Abstract
Estrogen-receptor status provides a major biomarker in breast cancer classification and has an important impact on prognosis and treatment options. The aim of this study was to investigate peptide profiles of invasive breast cancer with positive (n=39) and negative receptor status (n=41). Peptide profiles were generated by ‘Differential Peptide Display’, which is an offline-coupled combination of reversed-phase-HPLC and MALDI mass spectrometry. Mass spectrometric data were correlated with the immunohistochemically determined receptor state. Identification of peptides of interest was carried out by additional mass spectrometric methods (eg MALDI-TOF-TOF-MS-MS). Approximately 3000–7000 signals were detected per sample and thymosin alpha-1, an asparaginyl endopeptidase generated cleavage product of the ubiquitous acidic protein prothymosin-alpha, was found to differentiate the tumor samples according to their receptor status with the highest specificity. The concentration of Thymosin alpha-1 was found to be upregulated (n=37) in estrogen-negative cancer samples and downregulated (n=32) in estrogen-positive breast cancer samples. The expression of the precursor protein (Prothymosin-alpha) has been discussed previously as a prognostic factor in breast cancer. It is involved in the ER signal transduction pathway as an anti-coactivator-inhibitor. From our findings we conclude that Thymosin alpha-1 could serve as a surrogate marker in breast cancers and may indicate ER functionality.
Similar content being viewed by others
Main
In the Western world, breast cancer is the most prevalent cancer in women and the second leading cause of cancer death. In the past two decades, the molecular dissection of cancer has increased our understanding of the pathways that are altered in neoplastic cells and that have a growing impact in oncologic therapy. Complex genetic and proteomic alterations induce cancer growth and progression resulting in significant clinical heterogeneity among individual tumors and individual prognosis.
According to the College of American Pathologists, expression of estrogen receptor-alpha (ER) and progesterone receptor (PgR) belongs to the first category of prognostic factors besides tumor size, histological grade and lymph node status. The ER mediates the stimulatory effects of estrogens and the inhibitory effects of anti-estrogens in breast cancer and other hormone-dependent tissues. For patients with hormone-receptor (HR)-positive breast cancer (∼75% ductal invasive carcinoma), endocrine therapy (eg Tamoxifen) is the preferred treatment. Approximately 50–60% of patients with ER-positive tumors benefit from endocrine therapy.1 The ER expression is gradually lost during tumor progression, leading to a more undifferentiated state and so efficacy of endocrine therapy is limited. Panendocrine resistance2 is almost universally in breast cancer negative for ER and PgR content but also occurs in a significant fraction of breast cancer cases that express HR. PgR expression is ER-dependent and the presence of PgR in the breast indicates probable responsiveness to endocrine therapy.3, 4 The underlying mechanism of endocrine resistance is still obscure. Potentially critical links between the ER and cell-cycle or cell-survival pathways are disrupted, inducing usage of alternative pathways to serve functions previously subject to master regulation by the ER.
In recent years, peptidomic techniques have evolved, which allow the comprehensive analysis of native peptides in virtually every biological source. In this study, we applied a peptidomic technique called differential peptide display (DPD) to compare ER-positive and ER-negative tumor tissues (Table 1) in order to reveal specific profiles.
DPD5, 6, 7, 8 is a peptidomic technique to generate comprehensive peptide displays of usually >1500 peptides from a variety of biological samples including blood plasma, urine, cerebrospinal fluid, cell and tissue extracts or cell culture supernatants. For clarification, the size we refer to for the term ‘peptides’ covers a mass range of 750–15 000 Da.
Materials and methods
Tissue Samples
Freshly frozen as well as formalin-fixed, paraffin-embedded tissue of 80 cases of invasive ductal carcinomas were retrieved from the archive of the Institute of Pathology, Hannover Medical School. The samples were collected according to the guidelines of the local ethics committee.
Immunohistochemistry
Sections measuring 2 μm of carcinoma tissue were de-waxed by xylene, rehydrated by graded alcohol and epitope retrieval was carried out in a microwave oven (20 min; 100°C in 10 mM sodium citrate; pH 6.0). Sections were stained using the Shandon coverplate system in a Tecan Genesis Autostainer (ThermoShandon, Frankfurt a.M., Germany; Tecan, Deisenhofen, Germany). Tissue peroxidase activity was blocked by incubation with 3% hydrogen peroxide for 8 min. The primary antibodies against PgR (PgR 636), Her2neu (c-erbB-2), Ki67 (MIB-1) (all from Dako, Germany), p53 (DO-7) and for ER (6F11) (both from Nova Castra, Germany) were used. Biotinylated secondary antibodies were used for the catalyzed signal amplification technique (CSA, Dako). New fuchsin served as chromogen, and hemalaun served as counterstain.
Differential Peptide Display
Peptide extraction
Pro analysi guanidinium chloride was obtained from Merck KGaA (Darmstadt, Germany). Pro analysi glycine was obtained from Carl Roth GbmH & Co (Karlsruhe, Germany). Microcon centrifugal filter units with an MWCO of 50 000 were purchased from Millipore (Bedford, MA, USA). A measure of 10 mg of each frozen sample was solubilized in extraction buffer (6 M guanidinium chloride, 50 mM glycine, pH 2.6). The protein/peptide extraction was carried out at 99°C and 1100 r.p.m. in a thermomixer. After incubating for 10 min, the tubes were centrifuged at 18 000 g for 30 min at 10°C. The supernatants were transferred to the ultra centrifugal filter units and centrifuged (Z 233 MK-2, HERMLE Labortechnik GmbH, Wehingen, Germany) at 12 000 × g in a fixed angle rotor for 30 min.
Peptide Display
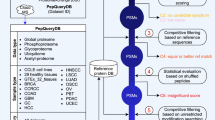
The methodology has been described by Schulte et al,8 Tammen et al,9 and Rai et al10 in detail. Briefly, peptides are extracted from biological sources and fractionated by means of reversed phase chromatography. Each fraction is subjected to MALDI-TOF-MS and the mass spectra of all 96 fractions are combined, resulting in a two-dimensional display of peptide masses named ‘Peptide Display’, where the abscissa displays the mass-to-charge ratio and the ordinate is determined by the retention time on the RP-HPLC and the signal intensity is depicted by the color saturation. Specific software (Spectromania®) has been designed to allow the visualization, processing and analysis of mass spectra.11 For the identification of peptides, peaks from individual HPLC fractions can be subjected to tandem MS identification by means of nESI-qTOF-MS sequencing or MALDI-TOF-TOF-MS resulting in peptide fragment spectra. These spectra serve to identify the corresponding peptide sequence by remote database searching.
Sample size and statistical data analysis
For this study, more than 200 peptide displays were generated from 80 different samples. Each display consisted of 96 MS spectra. Each sample was measured at least twice either on the STR-Platform or on the ABI4700 TOF-TOF analyzer. Detection of differentially expressed peptides was achieved by calculation of subtractive peptide displays or correlation analysis (DPD) by referring to mass spectrometric data. In addition, all mass spectra with the same fraction number were averaged, assuming that they contain an identical subset of peptides, yielding an average peptide display. The averaged spectra of this display served as a template for the definition of those peak coordinates, which can be clearly distinguished from baseline noise. Signal intensities of all peaks derived from each group were exported for comprehensive correlation data analysis with commercially available software. For quality assurance, each sample was spiked with a defined peptide mixture (500 fmol per sample).
Peptide Identification
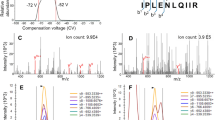
For MALDI-TOF/TOF-MS/MS identifications the archived MALDI target plates from the peptide display process were used. High accuracy survey MALDI-TOF-MS spectra were acquired in a fully automated fashion in the reflector mode of a 4700 proteomics analyzer (Applied Biosystems, Framingham, USA). All measurements were calibrated using a daily updated default calibration. Peptides were subsequently fragmented in the MS/MS mode using a collision energy of 1 keV and an intermediate gas pressure of approximately 3 × 10–7 Torr of ambient air (Figure 1). MS/MS spectra were noise filtered, peak de-isotoped and transferred into the MASCOT generic file format using the MascotDistiller software tool (Matrix Science Ltd, UK). Cascading searches on a MASCOT database engine (Matrix Science) including several post-translational modifications were performed by the Mascot Daemon software (Version 1.8 or higher) against the Swiss-Prot (Version 39 or higher, GeneBio, Geneva, Switzerland) database.
MALDI-TOF/TOF-MS/MS spectrum of PTMA_HUMAN 1–28. Peaks depicting identified fragment ions are labeled according to the standard fragment ion nomenclature. The peptide identification score (probability based MOWSE score using the MASCOT database engine) for this particular spectrum was highly significant (MOWSE score=75).
Western-blot analysis
Frozen tissue homogenates were denatured in SDS–PAGE sample buffer at 95°C for 5 min twice. A measure of 15 μg of protein (measured by Bradford assay) of each sample was analyzed by SDS–PAGE (Tris-HCl Gels 10–20%, Bio-Rad, Germany), transferred to 0.45 μm Nytran SPC (Schleicher und Schüll, Germany), and subjected to Western-blot analysis. The membrane was incubated with the primary antibody 2F11 (Alexis, Germany) (diluted 1:100) for 1 h at RT. After washing with PBS, the peroxidase-conjugated second antibody was added and the protein was visualized by chemiluminescence detection.
Results
In a first approach, samples were prepared in two different concentrations (3 and 7 mg wet weight equivalent) and subjected to peptide profiling by DPD (Figure 2 and 3). Every data point (approximately 1.4 million per display) of every display was then correlated to the receptor state. This analysis revealed that a peptide with m/z ratio of 3108 exhibited the highest correlation value. This signal could be identified as thymosin alpha-1 (prothymosin-alpha 1–28 with an acetylated N-terminus (N-acetylserine), THYA) by MALDI-TOF/TOF-MS-MS. For further characterization of the samples, an overview sequencing approach was applied (insert Table Figure 2). The analysis revealed extracellular as well as intracellular peptides.
Peptide displays from tumor specimens (mean display of ER-negative and ER-positive tumors). The x-axis displays the mass-to-charge ratio, the y-axis depicts the retention time of the peptides on the RP-HPLC. The signal intensity is indicated by color saturation. Between 3000 and 7000 signals are visualized per peptide display. Sequenced peptides are marked with Arabic numerals, which refer to the inserted table. Peptides no. 22 and 24 were added prior to processing of the samples for quality control purposes. Insert: The table depicts the precursor name, the putative origin of the peptide and the Swiss Prot accession number.
Mass spectrometric detection of THYA. The figure depicts a distinct region of peptide displays from ER-negative and ER-positive tumor samples. Each lane separates two different wet weight equivalents of tumor tissue. The signal of THYA is marked. Additionally, as a reference a signal corresponding to Thymosin beta 4 sulphoxid is tagged.
To validate this first result, 60 samples were again obtained from the Institute of Pathology. A total of 50% of the samples were blinded for the primary investigator. Blinded samples and samples with known receptor state were investigated especially for the occurrence of Thymosin alpha-1 and measured at least twice. Based on the intensity levels of Thymosin alpha-1 and visual inspection of each spectrum, the blinded samples were assigned to the putative receptor state. After assignment, the results were transferred back to the Institute of Pathology, logged and the results of the IHC analysis were disclosed. After disclosure of the IHC results the assignment was found to be correct in 26 out of 30 cases. Figures 4 and 5 show the intensity levels of THYA in correlation to the receptor state.
The figure shows a zoom view representing a distinct chromatographic area and m/z region (x-axis) from peptide displays. Lanes of peptide displays of independent experimental groups are shown (separated by solid lines). The outcome of the IHC (separated by dashed lines) is depicted on the right side. THYA (arrow) is highly upregulated in ER-negative tumor samples.
Intensity levels of THYA. Each graph depicts the signal intensities of THYA in different panels (a–d). Panel a corresponds to the discovery set. Panel b contains redundant measurements (eight ER-positive cases and 13 ER-negative cases). Panel c represents MALDI-TOF-TOF data and Panel e corresponds to samples that show a positive correlation between positive estrogen receptor status and occurrence of THYA. Open circles=ER negative samples, solid squares=ER positive samples.
The detailed data analysis assign 69 samples to be in accordance to the IHC analysis, which means that positive correlation is seen between the occurrence of THYA and negative estrogen receptor state. In seven ER-positive breast cancer specimens, THYA was detectable and in four ER-negative samples THYA was not detectable. These 11 samples were reanalyzed including de novo sampling and IHC analysis (Figure 5, panel d) and findings were confirmed.
To validate the results Western blots for THYA were performed. In most cases, the Western blot was not successful, resulting in non-detection of THYA and its precursor. In successful attempts two bands were displayed, corresponding to Prothymosin-alpha (PTa) and THYA in ER-negative breast carcinomas. In ER-positive breast carcinomas, exclusively, the precursor was detectable (Figure 6).
Discussion
The overview (sequencing) approach shed light on a common problem in tumor sample composition, namely the tumor heterogeneity. Among the extracted peptides, not only the tumor-associated peptides as Vimentin or Cytokeratin 8 were identified, but also peptides with origin outside of tumor cells were sequenced. Multiple peptides derived from hemoglobin as indicators of blood contamination. For this reason, it is insufficient to use pattern recognition. Sequencing and identifying peptides and protein associated with discriminating peaks obviate the heterogeneity problem and will improve biomarker reproducibility.
Gene transcriptional activation by ER is enhanced or repressed by interaction with regulatory factors, which function in a positive fashion (coactivators) or negative fashion (corepressor) and are believed to be interposed between the receptor and the basal transcription complex. These co-regulators exist as a part of large complexes that can be recruited by the ER and function as chromatin remodeling factors. For that reason, this tripartite action of the ER, involving the receptor, its ligands and co-regulator proteins, allows the precise regulation of the biological effects of these HRs on gene expression.
The DPD analysis of ER-positive and ER-negative breast cancer tissue samples revealed THYA as the peptide with the highest power to differentiate between the two groups. THYA is upregulated in ER-negative breast cancer tissue. A broad spread in THYA signal expression from the same sample during independent analysis rounds can be seen. This may have multiple reasons: heterogeneity of tumors, insufficient extraction of peptides, loss of material during processing or sample mix-up. To curate such errors multiple measurements were performed and all data, even with confounding results, are shown. Nevertheless, the data show rather good robustness and high reproducibility. On the other hand, the validation by Western blotting was not reproducible and error prone. It is established that highly conserved polypeptides like PTa and THYA are only poorly immunogenic. Furthermore, these small molecules hardly bind to membranes used routinely for Western blots.12 These difficulties become even more noticeable when the literature is screened for optimized detection methods. Only a few reports demonstrate immunogenic detection of PTa. However, after optimizing the conditions as described by Rodriguez et al,13 in some cases a separation and blotting of THYA and PTa (the precursor) was possible.
A link between PTa and breast cancer was first published in 1993.14 It is a highly acidic protein present in virtually every cell.15 Its high conservation in mammals and wide tissue distribution suggests an essential biological role and several descriptions16, 17, 18 point to its involvement in cell cycle. PTa expression, measured via radio immunoassay, is higher in tumor samples compared to normal breast tissue, and the expression levels of PTa correlate with the proliferation status and metastatic potential of tumors.19 The group of Katzenellenbogen published results describing the interaction of a repressor of estrogen activity (REA) with the liganded ER. REA suppresses ER-activated gene transcription by estrogens, and competes with coactivators for binding to ER.20, 21, 22 PTa was identified as a binding protein partner of REA and by binding to REA selectively activates ER. This interaction of PTa with REA enables coactivator interaction with ER, thus resulting in enhanced ER transcriptional activity. Furthermore, it was found that gene expression of PTa is rapidly increased by estrogens, highlighting a new role for PTa and REA as ER activity-modulating proteins. The repressor REA20, 21, 22 is inhibited by binding of PTa. Thus, PTa can be regarded as an anti-coactivator-inhibitor.
Thymosin alpha-1, a truncated N-terminal form of PTa is generated at least partially, by the lysosomal asparaginyl endopeptidase legumain.23 Breast cancer tissues are found to be positive for legumain,24 but so far there are no studies that have determined the legumain activity or expression levels in breast cancers in correlation to ER receptor state. Bianco and co-workers25 reported that increased intracellular PTa expression in the presence of estrogens is accompanied by increased nuclear and decreased cytoplasmic PTa localization if estrogens are present. As estrogen-dependent nuclear translocation in ER-negative tumors is diminished, PTa may be more susceptible to lysosomal processing by legumain in ER-negative breast cancers. This assumption may explain the processing of PTa leading to the occurrence of THYA in ER-negative tumor samples. This is supported by Sakabe et al26 who reported that natural and environmental estrogens modulate the production of THYA in ER-positive TECs (thymus epithelial cells) in a negative fashion. Thus, the emersion of THYA may be a result of the dysfunction and not of the expression of the ER.
The analysis of the clinical data revealed an accumulation of patients with negative progesterone status (Table 2) in the subgroup of patients with positive ER status and THYA, which is indicative of dysfunctional ER. Therefore, we conclude that the detection of THYA in estrogen-positive cases may indicate steroid-insensitive breast cancers or breast cancers with compromised response to regular steroid stimulation. Consequently, the analysis of breast tumor tissue for these specific molecules may be useful for stratification of patients who are likely to profit from anti-estrogen therapies or are in need of a more intensive therapy.
As peptides appear at the end of metabolic pathways, they are likely to subsume genetic changes as well as changes in protein expression, protein half-life and biochemical pathways. The half-life times of peptides is often increased as compared to proteins, as the number of sites for proteolytic degradation is already drastically reduced. In addition, peptides usually have higher permeability between tissue compartments and therefore the chance of detecting peptides apart from the tissue itself (eg in blood) is higher compared to proteins and nucleic acids. Data from previous studies27 suggest that truncated forms of PTa are capable of migrating from cells into the peripheral blood compartment. Therefore, we are currently conducting studies to determine whether THYA is detectable at different levels in plasma of breast cancer patients in correlation to the ER state. The determination must be specific for THYA as PTa may cross-react in an immunoassay. The level of THYA may indicate ER-negative breast cancer or the conversion to hormone-insensitive breast cancer during anti-estrogen (eg Tamoxifen) therapy.
Note
Peptidomics, Differential Peptide Display, Spectromania and BioVisioN are registered trademarks of BioVisioN AG, Hannover, Germany.
References
Osborne CK . Steroid hormone receptors in breast cancer management. Breast Cancer Res Treat 1998;51:227–238.
Ellis M . Overcoming endocrine therapy resistance by signal transduction inhibition. Oncologist 2004;9 (Suppl 3):20–26.
McGuire WL . Hormone receptors: their role in predicting prognosis and response to endocrine therapy. Semin Oncol 1978;5:428–433.
Clarke RB, Anderson E, Howell A . Steroid receptors in human breast cancer. Trends Endocrinol Metab 2004;15:316–323.
Schulz-Knappe P, Zucht HD, Heine G, et al. Peptidomics: the comprehensive analysis of peptides in complex biological mixtures. Comb Chem High Throughput Screen 2001;4:207–217.
Tammen H, Mohring T, Kellmann M, et al. Mass spectrometric phenotyping of Val34Leu polymorphism of blood coagulation factor XIII by differential peptide display. Clin Chem 2004;50:545–551.
Tammen H, Kreipe H, Hess R, et al. Expression profiling of breast cancer cells by differential peptide display. Breast Cancer Res Treat 2003;79:83–93.
Schulte I, Tammen H, Selle H, et al. Peptides in body fluids and tissues as markers of disease. Expert Rev Mol Diagn 2005;5:145–157.
Tammen H, Schulte I, Hess R, et al. Peptidomic analysis of human blood specimens: comparison between plasma specimens and serum by differential peptide display. Proteomics 2005;5:3362–3377.
Rai AJ, Gelfand CA, Haywood BC, et al. HUPO Plasma Proteome Project specimen collection and handling: towards the standardization of parameters for plasma proteome samples. Proteomics 2005;5:3262–3277.
Lamerz J, Selle H, Scapozza L, et al. Correlation-associated peptide networks of human cerebrospinal fluid. Proteomics 2005;5:2789–2798.
Klimentzou P, Paravatou-Petsotas M, Zikos C, et al. Development and immunochemical evaluation of antibodies Y for the poorly immunogenic polypeptide prothymosin alpha. Peptides 2006;5:183–193.
Rodriguez P, Vinuela JE, Alvarez-Fernandez L, et al. Overexpression of prothymosin alpha accelerates proliferation and retards differentiation in HL-60 cells. Biochem J 1998;331 (Part 3):753–761.
Tsitsiloni OE, Stiakakis J, Koutselinis A, et al. Expression of alpha-thymosins in human tissues in normal and abnormal growth. Proc Natl Acad Sci USA 1993;90:9504–9507.
Hannappel E, Huff T . The thymosins. Prothymosin alpha, parathymosin, and beta-thymosins: structure and function. Vitam Horm 2003;66:257–296.
Pineiro A, Cordero OJ, Nogueira M . Fifteen years of prothymosin alpha: contradictory past and new horizons. Peptides 2000;21:1433–1446.
Papamarcaki T, Tsolas O . Prothymosin alpha binds to histone H1 in vitro. FEBS Lett 1994;345:71–75.
Karetsou Z, Sandaltzopoulos R, Frangou-Lazaridis M, et al. Prothymosin alpha modulates the interaction of histone H1 with chromatin. Nucleic Acids Res 1998;26:3111–3118.
Tsitsilonis OE, Bekris E, Voutsas IF, et al. The prognostic value of alpha-thymosins in breast cancer. Anticancer Res 1998;18:1501–1508.
Delage-Mourroux R, Martini PG, Choi I, et al. Analysis of estrogen receptor interaction with a repressor of estrogen receptor activity (REA) and the regulation of estrogen receptor transcriptional activity by REA. J Biol Chem 2000;275:35848–35856.
Martini PG, Delage-Mourroux R, Kraichely DM, et al. Prothymosin alpha selectively enhances estrogen receptor transcriptional activity by interacting with a repressor of estrogen receptor activity. Mol Cell Biol 2000;20:6224–6232.
Montano MM, Ekena K, Delage-Mourroux R, et al. An estrogen receptor-selective coregulator that potentiates the effectiveness of antiestrogens and represses the activity of estrogens. Proc Natl Acad Sci USA 1999;96:6947–6952.
Chen JM, Dando PM, Rawlings ND, et al. Cloning, isolation, and characterization of mammalian legumain, an asparaginyl endopeptidase. J Biol Chem 1997;272:8090–8098.
Liu C, Sun C, Huang H, et al. Overexpression of legumain in tumors is significant for invasion/metastasis and a candidate enzymatic target for prodrug therapy. Cancer Res 2003;63:2957–2964.
Bianco NR, Montano MM . Regulation of prothymosin alpha by estrogen receptor alpha: molecular mechanisms and relevance in estrogen-mediated breast cell growth. Oncogene 2002;21:5233–5244.
Sakabe K, Okuma M, Karaki S, et al. Inhibitory effect of natural and environmental estrogens on thymic hormone production in thymus epithelial cell culture. Int J Immunopharmacol 1999;21:861–868.
Tammen H, Schorn K, Selle H, et al. Identification of peptide markers in a tumor graft model in immunodeficient mice. Comb Chem High Throughput Screen 2005;8:783–788.
Acknowledgements
This work was supported by Grant No. 203.19-32329-5-406 (Federal State of Lower Saxony).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Traub, F., Jost, M., Hess, R. et al. Peptidomic analysis of breast cancer reveals a putative surrogate marker for estrogen receptor-negative carcinomas. Lab Invest 86, 246–253 (2006). https://doi.org/10.1038/labinvest.3700385
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700385
Keywords
This article is cited by
-
Breast cancer proteomics: a review for clinicians
Journal of Cancer Research and Clinical Oncology (2011)
-
Legumain expression as a prognostic factor in breast cancer patients
Breast Cancer Research and Treatment (2007)
-
Peptidomic analysis of human peripheral monocytes persistently infected by Chlamydia trachomatis
Medical Microbiology and Immunology (2007)
-
The blood peptidome: a higher dimension of information content for cancer biomarker discovery
Nature Reviews Cancer (2006)