2006;12:1074–1079.
Miele L, Golde T, Osborne B . Notch signaling in cancer. Curr Mol Med 2006;6:905–918.
Berman JN, Look AT . Targeting transcription factors in acute leukemia in children. Curr Drug Targets 2007;8:727–737.
Tanigaki K, Honjo T . Regulation of lymphocyte development by Notch signaling. Nat Immunol 2007;8:451–456.
Bolos V, Grego-Bessa J, de la Pompa JL . Notch signaling in development and cancer. Endocr Rev 2007;28:339–363.
Roy M, Pear WS, Aster JC . The multifaceted role of Notch in cancer. Curr Opin Genet Dev 2007;17:52–59.
Lai EC . Notch signaling: control of cell communication and cell fate. Development 2004;131:965–973.
Sanchez-Irizarry C, Carpenter AC, Weng AP, et al. Notch subunit heterodimerization and prevention of ligand-independent proteolytic activation depend, respectively, on a novel domain and the LNR repeats. Mol Cell Biol 2004;24:9265–9273.
Gordon WR, Vardar-Ulu D, Histen G, et al. Structural basis for autoinhibition of Notch. Nat Struct Mol Biol 2007;14:295–300.
Wang W, Struhl G . Dorsophila Epsin mediates a select endocytic pathway that DSL ligands must enter to activate Notch. Development 2004;131:5367–5380.
Kopan R, Ilagan MX . Gamma-secretase: proteasome of the membrane? Nat Rev Mol Cell Biol 2004;5:499–504.
Artavanis-Tsakonas S, Rand MD, Lake RJ . Notch signaling: cell fate control and signal integration in development. Science 1999;284:770–776.
Rangarajan A, Talora C, Okuyama R, et al. Notch signaling is a direct determinant of keratinocyte growth arrest and entry into differentiation. EMBO J 2001;20:3427–3436.
Ronchini C, Capobianco AJ . Induction of cyclin D1 transcription and CDK2 activity by Notch(ic): implication for cell cycle disruption in transformation by Notch(ic). Mol Cell Biol 22001;1:5925–5934.
Baonza A, Freeman M . Control of cell proliferation in the Drosophila eye by Notch signaling. Dev Cell 2005;8:529–539.
Cheng P, Zlobin A, Volgina V, et al. Notch-1 regulates NF-kappaB activity in hemopoietic progenitor cells. J Immunol 2001;167:4458–4467.
Garces C, Ruiz-Hidalgo MJ, Font de Mora J, et al. Notch-1 controls the expression of fatty acid-activated transcription factors and is required for adipogenesis. J Biol Chem 1997;272:29729–29734.
Nickoloff BJ, Qin JZ, Chaturvedi V, et al. Jagged-1 mediated activation of notch signaling induces complete maturation of human keratinocytes through NF-kappaB and PPARgamma. Cell Death Differ 2002;9:842–855.
Sarmento LM, Huang H, Limon A, et al. Notch1 modulates timing of G1-S progression by inducing SKP2 transcription and p27 Kip1 degradation. J Exp Med 2005;202:157–168.
Klinakis A, Szabolcs M, Politi K, et al. Myc is a Notch1 transcriptional target and a requisite for Notch1-induced mammary tumorigenesis in mice. Proc Natl Acad Sci USA 2006;103:9262–9267.
Weng AP, Millholland JM, Yashiro-Ohtani Y, et al. c-Myc is an important direct target of Notch1 in T-cell acute lymphoblastic leukemia/lymphoma. Genes Dev 2006;20:2096–2109.
Tergaonkar V, Perkins ND . p53 and NF-kappaB crosstalk: IKKalpha tips the balance. Mol Cell 2007;26:158–159.
Okamoto T, Sanda T, Asamitsu K . NF-kappa B signaling and carcinogenesis. Curr Pharm Des 2007;13:447–462.
Van Waes C . Nuclear factor-kappaB in development, prevention, and therapy of cancer. Clin Cancer Res 2007;13:1076–1082.
Inoue J, Gohda J, Akiyama T, et al. NF-kappaB activation in development and progression of cancer. Cancer Sci 2007;98:268–274.
Melisi D, Chiao PJ . NF-kappa B as a target for cancer therapy. Expert Opin Ther Targets 2007;11:133–144.
Camandola S, Mattson MP . NF-kappa B as a therapeutic target in neurodegenerative diseases. Expert Opin Ther Targets 2007;11:123–132.
O'Sullivan B, Thompson A, Thomas R . NF-kappa B as a therapeutic target in autoimmune disease. Expert Opin Ther Targets 2007;11:111–122.
Campbell KJ, Perkins ND . Regulation of NF-kappaB function. Biochem Soc Symp 2006;73:165–180.
Perkins ND . Integrating cell-signalling pathways with NF-kappaB and IKK function. Nat Rev Mol Cell Biol 2007;8:49–62.
Beg AA, Baldwin Jr AS . The I kappa B proteins: multifunctional regulators of Rel/NF-kappa B transcription factors. Genes Dev 1993;7:2064–2070.
Beg AA, Ruben SM, Scheinman RI, et al. I kappa B interacts with the nuclear localization sequences of the subunits of NF-kappa B: a mechanism for cytoplasmic retention. Genes Dev 1992;6:1899–1913.
Inoue J, Kerr LD, Rashid D, et al. Direct association of pp40/I kappa B beta with rel/NF-kappa B transcription factors: role of ankyrin repeats in the inhibition of DNA binding activity. Proc Natl Acad Sci USA 1992;89:4333–4337.
Wang C, Deng L, Hong M, et al. TAK1 is a ubiquitin-dependent kinase of MKK and IKK. Nature 2001;412:346–351.
Yamamoto Y, Verma UN, Prajapati S, et al. Histone H3 phosphorylation by IKK-alpha is critical for cytokine-induced gene expression. Nature 2003;423:655–659.
Sakurai H, Suzuki S, Kawasaki N, et al. Tumor necrosis factor-alpha-induced IKK phosphorylation of NF-kappaB p65 on serine 536 is mediated through the TRAF2, TRAF5 and TAK1 signaling pathway. J Biol Chem 22003;78:36916–36923.
Hoberg JE, Yeung F, Mayo MW . SMRT derepression by the IkappaB kinase alpha: a prerequisite to NF-kappaB transcription and survival. Mol Cell 2004;16:245–255.
Park KJ, Krishnan V, O'Malley BW, et al. Formation of an IKKalpha-dependent transcription complex is required for estrogen receptor-mediated gene activation. Mol Cell 2005;18:71–82.
Wu RC, Qin J, Hashimoto Y, et al. Regulation of SRC-3 (pCIP/ACTR/AIB-1/RAC-3/TRAM-1) coactivator activity by I kappa B kinase. Mol Cell Biol 2002;22:3549–3561.
Albanese C, Wu K, D'Amico M, et al. IKKalpha regulates mitogenic signaling through transcriptional induction of cyclin D1 via Tcf. Mol Biol Cell 2003;14:585–599.
Carayol N, Wang CY . IKKalpha stabilizes cytosolic beta-catenin by inhibiting both canonical and non-canonical degradation pathways. Cell Signal 2006;18:1941–1946.
Kwak YT, Li R, Becerra CR, et al. IkappaB kinase alpha regulates subcellular distribution and turnover of cyclin D1 by phosphorylation. J Biol Chem 2005;280:33945–33952.
Wegener E, Oeckinghaus A, Papadopoulou N, et al. Essential role for IkappaB kinase beta in remodeling Carma1-Bcl10-Malt1 complexes upon T cell activation. Mol Cell 2006;23:13–23.
Yang F, Tang E, Guan K, et al. IKK beta plays an essential role in the phosphorylation of RelA/p65 on serine 536 induced by lipopolysaccharide. J Immunol 2003;170:5630–5635.
Hu MC, Lee DF, Xia W, et al. IkappaB kinase promotes tumorigenesis through inhibition of forkhead FOXO3a. Cell 2004;117:225–237.
Gringhuis SI, Garcia-Vallejo JJ, van Het HB, et al. Convergent actions of I kappa B kinase beta and protein kinase C delta modulate mRNA stability through phosphorylation of 14-3-3 beta complexed with tristetraprolin. Mol Cell Biol 2005;25:6454–6463.
Beinke S, Robinson MJ, Hugunin M, et al. Lipopolysaccharide activation of the TPL-2/MEK/extracellular signal-regulated kinase mitogen-activated protein kinase cascade is regulated by IkappaB kinase-induced proteolysis of NF-kappaB1 p105. Mol Cell Biol 2004;24:9658–9667.
Waterfield M, Jin W, Reiley W, et al. IkappaB kinase is an essential component of the Tpl2 signaling pathway. Mol Cell Biol 2004;24:6040–6048.
Oswald F, Liptay S, Adler G, et al. NF-kappaB2 is a putative target gene of activated Notch-1 via RBP-Jkappa. Mol Cell Biol 1998;18:2077–2088.
Oakley F, Mann J, Ruddell RG, et al. Basal expression of IkappaBalpha is controlled by the mammalian transcriptional repressor RBP-J (CBF1) and its activator Notch1. J Biol Chem 2003;278:24359–24370.
Bash J, Zong WX, Banga S, et al. Rel/NF-kappaB can trigger the Notch signaling pathway by inducing the expression of Jagged1, a ligand for Notch receptors. EMBO J 1999;18:2803–2811.
Moran ST, Cariappa A, Liu H, et al. Synergism between NF-kappaB1/p50 and Notch2 during the development of marginal zone B lymphocytes. J Immunol 2007;179:195–200.
Guan E, Wang J, Laborda J, et al. T cell leukemia-associated human Notch/translocation-associated Notch homologue has I kappa B-like activity and physically interacts with nuclear factor-kappa B proteins in T cells. J Exp Med 1996;183:2025–2032.
Wang J, Shelly L, Miele L, et al. Human Notch-1 inhibits NF-kappa B activity in the nucleus through a direct interaction involving a novel domain. J Immunol 2001;167:289–295.
Palaga T, Miele L, Golde TE, et al. TCR-mediated Notch signaling regulates proliferation and IFN-gamma production in peripheral T cells. J Immunol 2003;171:3019–3024.
Shin HM, Minter LM, Cho OH, et al. Notch1 augments NF-kappaB activity by facilitating its nuclear retention. EMBO J 22006;5:129–138.
Jang MS, Miao H, Carlesso N, et al. Notch-1 regulates cell death independently of differentiation in murine erythroleukemia cells through multiple apoptosis and cell cycle pathways. J Cell Physiol 2004;199:418–433.
Bellavia D, Campese AF, Alesse E, et al. Constitutive activation of NF-kappaB and T-cell leukemia/lymphoma in Notch3 transgenic mice. EMBO J 2000;19:3337–3348.
Bellavia D, Campese AF, Checquolo S, et al. Combined expression of pTalpha and Notch3 in T cell leukemia identifies the requirement of preTCR for leukemogenesis. Proc Natl Acad Sci USA 2002;99:3788–3793.
Vacca A, Felli MP, Palermo R, et al. Notch3 and pre-TCR interaction unveils distinct NF-kappaB pathways in T-cell development and leukemia. EMBO J 2006;25:1000–1008.
Vilimas T, Mascarenhas J, Palomero T, et al. Targeting the NF-kappaB signaling pathway in Notch1-induced T-cell leukemia. Nat Med 2007;13:70–77.
Fernandez-Majada V, Aguilera C, Villanueva A, et al. Nuclear IKK activity leads to dysregulated Notch-dependent gene expression in colorectal cancer. Proc Natl Acad Sci USA 2007;104:276–281.
Aguilera C, Hoya-Arias R, Haegeman G, et al. Recruitment of IkappaBalpha to the hes1 promoter is associated with transcriptional repression. Proc Natl Acad Sci USA 2004;101:16537–16542.
Wang Y, Chan SL, Miele L, et al. Involvement of Notch signaling in hippocampal synaptic plasticity. Proc Natl Acad Sci USA 2004;101:9458–9462.
Wang Z, Banerjee S, Li Y, et al. Down-regulation of Notch-1 inhibits invasion by inactivation of nuclear factor-kappaB, vascular endothelial growth factor and matrix metalloproteinase-9 in pancreatic cancer cells. Cancer Res 2006;66:2778–2784.
Wang Z, Zhang Y, Banerjee S, et al. Inhibition of nuclear factor kappaB activity by genistein is mediated via Notch-1 signaling pathway in pancreatic cancer cells. Int J Cancer 2006;118:1930–1936.
Ramdass B, Maliekal TT, Lakshmi S, et al. Coexpression of Notch1 and NF-kappaB signaling pathway components in human cervical cancer progression. Gynecol Oncol 2006;104:352–361.
Acknowledgements
We are supported by PO1 AG025531.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Osipo, C., Golde, T., Osborne, B. et al. Off the beaten pathway: the complex cross talk between Notch and NF-κB. Lab Invest 88, 11–17 (2008). https://doi.org/10.1038/labinvest.3700700
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700700
Keywords
This article is cited by
-
Cooperative NF-κB and Notch1 signaling promotes macrophage-mediated MenaINV expression in breast cancer
Breast Cancer Research (2023)
-
Multi-objective optimization reveals time- and dose-dependent inflammatory cytokine-mediated regulation of human stem cell derived T-cell development
npj Regenerative Medicine (2022)
-
A DL-4- and TNFα-based culture system to generate high numbers of nonmodified or genetically modified immunotherapeutic human T-lymphoid progenitors
Cellular & Molecular Immunology (2021)
-
Preactivation of Notch1 in remote ischemic preconditioning reduces cerebral ischemia-reperfusion injury through crosstalk with the NF-κB pathway
Journal of Neuroinflammation (2019)
-
MiR-182 promotes cancer invasion by linking RET oncogene activated NF-κB to loss of the HES1/Notch1 regulatory circuit
Molecular Cancer (2017)