Abstract
University Health Network (UHN) Pathology, in its capacity of providing neuro-oncologic care, now utilizes a laboratory information system (LIS), which was instituted in September 2001. For the 75 years preceding the LIS, more than 50 000 pathology reports exist in paper format. High-throughput automated scanning of the paper archives was employed to add the most recent 30 years of paper records (30 000 neuropathology specimens) to the LIS. The searchable portable document format (PDF) files generated from the scans were filtered through a multi-tiered process driven by Java computer programs that selected relevant patient and diagnostic information. A second series of programs queried the neuropathologist-assigned diagnoses and successfully converted these to the standardized World Health Organization (WHO) format. This was achieved with a master list of key site and diagnostic terms, and prioritization rules that were determined on a trial and error basis. Categorization, verification, and consolidation were completed within 3 months and on a C$10 000 budget.
Similar content being viewed by others
Main
Databases of pathology specimens can have enormous benefits for clinical use and long-term population-based research. In addition to clear economizing effects, the ease of access and searchability provided by well-organized collections of pathologic cases is highly advantageous for retrospective analyses and augmenting of sample sizes to increase the power of observation.1 Various data mining methods could also be employed to extract previously unrecognized patterns within large data sets.2 While databases directed toward specific research tasks and clinical outcomes have previously been described,3, 4, 5 the electronic construction of a comprehensive pathology database from print archives has not been reported thus far. The challenge in constructing such collections lies in the organization of large amounts of data and the feasibility of digitizing paper-based records by optical character recognition (OCR). In this project, we developed a methodical and practical approach for the organization and treatment of retrospective data from print archives.
University Health Network (UHN) in Toronto provides most of the neuro-oncologic care for the southwest region of the province of Ontario. Archival material comprising paper records and corresponding glass slides and paraffin blocks for our center is available from 1932 to September 2001, at which point a laboratory information system (LIS) was implemented. Notably, the majority of the more than 50 000 records in the paper collection (up to 1993) are complemented with surgical operative reports. Here, we describe the construction of a 30 000 entry neuropathologic database from the last 30 years of paper archives at UHN and its combination with the LIS as a model for data sorting, filtration, integration, and later modification geared toward specific research tasks. Furthermore, individual entries are tagged using key diagnostic criteria to allow for greater organization within the database. The feasibility and efficacy of data annotations have been described previously.6, 7, 8
A cumulative 700 h of human and computer operation, a budget of C$10 000, OCR software, and a Java programming environment were allocated toward this project over a year-long period.
MATERIALS AND METHODS
High-throughput scanning was performed by LASON Canada Inc. (Toronto, ON, Canada), a document management company, generating searchable portable document format (PDF) files (Table 1). Full text recognition was carried out on the files using the commercially available OCR software ScanSoft OmniPage Professional 15 (Nuance Communications, Burlington, MA, USA). Subsequently, a first tier of Java programs designed at our lab distilled patient demographic (ie case number, accession date, patient name, age, gender, medical record number, and pathologist) and diagnostic information (ie clinical history, neuropathologic diagnosis, microscopic description, and comment). Standardization and filtration according to the World Health Organization (WHO)9 classification were then performed. To that end, each entry within the database was automatically tagged with key neuropathologic diagnostic and site-of-occurrence terms. All input and output data utilized by the Java programs were mediated through simple text files. The organized and filtered results were first imported into a spreadsheet, after which the data could be exported into higher level database environments depending on the research and/or clinical needs.
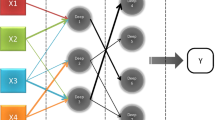
Reports obtained from the LIS were also organized and abridged in a similar manner and added to the data extracted from the scanned archives. To examine the searchability of the database and its potential in the selection of research-specific cases, we devised a sample algorithm and corresponding Java program which used the WHO diagnostic tags along with other criteria to generate a filtered list of malignant glioma and metastatic carcinoma cases (Figure 1). To protect patient confidentiality, cases were de-identified (patient names and medical record numbers were blocked).
RESULTS
From 49 326 scanned archived pages, in addition to reports extracted from the LIS (September 2001 to January 2007), a total of 29 080 cases were appended to the database. A cumulative 700 h of human and computer operation have been clocked. Of the approximately 450 h of direct human intervention, 58% of the time was allocated toward programming and systematization, while the rest was distributed between manual verification and consolidation of extracted data.
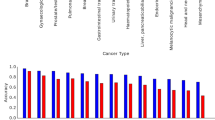
On the basis of the statistics of the scanned pages and extracted information, trends for the number of archived neurosurgical pages and neuropathology cases were compared over the period 1971–2001 (Figure 2). Although the pathology specimens show a general rise in number, the pattern in paper volume is unremarkable. The steady decrease in the number of archived pages after 1985 is largely due to the gradual omission of operative records from the archive. Furthermore, a chart depicting the distribution of neuropathologic diagnoses contained in the database is provided (Figure 3).
Possible errors in the digitization of records necessitated a manual cross-check of the extracted data (Table 1). This proved to be a rate-limiting step. Verification of dates, names, and diagnoses against scanned paper records required allocation of approximately 1 h for every 150 cases or 40% of the total human time input.
DISCUSSION
We have demonstrated a novel and highly economical method for the creation of a fully searchable database of pathology records from paper archives with a minimum of human input and correction time. The process does not require application in contiguous intervals, and as such can be utilized on multiple platforms. Access to the data collection can provide many opportunities for epidemiologic and retrospective research. Reviews on specific central nervous system neoplasms and the corresponding neurosurgical approaches are among a number of current studies made possible on account of the database. Correlation of tumors with demographic data such as age and gender, and analysis of the associated changes over time in such relations have also been undertaken.
The manual verification process was the most time-consuming stage. Breaking down the task into smaller segments with a more efficient redundancy function could have improved the outcome. The relatively low quality of some older documents might have been the main contributing factor to the poor OCR output of the corresponding reports. Scanning was performed at 200 dots per inch (DPI), but could be extended to 600 DPI without much compromise in scanning speed. A larger budget could allow for the use of digital camera equipment, which greatly increases the quality and speed of the scans without the necessity of cutting the paper volumes.10 However, since the quality of the 200 DPI scans and the format of reports might vary in a predictable fashion, more complex programs with learnability functions could be devised to maintain the low cost of the scanning process. This could allow for the automatic flagging of recurrent errors. The feasibility of this approach is currently under investigation. Alternatively, teachable programs could automate the manual checking of the digitized data following expert supervision on the initial runs.
In addition to the database applications currently in use, a multitude of other research tasks await assessment. The importance of pathological classifications and the introduction of immunohistochemical techniques could be evaluated from both a medical and economical point of view. Furthermore, the availability of surgical operative reports for the majority of cases in our collection can enhance the study of surgical methods. Linkage of the neuropathology database to other clinical data banks such as cancer registries can allow for more comprehensive epidemiologic studies. Additionally, linkage with tumor banks may enable the annotation of the database with genomic information. Higher level database environments can allow for such interconnections among databases in multiple institutions, and thereby facilitate the analysis of familial syndromes and cooccurrence of specific malignancies from multiple dimensions. Moreover, the database environment could be adapted if necessary to optimally suit the requirements of the researcher. Finally, to sustain the high research potential provided by the database, an efficient mechanism for constant addition of new cases and ongoing revision of previous records based on updated diagnostic information is imperative.
References
Schmidt R, Simmons K, Grimm E, et al. Integration of scanned document management with the anatomic pathology laboratory information system: analysis of benefits. Am J Clin Pathol 2006;126:678–683.
Delen D, Walker G, Kadam A . Predicting breast cancer survivability: a comparison of three data mining methods. Artif Intell Med 2005;34:113–127.
Manley S, Mucci N, De Marzo A, et al. Relational database structure to manage high-density tissue microarray data and images for pathology studies focusing on clinical outcome. Am J Pathol 2001;159:837–843.
Naf D, Krupke D, Sundberg J, et al. The mouse tumor biology database: a public resource for cancer genetics and pathology of the mouse. Cancer Res 2002;62:1235–1240.
Patel A, Gupta D, Seligson D, et al. Availability and quality of paraffin blocks identified in pathology archives: a multi-institutional study by the Shared Pathology Informatics Network (SPIN). BMC Cancer 2007;7:37.
Liu K, Mitchell K, Chapman W, et al. Automating tissue bank annotation from pathology reports—comparison to a gold standard expert annotation set. AMIA Annu Symp Proc 2005; 460–464.
Mitchell K, Becich M, Berman J, et al. Implementation and evaluation of a negation tagger in a pipeline-based system for information extract from pathology reports. Medinfo 2004;11 (Part 1):663–667.
Berman J . Automatic extraction of candidate nomenclature terms using the doublet method. BMC Med Inform Decis Mak 2005;5:35.
World Health Organization classification of tumours. Pathology and Genetics of Tumours of the Nervous System. IARC Press: Lyon, 2000.
America AN . BookDrive DIY Scanner. (cited September 16, 2007); Available from http://www.atiz.com/Download/brochure_diy.pdf.
Acknowledgements
Grants from the Department of Laboratory Medicine and Pathobiology, University of Toronto and Division of Neurosurgery, University Health Network.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/duality of interest
The authors have no duality of interest to declare.
Rights and permissions
About this article
Cite this article
Ehsani, S., Kiehl, TR., Bernstein, A. et al. Creation of a retrospective searchable neuropathologic database from print archives at Toronto's University Health Network. Lab Invest 88, 89–93 (2008). https://doi.org/10.1038/labinvest.3700694
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700694