Abstract
Malignant gliomas (astrocytomas) are lethal tumors that invade the brain. Invasive cell migration is initiated by extension of pseudopodia into interstitial spaces. In this study, U87 glioma cells formed pseudopodia in vitro as cells pushed through 3 μm pores of polycarbonate membranes. Harvesting pseudopodia in a novel two-step method provided material for proteomic analysis. Differences in the protein profiles of pseudopodia and whole cells were found using differential gel electrophoresis (DIGE) and immunoblotting. Proteins from two-dimensional (2D) gels with MR's of 20–100 kDa and pI's of 3.0–10.0 were identified by peptide mass fingerprinting analysis using mass spectrometry. For DIGE, lysates of pseudopodia and whole cells were each labeled with electrophilic forms of fluorescent dyes, Cy3 or Cy5, and analyzed as mixtures. Analysis was repeated with reciprocal labeling. Differences in protein distributions were detected by manual inspection and computer analysis. Topographical digital maps of the scanned gels were used for algorithmic spot matching, normalization of background, quantifying spot differences, and elimination of artifacts. Pseudopodial proteins in Coomassie-stained 2D gels included isoforms of glycolytic enzymes as the largest group, seven of 24 proteins. Peptide mass fingerprint analysis of DIGE gels demonstrated increased isoforms of annexin (Anx) I, AnxII, enolase, pyruvate kinase, and aldolase, and decreased mitochondrial manganese superoxide dismutase and transketolase in pseudopodia. Specific antibodies showed restricted immunoreactivity of the hepatocyte growth factor (HGF) α chain to pseudopodia, indicating localization of its active form. Met (the HGF receptor), actin, and total AnxI were increased in pseudopodial lysates on immunoblots. Increased constituents of the pseudopodial proteome in glioma cells, identified in this study as actin, HGF, Met, and isoforms of AnxI, AnxII, and several glycolytic enzymes, represent therapeutic targets to consider for suppression of tumor invasion.
Similar content being viewed by others
Main
Tumor cells of diffuse astrocytomas (gliomas) are invasive, regardless of tumor grade. They migrate in response to certain cytokine growth factors, such as hepatocyte growth factor (HGF), also known as scatter factor.1, 2, 3, 4 Glioma cells preferentially move through interstitial spaces formed in the white matter by axons and glial processes, possibly widened by tumor-induced edema and proteolysis. Once embryonic development of the brain is complete, most normal astrocytic cells maintain stable vascular contacts and do not migrate, with the exception of occasional, limited cell movement during some types of reactive gliosis. Normal astrocytes characteristically have a star shape due to slender cytoplasmic extensions stabilized by attachments with the vasculature and other cells. Some plasticity is retained in their fine, sheet-like perisynaptic processes that respond to neuronal activity.5 In contrast, tumor cells produce dynamic pseudopodial cell extensions that mediate migration with only transitory vascular and neuronal contacts. Signal transduction, membrane formation and turnover, cytoskeletal remodeling, adhesion, de-adhesion, and energy production occur within the pseudopodial leading edge during invasion. The proteins involved in mediating pseudopodial extension offer potential therapeutic targets to suppress tumor cell invasion.
Proteomic investigations of migrated pseudopodia rely on obtaining sufficient lysate material for study. Obtaining migrated cellular material by scraping it from wafer-thin filters or chemically releasing it from chamber well surfaces in migration assays often results in low yields. To address this issue, we used a novel two-step method to harvest migrated pseudopodia from filters. Pseudopodia were transferred from the filters onto glass slides and were then harvested in urea lysate buffer that crystallized so that all pseudopodial material could be retrieved by scraping.
A motile, human malignant glioma cell line, U87, was studied. Quantities of pseudopodial lysates were sufficient for two-dimensional (2D) gel electrophoresis followed by peptide mass fingerprint (PMF) analysis. Numerous abundant proteins detected with Coomassie blue staining were identified. Some have been previously associated with tumor progression. On differential gel electrophoresis (DIGE) analysis of whole cells and pseudopodia, several of these proteins, or some of their isoforms, demonstrated increased pseudopodial distributions. Immunoreactivity for the α chain of activated HGF was detected only in pseudopodial lysates. The HGF receptor, Met, was present in both whole cells and pseudopodia and was increased in pseudopodia. Differential localization of certain proteins, including an activated signaling network for a motility cytokine in the pseudopodia, supports this methodology for defining the functional pseudopodial proteome.
Materials and methods
Cell Culture
Human U87 MG astrocytoma cells (American Type Culture Collection, Rockville, MD, USA), were maintained in minimal essential media (MEM) Eagle (Cellgro, Mediatech, Herndon, VA, USA) with 10% fetal bovine serum (FBS) (Invitrogen, Carlsbad, CA, USA). Half volumes of media were changed the day prior to each harvest of pseudopodia to normalize their metabolic state from batch to batch.
Harvesting Pseudopodia and Whole Cells
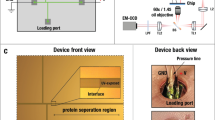
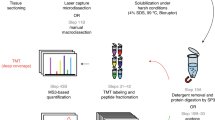
Confluent cells were trypsinized on the day of harvest and allowed to recover in 10% FBS for 2 h at 37°C. Cells were centrifuged (500 rpm, 4.5 min) and resuspended (1.5 × 106/ml) with minimal pipetting in motility media, which consisted of phosphate-buffered saline (PBS) containing 0.27% CaCl2·2H2O, 0.02% MgCl2·6H2O, 1% FBS, 5 mM sodium pyruvate, 1 mg/ml glucose, and 0.1% bovine serum albumin (BSA), unless stated otherwise. Four-well Boyden separation chambers (Neuro Probe, Gaithersburg, MD, USA) were assembled with 9 μm thick polycarbonate filters (Neuro Probe) containing 3 μm diameter pores, with and without a coating of 0.01% porcine gelatin. The filters were inserted to separate upper and lower wells. The lower wells contained motility media. Cell suspensions were added to the upper wells and each chamber was placed in a large Petri dish on a moist Kimwipe and incubated at 37°C in a 5% CO2 tissue culture incubator. To harvest pseudopodia, filters were removed at 4.5–5 h. In the first step, each filter was immersed in methanol for 15 s and placed with migrated cell materials down (attached to the undersurface of filters) on a 5 × 7.5 cm glass slide. Nonmigrated cell materials were completely wiped from the top of each filter with large Kimwipes. Fresh Kimwipes were used to press each filter firmly against a glass slide for a few seconds to promote adherence of migrated pseudopodia to the glass. The filter was then gently peeled off the slide with forceps as shown (Figure 1). If the pseudopodia did not adhere on the first attempt, the filter was resuspended in methanol a second time and the process was repeated on a new slide. The glass slides with attached pseudopodia were stored at −80°C in non-air tight plastic containers that allowed rapid freezing. During the second step, migrated materials on several glass slides were solubilized in lysate buffer consisting of 6 M urea, 4% 3-[(3-chloamidopropyl)dimethylammonio]-1-propanesulfonate, 2 M thiourea, 20 mM dithiothreitol, 1.6 mM 4-(2-hydroxyethyl)piperazine-1-ethanesulfonic acid, pH 8.0, and removed by pipetting. After allowing the residual lysate to dry and crystallize, it was retrieved with razor blades to complete the harvesting process. Pseudopodia from approximately 40–60 million cells were combined to form each batch of lysate, 1–3 mg/ml total protein. Whole cells were allowed to adhere to the filters and then placed on the glass slides, as described for pseudopodia, and harvested in a similar manner. The ratio of cell numbers used to obtain lysates of whole cells compared to pseudopodia was approximately 1:12. Most of the ‘whole’ cells were not in contact with pores that would have permitted loss of their protrusions when harvested from the filters.
Method for harvesting pseudopodia. U87 glioma cells placed in the upper wells of four-well Boyden separation chambers pushed their cytoplasm through 3 μm pores of 9 μm thick polycarbonate filters, during 4.5–5 h incubations. The chambers were disassembled and filters were removed with forceps. Following a 15 s dip in methanol, each filter was placed on a 5 × 7.5 cm glass slide with migrated materials trapped between the filter and glass. Unmigrated cell materials were wiped off the upper surface of the filter with large Kimwipes. Fresh Kimwipes were used to press the filter firmly against the underlying glass slide. The filter was then gently peeled away, leaving migrated pseudopodia attached to the slide. Urea lysis buffer was used to harvest the pseudopodia as a liquid combined with the dried residual material that was scraped off the slides following crystallization. Pseudopodia from multiple slides were batched.
Traditional 2D Gel Electrophoresis
The lysates were treated with a ReadyPrep 2D Cleanup Kit (BioRad, Hercules, CA, USA) to remove ions, DNA, RNA, etc. Their protein contents were measured using a 2D Quant kit (Amersham Biosciences, Piscataway, NJ, USA). After overnight rehydration in buffer consisting of 7 M urea, 2 M thiourea, 4% 3-[(3-chloamidopropyl)dimethylammonio]-1-propanesulfonate, 25 mM dithiothreitol, and 0.2% ampholyte, the protein samples, 75 μg, were loaded on a 17 cm immobilized pH gradient (IPG) strip (BioRad), nonlinear pH 3–10. Focusing was accomplished with 250 V for 15 min, 10 000 V for 3 h, and a third step of 60 000 total V h, and then maintained at 500 V, as needed. The resolved proteins were equilibrated in sodium dodecyl sulfate (SDS) buffer with reducing agents present and separated with SDS polyacrylamide gel electrophoresis (PAGE) using 8–16% gradient Tris-HCl gels in a Protean II apparatus (BioRad) operated at 2, 5, and 10 W per gel for 8 h. Coomassie blue stain was used to detect abundant proteins. Spots were manually retrieved with pipette tips, digested with trypsin, and submitted for PMF analysis.
DIGE
Procedures described above were followed with the following modifications. The electrophilic fluorescent dyes, PrCy3-N-hydroxysuccinimide ester and MeCy5-N-hydroxysuccinimide ester (Cy3 and Cy5, respectively), were synthesized according to a previously published method6 and stored as single-use aliquots in anhydrous N,N-dimethylformamide at −80°C. Pseudopodial and whole cell lysates were separately mixed with volumes of each dye solution to produce balanced covalent labeling of 1–5% of the available lysine side chains, without significant alteration of the protein isoelectric points (pI's). Quenching was accomplished with 40% aqueous methylamine, pH 8.6. Paired pseudopodial and whole cell lysates were mixed together for focusing and subsequent separation by SDS-PAGE. The gels were repeated with reciprocally labeled pseudopodial and whole cell lysates. The gels were viewed in a fluorescent gel imaging device built around a CCD camera (CH350 model, 16 bit chip, Photometrics, Munchen, Germany) with an integral gel cutting tool. Imaging was performed at two excitation wavelengths, 545±10 and 635±15 nm for Cy3 and Cy5, respectively. Image manipulation and viewing was done with Image J (National Institutes of Health, Bethesda, MD, USA) and V++ Precision Digital Imaging System software (Digital Optics, Auckland, New Zealand). Two-frame, ‘flash’ movies of the overlying Cy3 and Cy5 images were viewed in a continuous loop to visually detect differences in the protein signatures between the two samples separated in the same gels. The repeat gels of reciprocally stained lysates revealed any differences due to dye uptake. Sample-dependent differences in protein distributions were confirmed and quantified using an image analysis software package, DeCyder Differential Analysis Software, version 5.0 (DeCyder®, Amersham Biosciences). DeCyder analyzed images as fragments, summed to generate a composite image, with proprietary algorithms that detected overlapping, differently colored images within the same gel to match spots and subtract background for normalization. DeCyder created measurement masks, each outlining an area of the gel containing a protein spot, that were applied to each of the matching fragments in the paired fluorescent images of each gel. The masks defined areas in which the pixel values were integrated to create three dimensional topographical maps with peaks representing each protein spot. An estimate of the local background for each spot determined the base values across the masked area. Output from the image analysis, designated as fluorescence intensity, was the sum of pixel values in the mask area minus the background. Volumes of the peaks representing the relative strengths of fluorescent signals from matched spots were compared. Although a threshold was set at 1.5-fold to screen for differences, only proteins with ≥2.0-fold differences were reported. Visual inspection confirmed the differences indicated by the DeCyder software. Dust and artifacts were detected as peaks with slope values of less than one or as clusters of sharp spikes. Images of some protein spots were also scanned and digitized to compare total pixels, as described below for immunoblots. Protein spots that differed on DIGE between the labeled lysates were harvested from the gels using a robotic gel picker integrated with the high resolution CCD camera, digested with trypsin, and submitted for mass spectrometric PMF analysis.
Protein Identification
Following trypsin digestion, the peptides derived from each protein spot were analyzed with matrix assisted laser desorption ionization time of flight mass spectrometry (MALDI-TOF-MS) in a 4700 Proteomics Analyzer with TOF/TOF Optics (Applied Biosystems, Foster City, CA, USA). Analysis of data with GPS Explorer Protein Analysis Software on a Remote Client workstation (Applied Biosystems) provided automated acquisition of optimized mass spectra and the derivation of monoisotopic PMF information. Searches of the NCBI nonredundant database, based on the peptide mass results using MASCOT (Matrix Science, Boston, MA, USA) via the GPS Explorer work station and MS-Fit (University of California, San Francisco MS Facility) via the internet identified proteins with matching PMFs. The MASCOT parameters were set to include all species (with the exception of spots from the gel region containing BSA), tolerance of one missed trypsin cleavage per protein, allowance of protein modification by methionine oxidation, peptide tolerance of 50 ppm (ie an example mass accuracy of 1000±0.05 Da), and restriction of peptides to 700–4000 Da. The MS-Fit parameters included tolerance of one missed trypsin cleavage per protein, cysteine modification by acrylamide, oxidation of methionine, and changes at the amino termini, such as conversion of glutamine to pyro-glutamate and acetylation. Protein identifications were accepted when the observed and predicted pI's and MR's were consistent and scores indicated nonrandom identifications at a significance level of P<0.05.
Densitometry of Immunoblots
Differential localization of proteins was also analyzed with specific antibodies. Lysates of whole cells and pseudopodia, 10 μg each, were electrophoresed in separate lanes of 10% polyacrylamide 1D gels under reducing conditions. Gel contents were transferred onto polyvinylidene difluoride membranes (Invitrogen), blocked (Detector Block, Protein Detector Western Blot Kit LumiGLO System, Kirkegaard & Perry Laboratories, Gaithersburg, MD, USA), and reacted with anti-human HGF (2 μg/ml) (Clone 24612.11, Sigma, St. Louis, MO, USA), anti-Met (1:1000) (Clone 25H2, Cell Signaling, Beverly, MA, USA), anti-actin (1:1000) (Sigma), and anti-AnxI, (0.2 μg/ml) (Clone EH17a, Santa Cruz Biotechnology, Inc.). The secondary antibodies, 1:1000 (Kirkegaard & Perry Laboratories) were horseradish peroxidase-labeled anti-mouse for reactions with HGF, Met, and AnxI, and anti-rabbit for actin. Protein standards (MultiMark, Invitrogen) were loaded with 8 μg per lane. Immunoreactive bands were visualized with horseradish peroxidase conversion of a luminal-based solution to produce chemiluminescence, scanned (Epson Perfection 2450 PHOTO transparency scanner, Epson America, Long Beach, CA, USA) and digitized (UN-SCAN-It gel, Silk Scientific, Orem, UT, USA). Band densities were corrected for adjacent background. Staining portions of the gels with Coomassie blue (Novex Colloidal Blue Stain Kit, Invitrogen) was used to confirm equal loading of pseudopodial and whole cell lysates.
Results
Characterization of Pseudopodia in U87 Cells
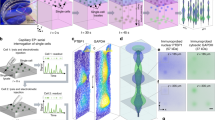
The cellular material that migrated through 3 μm diameter pores of filters, with and without a 0.01% gelatin (denatured collagen) coating, was stained and examined microscopically. Nuclei were not found after 5 h incubations at 37°C. Only a few thick, fingerlike pseudopodia had extended through some pores of the uncoated filters (Figure 2a). With gelatin present on the filters, migrated pseudopodia at 5 h formed discontinuous, thin sheets of various sizes that covered almost all the filters’ surfaces, involving many (more than half) of the pores. After the filters were removed, the slides and filters were separately stained with Diff Quik. Pseudopodia on the glass slides retained their appearance as thin, discontinuous sheets (Figure 2b) seen prior to filter removal. In separate assays, the nuclei also failed to migrate through the pores after 6 h. After overnight incubations (15 h), however, some nuclei had migrated through the filter pores, as shown in Figure 2c, thus verifying that pseudopodial formation at 4.5–5 h included the leading edge of migrating cells. Filters removed from the slides (Figure 2b and c) did not stain, indicating that no cell material remained (not shown).
Morphology of stained migrated U87 glioma cell materials. (a) Pseudopodia that formed on an uncoated filter. Darkly stained cytoplasmic material formed a few fingerlike pseudopodia during 4.5–5 h incubations without a matrix protein present. Pores, 3 μm in diameter, were evenly distributed in the overlying filter. No nuclei were present; 250-fold magnification. (b) Pseudopodia formed on a filter coated with 0.01% gelatin (denatured-collagen). Under the same conditions described above, the morphology and quantity of migrated cell material changed markedly when the filters were coated with matrix material. The porous filter was removed as described in Figure 1, allowing an unobstructed view. No residual material remained on the removed filters that were stained and examined. Before and after filter removal, migrated cell material appeared as irregularly shaped, lightly stained sheets that varied in size and covered most of the glass surface. No nuclei were present; 500-fold magnification. (c) Cell materials that migrated overnight. During overnight (15 h) incubations, nuclei migrated through the 3 μm pores of gelatin-coated filters, indicating that material found at 4.5–5 h included the leading edge; 500-fold magnification. Diff Quik stains for (a–c). The migration media is described in the text.
Pseudopodial Proteins Detected with Coomassie Blue Staining
Within ranges of 20–100 kD and pI's of 3–10, isoforms of 24 proteins were identified in pseudopodial lysates separated on 2D gels and stained with Coomassie blue. The largest group of stained proteins included metabolic enzymes catalyzing seven consecutive steps of the glycolytic pathway (aldolase, triose phosphate isomerase, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), phosphoglycerate kinase, phosphoglycerate mutase, enolase, and pyruvate kinase (PK)). A total of 17 isoforms of these glycolytic enzymes represented 46% of the total number of protein isoforms (37) identified on Coomassie blue-stained gels. Proteins involved with the cytoskeleton (actin, tropomyosin, vimentin, tubulin and moesin), heat shock protein (HSP)s (27, 60, 70, and 90 kDa), and annexin (Anx)s (I, II, and V) were also identified. Proteins that appeared as single isoforms on the gels, such as valosin-containing protein, HSP90, HSP60, calumenin precursor, tropomyosin, ubiquitin carboxyl-terminal esterase LI, HSP27, phosphoglycerate mutase and tropomyosin, had predicted pI's of 5.14, 5.3, 5.7, 4.4, 4.67, 5.33, 5.98, 6.67 and 6.45, respectively, all of which approximately corresponded with the observed pI's of the migrated spots on the gel shown in Figure 3. These and other proteins are listed in Table 1 with their database identification number, confidence scores, and other parameters. The protein expression patterns of whole cell lysates harvested at 2 and 5 h were similar.
Representative 2D gel of the relatively abundant proteins in U87 pseudopodial lysates. Coomassie blue-stained gels, 8–16% polyacrylamide, containing pseudopodial lysate showed the distribution of 22 proteins identified with PMF analysis. A total of 17 isoforms of glycolytic enzymes represented 46% of the total number of protein isoforms (37) seen. Table 1 lists 24 pseudopodial proteins with their parameters, identified by PMF analysis using MALDI-TOF-MS. Exogenous BSA, present in box 5, is not listed. The pH ranged from 3 to 10, left and right margins of the gel, respectively. The expected pI's of proteins that migrated as single isoforms are stated in the Results section.
Differential Distributions of Proteins Detected with DIGE
Image analysis of fluorescently labeled lysates loaded together on 2D gels identified different distributions of protein isoforms in pseudopodia compared to whole cells. A representative DIGE gel is shown (Figure 4) with its DeCyder analysis portrayed topographically in Figure 5 and the quantified differences listed in Table 2. The signals for two proteins, transketolase and mitochondrial manganese superoxide dismutase (MnSOD), were significantly decreased in pseudopodia compared to whole cells (Figure 5a). Abundant proteins that increased at least two-fold in pseudopodia and were identified on the DIGE gel shown (Figure 4) included isoforms of AnxI, enolase, and PK (Figure 5b). Inclusion of data from additional gels indicated that aldolase and AnxII were also elevated in pseudopodia. Although the DeCyder-determined ratio for aldolase was only 1.94 (almost significant) in the DIGE gel shown in Figure 4, its ratio, 2.09, was above the threshold of significance in another DIGE gel. A protein spot in Figure 4's gel that corresponded to the placement of AnxII isoforms in Figure 3 had a DeCyder ratio of 2.48 but did not yield satisfactory material for mass spectrometry. In another DIGE gel, a protein spot in the same location was identified as AnxII and demonstrated a DeCyder ratio of 1.88. The left edge and lower mid region of the large, irregularly shaped spot identified as actin yielded DeCyder ratios greater than two-fold, but distinct isoforms of actin did not resolve for clearcut analysis. In a separate analysis using densitometric data from the scanned fluorescent signal, total actin increased by 38% in the pseudopodial lysate on DIGE. Keratins were found preferentially in pseudopodia, but may have resulted from hair and skin contamination that accumulated in lysates when several slides were stored and batched during harvests of pseudopodia. Increased BSA, an exogenous protein in the media, was not considered biologically significant. BSA may have bound to glass surfaces not covered by pseudopodia or it may have preferentially adhered to the surfaces of pseudopodia. Lack of successful identification with MALDI-TOF-MS for many of the fluorescent protein spots that yielded significant DeCyder ratios was mostly attributed to insufficient amounts of protein. None of the pseudopodial proteins with sufficient abundance to be detected and identified on gels stained with Coomassie blue (Table 1) were found to be decreased compared to whole cells on DIGE gels.
Representative DIGE images of paired whole cell and pseudopodial lysates shown as inverse images of the fluorescent stains. (a) Whole cell lysate labeled with Cy5 demonstrated relatively increased quantities of transketolase (arrow 1) and MnSOD (arrow 2) compared to the pseudopodial lysates. (b) Pseudopodial lysate labeled with Cy3 demonstrated relatively increased quantities of two isoforms for AnxI, one isoform for enolase, two isoforms for PK, and merged isoforms for actin (see text), as indicated with arrows, 1 and 2, 3, 4 and 5, and 7, respectively. Increases in the isoforms for aldolase and AnxII (positions indicated on the Coomassie-stained 2D gel in Figure 3) are discussed in the text. Apparent increases in exogenous BSA, arrow 6, and keratin (probable contaminant) were not considered to be biologically relevant. Selected matching protein spots with digitized representations are topographically compared in Figure 5 and the differences are quantified in Table 2. The background was normalized for both images. The 2D gel parameters were similar to those described for Figure 3.
The topographically rendered densitometry results for selected paired spots in the DIGE gel shown in Figure 4. Pairs of encircled (black) peaks, pseudopodial (left) and whole cells (right), represent the spots highlighted on the inserts of gel images. Arrows indicate the corresponding pseudopodial protein spots. Proteins were identified with MALDI-TOF-MS. (a) Two proteins, transketolase and MnSOD, were decreased in pseudopodia compared to whole cells, indicated as 1 and 2, respectively. (b) Selected proteins that were increased in pseudopodia compared to whole cells are indicated as follows, isoforms of AnxI as 1 and 2, enolase as 3, isoforms of PK as 4 and 5. These proteins are further described in Table 2.
Differential Distributions of Proteins on Immunoblots of 1D Gels
Analysis of paired pseudopodial and whole cell lysates demonstrated immunoreactivity to the 69 kDa α-chain of activated HGF only in pseudopodial lystates. The antibody was raised against recombinant human HGF. Met immunoreactivity, apparent as a 145 kDa band, was present in both types of lysates and increased up to 79% in the pseudopodia. Increased immunoreactivity was present for total AnxI and actin, 35 and 66%, respectively, in pseudopodia (Figure 6a). Equalized loading of lysates was also demonstrated with Coomassie blue staining (Figure 6b).
Representative immunoblots comparing several proteins in lysates of pseudopodia (Ps) and whole cells (WC) with equalized gel loading. (a) Proteins were separated on 1D, 10% polyacrylamide SDS gels under reducing conditions, blotted, and reacted with specific antibodies. An antibody raised against human recombinant HGF reacted with the 69 kDa α-chain of activated HGF in Ps but no reactivity was found in lysates of WC. A specific antibody for Met, the receptor for HGF, reacted with 145 kDa bands representing the β-subunit of Met in both lysates of Ps and WC with increased (79%) reactivity in Ps. Antibodies specific for total AnxI and actin reacted with 40 and 42 kDa bands, respectively, demonstrating 35 and 66% increased reactivities, respectively, in Ps. (b) Equalized loading of Ps and WC lysates was demonstrated in a portion of the gel sliced from the one transferred and immunoblotted with anti-AnxI and anti-actin shown in (a): Coomassie blue stain.
Discussion
Malignancy of many tumors is defined by invasive and/or metastatic behavior. Diffuse astrocytomas invade widely through surrounding central nervous system tissue. These tumors migrate preferentially through white matter tracts formed by axons, glial processes, and the vasculature. Although white matter of the brain is a dense meshwork, it can be loosened by tumor edema and contains fewer blood vessels and neural processes than gray matter. The process of cell locomotion is often studied in single cells that crawl independently on flat surfaces in vitro. Their cytoplasmic protrusions have been named lamellipodia, filopodia, ruffles, spikes and blebs. Some of these structures have been characterized by the proteins involved in their formation.7, 8 However, during brain invasion, tumor cells migrate mostly through the 3D, interstitial spaces of white matter. The term ‘pseudopodia’ has been used to describe tumor cell protrusions into the 5–7 μm openings of micropipette tips9 and protrusions through 1 μm pores of filters.10 In the current study, pseudopodia were formed by the tumor cell protrusions through 3 μm diameter pores of filters.
Microscopic examination of migrated pseudopodia at different time points confirmed inclusion of the leading edge in the harvested material by demonstrating nuclear migration at a later time point. The 2D analysis of pseudopodia formed by motile glioma cells provided a proteomic view, with some constituents noted for their association with high grade gliomas and progression in other types of tumors (Table 1). Immunoreactions with specific antibodies documented the potential for localized signaling in pseudopodia by an activated, well-known glioma migration factor, HGF. DIGE detected changes in several constituents of the pseudopodial proteome compared to whole cells that potentially include important effectors of migration signaling.
Both HGF and Met are known to be produced by glial cells, in vivo and in vitro, including the U87 cell line.2, 4, 11, 12, 13, 14 Expression of HGF and Met are associated with tumor progression.3, 4, 15, 16 In other types of malignant cells, downstream events induced by HGF signaling have been shown to include increased glycolysis17 and phoshorylation of AnxI.18 This study revealed the potential for HGF signaling to mediate similar changes within the leading edge of U87 glioma cells.
Malignant astrocytic tumors are known to be glycolytic, possibly attributed to decreased numbers of functional mitochondria.19 Several glycolytic enzymes, including GAPDH, PK, and phosphoglycerate kinase, have been found on the cytoplasmic face of the plasma membrane in malignant astrocytoma cells.20 Phosphoglycerate mutase was previously found to be increased in high-grade vs low-grade gliomas.21 In the current study, the abundance of at least seven enzymes that catalyze consecutive steps in glycolysis (aldolase through PK reactions) in U87 cells is consistent with the prominence of glycolytic metabolism in gliomas. In previous studies, glycolysis has been shown to provide energy for the migration of malignant cells when mitochondria were inhibited. With mitochondrial inhibitors added to force reliance on glycolytic energy, we showed that A2058 melanoma and U87 glioma cells migrated at levels comparable to migration in normal conditions.22, 23 Others demonstrated the presence of GAPDH in pseudopodia and found that glycolysis was necessary for the formation of protrusions in an invasive, transformed epithelial MDCK cell line.10
Site-directed actin polymerization in response to signaling is needed for the formation of cell protrusions.24 Glycolytic enzymes are known to bind cytoskeletal proteins, including actin filaments, microtubules, and tropomyosin, in addition to existing as unbound cytoplasmic proteins. Numerous studies have demonstrated interactions between the cytoskeleton and glycolytic enzymes, especially GAPDH, aldolase and PK.25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38 In this study, quantities of these glycolytic enzymes, as well as actin, tubulin and tropomyosin, were sufficient in U87 pseudopodia to be detected with Coomassie blue staining, thus establishing the potential for the energy-generating portion of the glycolytic pathway to energize cytoskeletal proteins for migration within U87 pseudopodia. Polymerization of actin and tubulin shares a similar use of energy from nucleotide triphosphate hydrolysis.39, 40, 41, 42, 43, 44 Increased actin in pseudopodia, as found in this study, has been previously documented by others.10, 45 Increased isoforms for enolase, aldolase, and PK detected with DIGE in U87 pseudopodia may help to energize the leading edge.
Several types of annexins, I, II, and V, were found in the U87 pseudopodia. The Anx family is comprised of numerous gene products, 12 in vertebrates, with various functions proposed for controlling membrane structure and transport, vesicle secretion, anti-inflammation, prostaglandin metabolism, apoptosis, and other functions.46 AnxI is also known as lipocortin I, calpactin II, epidermal growth factor receptor substrate, and chromobindin 9.47 Two isoforms of AnxI were increased in pseudopodia compared to whole cells as determined by DIGE, and total AnxI was modestly increased when compared on immunoblots. Previous studies have identified AnxI within CNS tumors, including astrocytomas,48 and in the anterior pituitary gland.49 Involvement of AnxI in cell surface changes induced by steroids has been demonstrated in C6 rat astrocytoma cells.50, 51 AnxI is also involved in the regulation of prostaglandins and nitric oxide.52, 53 Previous associations of AnxI with tumor progression were found with 2D electrophoresis in studies that showed its differential expression in chemoresistant gastric carcinoma cell lines,54 a cell line established from a metastasis compared to primary squamous carcinoma of the larynx,55 and in tissue containing metastasized breast carcinoma cells compared to normal tissue.56 Immunohistochemical studies of breast and hepatocellular carcinoma also indicated that AnxI is associated with tumor progression.57, 58, 59 The contribution of AnxI to pseudopodial extension during migration may explain AnxI's association with tumor progression and HGF signaling seen in other cells18 and with activated HGF in U87 pseudopodia. Also, AnxII was previously identified more frequently in high-grade than low-grade gliomas, 50 of 65 and three of 10 tumors, respectively, following 2D electrophoresis of tissue samples.21
Sensitivity for visualizing stained proteins in 2D gels, approximately 40 ng for Coomassie blue, is increased by using fluorescent dyes and silver nitrate to 5 and 0.5 ng, respectively. However, our high success rate for identifying proteins detected with Coomassie blue fell to a much lower rate for spots demonstrated with the fluorescent dyes. Although the relatively low abundance of signaling proteins that control cell migration prevents their proteomic study by current 2D gel technologies,60 specific antibodies on immunblots detected a striking localization of active HGF to pseudopodia that also contained increased Met. Combining DIGE with specific antibody reactivities on immunoblots demonstrated significant increases in isoforms for actin, enolase, PK, and AnxI, and probable increases of aldolase and AnxII in the pseudopodia. Identifying the functional proteome of tumor pseudopodia will help to elucidate mechanisms of cell migration in tissue and will aid in the selection of biomarkers and molecular treatment targets for malignant gliomas.
References
Brockmann MA, Ulbricht U, Gruner K, et al. Glioblastoma and cerebral microvascular endothelial cell migration in response to tumor-associated growth factors. Neurosurgery 2003;52:1391–1399.
Koochekpour S, Jeffers M, Rulong S, et al. Met and hepatocyte growth factor/scatter factor expression in human gliomas. Cancer Res 1997;57:5391–5398.
Lamszus K, Schmidt NO, Jin L, et al. Scatter factor promotes motility of human glioma and neuromicrovascular endothelial cells. Int J Cancer 1998;75:19–28.
Lamszus K, Laterra J, Westphal M, et al. Scatter factor/hepatocyte growth factor (SF/HGF) content and function in human gliomas. Int J Dev Neurosci 1999;17:517–530.
Derouiche A, Frotscher M . Peripheral astrocyte processes: monitoring by selective immunostaining for the actin-binding ERM proteins. Glia 2001;36:330–341.
Unlu M, Morgan ME, Minden JS . Difference gel electrophoresis: a single gel method for detecting changes in cell extracts. Electrophoresis 1997;18:2071–2077.
Horwitz AR, Parsons JT . Cell migration–movin’ on. Science 1999;286:1102–1103.
Ridley AJ, Schwartz MA, Burridge K, et al. Cell migration: integrating signals from front to back. Science 2003;302:1704–1709.
Dong C, Aznavoorian S, Liotta LA . Two phases of pseudopod protrusion in tumor cells revealed by a micropipette. Microvasc Res 1994;47:55–67.
Nguyen TN, Wang HJ, Zalzal S, et al. Purification and characterization of beta-actin-rich tumor cell pseudopodia: role of glycolysis. Exp Cell Res 2000;258:171–183.
Abounader R, Ranganathan S, Lal B, et al. Reversion of human glioblastoma malignancy by U1 small nuclear RNA/ribozyme targeting of scatter factor/hepatocyte growth factor and c-met expression. J Natl Cancer Inst 1999;91:1548–1556.
Hollborn M, Krausse C, Iandiev I, et al. Glial cell expression of hepatocyte growth factor in vitreoretinal proliferative disease. Lab Invest 2004;84:963–972.
Yamada T, Yoshiyama Y, Tsuboi Y, et al. Astroglial expression of hepatocyte growth factor and hepatocyte growth factor activator in human brain tissues. Brain Res 1997;762:251–255.
Welch WC, Kornblith PL, Michalopoulos GK, et al. Hepatocyte growth factor (HGF) and receptor (c-met) in normal and malignant astrocytic cells. Anticancer Res 1999;19:1635–1640.
Schmidt NO, Westphal M, Hagel C, et al. Levels of vascular endothelial growth factor, hepatocyte growth factor/scatter factor and basic fibroblast growth factor in human gliomas and their relation to angiogenesis. Int J Cancer 1999;84:10–18.
Yano H, Hara A, Murase S, et al. Expression of hepatocyte growth factor and matrix metalloproteinase-2 in human glioma. Brain Tumor Pathol 2001;18:7–12.
Kaplan O, Firon M, Vivi A, et al. HGF/SF activates glycolysis and oxidative phosphorylation in DA3 murine mammary cancer cells. Neoplasia 2000;2:365–377.
Skouteris GG, Schroder CH . The hepatocyte growth factor receptor kinase-mediated phosphorylation of lipocortin-1 transduces the proliferating signal of the hepatocyte growth factor. J Biol Chem 1996;271:27266–27273.
Oudard S, Boitier E, Miccoli L, et al. Gliomas are driven by glycolysis: putative roles of hexokinase, oxidative phosphorylation and mitochondrial ultrastructure. Anticancer Res 1997;17:1903–1911.
Daum G, Keller K, Lange K . Association of glycolytic enzymes with the cytoplasmic side of the plasma membrane of glioma cells. Biochim Biophys Acta 1988;939:277–281.
Iwadate Y, Sakaida T, Hiwasa T, et al. Molecular classification and survival prediction in human gliomas based on proteome analysis. Cancer Res 2004;64:2496–2501.
Beckner ME, Stracke ML, Liotta LA, et al. Glycolysis as primary energy source in tumor cell chemotaxis. J Natl Cancer Inst 1990;82:1836–1840.
Beckner ME, Pollack IF . Treatment of hypoxia-resistant astrocytoma cell migration with glucose inhibitors. Neuro-Oncology 2003;5:280.
Carlier MF, Wiesner S, Le Clainche C, et al. Actin-based motility as a self-organized system: mechanism and reconstitution in vitro. C R Biol 2003;326:161–170.
Bronstein WW, Knull HR . Interaction of muscle glycolytic enzymes with thin filament proteins. Can J Biochem 1981;59:494–499.
Durrieu C, Bernier-Valentin F, Rousset B . Binding of glyceraldehyde 3-phosphate dehydrogenase to microtubules. Mol Cell Biochem 1987;74:55–65.
Durrieu C, Bernier-Valentin F, Rousset B . Microtubules bind glyceraldehyde 3-phosphate dehydrogenase and modulate its enzyme activity and quaternary structure. Arch Biochem Biophys 1987;252:32–40.
Masters C . Interactions between glycolytic enzymes and components of the cytomatrix. J Cell Biol 1984;99 (1 Part 2):222s–225s.
Karkhoff-Schweizer R, Knull HR . Demonstration of tubulin-glycolytic enzyme interactions using a novel electrophoretic approach. Biochem Biophys Res Commun 1987;146:827–831.
Mejean C, Pons F, Benyamin Y, et al. Antigenic probes locate binding sites for the glycolytic enzymes glyceraldehyde-3-phosphate dehydrogenase, aldolase and phosphofructokinase on the actin monomer in microfilaments. Biochem J 1989;264:671–677.
Minaschek G, Groschel-Stewart U, Blum S, et al. Microcompartmentation of glycolytic enzymes in cultured cells. Eur J Cell Biol 1992;58:418–428.
Somers M, Engelborghs Y, Baert J . Analysis of the binding of glyceraldehyde-3-phosphate dehydrogenase to microtubules, the mechanism of bundle formation and the linkage effect. Eur J Biochem 1990;193:437–444.
Sirover MA . New insights into an old protein: the functional diversity of mammalian glyceraldehyde-3-phosphate dehydrogenase. Biochim Biophys Acta 1999;1432:159–184.
Walsh JL, Knull HR . Heteromerous interactions among glycolytic enzymes and of glycolytic enzymes with F-actin: effects of poly(ethylene glycol). Biochim Biophys Acta 1988;952:83–91.
Walsh JL, Keith TJ, Knull HR . Glycolytic enzyme interactions with tubulin and microtubules. Biochim Biophys Acta 1989;999:64–70.
Volker KW, Knull HR . Glycolytic enzyme–tubulin interactions: role of tubulin carboxy terminals. J Mol Recogn 1993;6:167–177.
Volker KW, Reinitz CA, Knull HR . Glycolytic enzymes and assembly of microtubule networks. Comp Biochem Physiol B Biochem Mol Biol 1995;112:503–514.
Volker KW, Knull H . A glycolytic enzyme binding domain on tubulin. Arch Biochem Biophys 1997;338:237–243.
Carlier MF, Pantaloni D, Evans JA, et al. The hydrolysis of ATP that accompanies actin polymerization is essentially irreversible. FEBS Lett 1988;235:211–214.
Carlier MF . Nucleotide hydrolysis regulates the dynamics of actin filaments and microtubules. Philos Trans R Soc London B Biol Sci 1992;336:93–97.
Korn ED, Carlier MF, Pantaloni D . Actin polymerization and ATP hydrolysis. Science 1987;238:638–644.
Dufort PA, Lumsden CJ . How profilin/barbed-end synergy controls actin polymerization: a kinetic model of the ATP hydrolysis circuit. Cell Motil Cytoskeleton 1996;35:309–330.
Loisel TP, Boujemaa R, Pantaloni D, et al. Reconstitution of actin-based motility of Listeria and Shigella using pure proteins. Nature 1999;401:613–616.
Mitchison TJ . Evolution of a dynamic cytoskeleton. Philos Trans R Soc London B Biol Sci 1995;349:299–304.
Le PU, Nguyen TN, Drolet-Savoie P, et al. Increased beta-actin expression in an invasive moloney sarcoma virus-transformed MDCK cell variant concentrates to the tips of multiple pseudopodia. Cancer Res 1998;58:1631–1635.
Rescher U, Gerke V . Annexins—unique membrane binding proteins with diverse functions. J Cell Sci 2004;117 (Part 13):2631–2639.
Fernandez MP, Morgan RO . Annexin Sequence Database, The New Nomenclature. European Annexin Homepage 2004;www.annexins.org.
Johnson MD, Kamso-Pratt J, Pepinsky RB ; et al. Lipocortin-1 immunoreactivity in central and peripheral nervous system glial tumors. Hum Pathol 1989;20:772–776.
Turgeon JL, Cooper RH, Waring DW . Membrane-specific association of annexin I and annexin II in anterior pituitary cells. Endocrinology 1991;128:96–102.
McLeod JD, Goodall A, Jelic P, et al. Changes in the cellular distribution of lipocortin-1 (Annexin-1) in C6 glioma cells after exposure to dexamethasone. Biochem Pharmacol 1995;50:1103–1107.
Mizuno H, Uemura K, Moriyama A, et al. Glucocorticoid induced the expression of mRNA and the secretion of lipocortin 1 in rat astrocytoma cells. Brain Res 1997;746:256–264.
Cirino G, Flower RJ . Human recombinant lipocortin 1 inhibits prostacyclin production by human umbilical artery in vitro. Prostaglandins 1987;34:59–62.
Minghetti L, Nicolini A, Polazzi E, et al. Down-regulation of microglial cyclooxygenase-2 and inducible nitric oxide synthase expression by lipocortin 1. Br J Pharmacol 1999;126:1307–1314.
Sinha P, Hutter G, Kottgen E, et al. Increased expression of annexin I and thioredoxin detected by two-dimensional gel electrophoresis of drug resistant human stomach cancer cells. J Biochem Biophys Methods 1998;37:105–116.
Wu W, Tang X, Hu W, et al. Identification and validation of metastasis-associated proteins in head and neck cancer cell lines by two-dimensional electrophoresis and mass spectrometry. Clin Exp Metast 2002;19:319–326.
Cicek M, Samant RS, Kinter M, et al. Identification of metastasis-associated proteins through protein analysis of metastatic MDA-MB-435 and metastasis-suppressed BRMS1 transfected-MDA-MB-435 cells. Clin Exp Metast 2004;21:149–157.
Pencil SD, Toth M . Elevated levels of annexin I protein in vitro and in vivo in rat and human mammary adenocarcinoma. Clin Exp Metast 1998;16:113–121.
Masaki T, Tokuda M, Ohnishi M, et al. Enhanced expression of the protein kinase substrate annexin in human hepatocellular carcinoma. Hepatology 1996;24:72–81.
Ahn SH, Sawada H, Ro JY, et al. Differential expression of annexin I in human mammary ductal epithelial cells in normal and benign and malignant breast tissues. Clin Exp Metast 1997;15:151–156.
Lim MS, Elenitoba-Johnson KS . Proteomics in pathology research. Lab Invest 2004;84:1227–1244.
Diaz VM, Hurtado M, Thomson TM, et al. Specific interaction of tissue-type plasminogen activator (t-PA) with annexin II on the membrane of pancreatic cancer cells activates plasminogen and promotes invasion in vitro. Gut 2004;53:993–1000.
Hajjar KA, Acharya SS . Annexin II and regulation of cell surface fibrinolysis. Ann NY Acad Sci 2000;902:265–271.
Roshy S, Sloane BF, Moin K . Pericellular cathepsin B and malignant progression. Cancer Metast Rev 2003;22:271–286.
Jones PG, Moore GJ, Waisman DM . A nonapeptide to the putative F-actin binding site of annexin-II tetramer inhibits its calcium-dependent activation of actin filament bundling. J Biol Chem 1992;267:13993–13997.
Roseman BJ, Bollen A, Hsu J, et al. Annexin II marks astrocytic brain tumors of high histologic grade. Oncol Res 1994;6:561–567.
Allen RT, Hunter Jr W, Agrawal DK . Morphological and biochemical characterization and analysis of apoptosis. J Pharmacol Toxicol Methods 1997;37:215–228.
Townsend DM, Tew KD . The role of glutathione-S-transferase in anti-cancer drug resistance. Oncogene 2003;22:7369–7375.
Bagatell R, Beliakoff J, David CL, et al. Hsp90 inhibitors deplete key anti-apoptotic proteins in pediatric solid tumor cells and demonstrate synergistic anticancer activity with cisplatin. Int J Cancer 2005;113:179–188.
Bagatell R, Whitesell L . Altered Hsp90 function in cancer: a unique therapeutic opportunity. Mol Cancer Ther 2004;3:1021–1030.
Beliakoff J, Bagatell R, Paine-Murrieta G, et al. Hormone-refractory breast cancer remains sensitive to the antitumor activity of heat shock protein 90 inhibitors. Clin Cancer Res 2003;9:4961–4971.
Workman P . Combinatorial attack on multistep oncogenesis by inhibiting the Hsp90 molecular chaperone. Cancer Lett 2004;206:149–157.
Myllyharju J . Prolyl 4-hydroxylases, the key enzymes of collagen biosynthesis. Matrix Biol 2003;22:15–24.
Pickart CM, Rose IA . Ubiquitin carboxyl-terminal hydrolase acts on ubiquitin carboxyl-terminal amides. J Biol Chem 1985;260:7903–7910.
Asai T, Tomita Y, Nakatsuka S, et al. VCP (p97) regulates NFkappaB signaling pathway, which is important for metastasis of osteosarcoma cell line. Jpn J Cancer Res 2002;93:296–304.
Tsujimoto Y, Tomita Y, Hoshida Y, et al. Elevated expression of valosin-containing protein (p97) is associated with poor prognosis of prostate cancer. Clin Cancer Res 2004;10:3007–3012.
Yamamoto S, Tomita Y, Hoshida Y, et al. Expression level of valosin-containing protein is strongly associated with progression and prognosis of gastric carcinoma. J Clin Oncol 2003;21:2537–2544.
Yamamoto S, Tomita Y, Nakamori S, et al. Elevated expression of valosin-containing protein (p97) in hepatocellular carcinoma is correlated with increased incidence of tumor recurrence. J Clin Oncol 2003;21:447–452.
Yamamoto S, Tomita Y, Hoshida Y, et al. Expression of valosin-containing protein in colorectal carcinomas as a predictor for disease recurrence and prognosis. Clin Cancer Res 2004;10:651–657.
Yamamoto S, Tomita Y, Hoshida Y, et al. Increased expression of valosin-containing protein (p97) is associated with lymph node metastasis and prognosis of pancreatic ductal adenocarcinoma. Ann Surg Oncol 2004;11:165–172.
Yamamoto S, Tomita Y, Hoshida Y, et al. Expression level of valosin-containing protein (p97) is correlated with progression and prognosis of non-small-cell lung carcinoma. Ann Surg Oncol 2004;11:697–704.
Yamamoto S, Tomita Y, Hoshida Y, et al. Expression level of valosin-containing protein (VCP) as a prognostic marker for gingival squamous cell carcinoma. Ann Oncol 2004;15:1432–1438.
Yamamoto S, Tomita Y, Hoshida Y, et al. Expression level of valosin-containing protein (p97) is associated with prognosis of esophageal carcinoma. Clin Cancer Res 2004;10:5558–5565.
Acknowledgements
We thank April E Engram, Fatima Burovic, and Beth Arnold for technical assistance. We thank Sadie Aznavoorian Cheshire, PhD (Prism Medical Communications, Inc., Morristown, NJ, USA) for helpful comments. We gratefully acknowledge financial support from The Nick Eric Wichman Foundation, Ellicott City, MD, USA and The Walter L Copeland Fund for Cranial Research of the Pittsburgh Foundation, Pittsburgh, PA, USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Beckner, M., Chen, X., An, J. et al. Proteomic characterization of harvested pseudopodia with differential gel electrophoresis and specific antibodies. Lab Invest 85, 316–327 (2005). https://doi.org/10.1038/labinvest.3700239
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700239
Keywords
This article is cited by
-
Protrusion-localized STAT3 mRNA promotes metastasis of highly metastatic hepatocellular carcinoma cells in vitro
Acta Pharmacologica Sinica (2016)
-
A Metabolic Inhibitory Cocktail for Grave Cancers: Metformin, Pioglitazone and Lithium Combination in Treatment of Pancreatic Cancer and Glioblastoma Multiforme
Biochemical Genetics (2016)
-
The expanding role of metformin in cancer: an update on antitumor mechanisms and clinical development
Targeted Oncology (2016)
-
Guanine nucleotide exchange factor Dock7 mediates HGF-induced glioblastoma cell invasion via Rac activation
British Journal of Cancer (2014)
-
Direct RNA sequencing mediated identification of mRNA localized in protrusions of human MDA-MB-231 metastatic breast cancer cells
Journal of Molecular Signaling (2013)