Abstract
In this paper, biospecific interaction analysis (BIA) employing surface plasmon resonance (SPR) and biosensor technologies was applied to the analysis of multiple mutations of the human beta-globin gene. To this aim, large target polymerase chain reaction (PCR) products were immobilized on sensor chips and then probes detecting beta°39 (C>T), beta°IVSI-1 (G>A), beta+IVSI-6 (T>C) and beta+IVSI-110 (G>A) thalassemia mutations were sequentially injected. In this study, a total of ten normal and seven heterozygous subjects, and six homozygous patients were considered. The results obtained allow to conclude that discrimination between normal subjects, heterozygous, and homozygous patients is readily achieved for all the four mutations by PCR amplification of genomic DNA containing all the regions corresponding to the same mutations, immobilization of the same PCR products, and hybridization. To our knowledge the procedure described here is the first reported on the use of SPR-based BIA and biosensor technology for multiple detections of point mutations.
Similar content being viewed by others
Main
We have recently demonstrated that biosensor technologies for biospecific interaction analysis (BIA)1, 2, 3, 4, 5, 6, 7 are suitable approaches for detecting point mutations involved in human diseases. For instance, we have recently published two reports on the use of surface plasmon resonance (SPR)-based BIA and oligonucleotide (ODN) probes to detect cystic fibrosis deltaF508 and W1282X mutations8, 9, 10 present in target DNA amplified by polymerase chain reaction (PCR).11 The conclusions of these two studies were that (a) the ability of ODN probes to discriminate efficiently in SPR-based BIA between target sequences differing in terms of a single nucleotide largely depends on length; (b) short ODNs (9–10 mers) perform differential hybridization thereby being suitable for diagnostics, but, unfortunately, they could be inefficient in hybridizing with single–stranded PCR products exhibiting heavy secondary structures; (c) longer oligonucleotide probes (12–13 mers) are not informative during the hybridization step (they hybridize to both normal and mutated DNAs), but could be used, with excellent results, in the dissociation phase, since stability of mismatched hybrids is very low; and (d) 15–20 mer ODN probes do not discriminate between full matched and mismatched target DNA in BIA experiments, yielding in both cases stable hybrids. The hypothesis of our studies was that the molecular BIA diagnosis based on SPR strongly depends on the secondary structure of the target PCR product. In a more recent paper,9 we demonstrated that short peptide nucleic acid (PNA)12 probes (9 mers) were efficient, unlike the respective ODN probes, in hybridizing to target PCR products, the Tm of DNA/PNA hybrids being higher than that of DNA/DNA hybrids, due to the fact that they are not negatively charged and, therefore, no electrostatic repulsion occurs during DNA/PNA hybrid formation.13, 14
The most recent findings in SPR-based analysis of PCR products have been discussed in a recent review article from our laboratory.15 The suitability of SPR-based BIA for the detection of point mutations has been recently confirmed in molecular diagnosis of the beta°39 (C>T) thalassemia mutation of the human beta-globin gene16, 17 involved in a severe type of beta°-thalassemia.18
However, to our knowledge, no attempt has been made to perform SPR-based BIA for multiple clinically relevant mutations in genetic diseases. This is of great relevance in the molecular diagnosis of the highly heterogeneous beta-thalassemia syndromes, in which double heterozygotes are frequently found.
It should be pointed out that the possibility to perform multiplex analyses on the same sensor chip is not an easy approach, due to the fact that probes interacting in regions of similar PCR products exhibiting different secondary structures might display highly variable hybridization efficiency, as recently reported by our group in several papers and reviews.8, 9, 10, 15, 18, 19, 20
The aim of this study was to verify whether SPR-based BIA could be employed in multiplex analysis for real-time detection of several mutations of the human beta-globin gene causing beta-thalassemia.
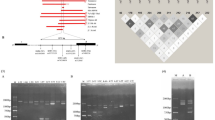
In the designed BIA approach format, biotinylated PCRs were performed using, as target, genomic DNA from normal subjects and from informative patients. The PCR products were designed to contain gene regions involved in four beta-thalassemia mutations: beta°39 (C>T), beta°IVSI-1 (G>A),21 beta+IVSI-6 (T>C),21 and beta+IVSI-110 (G>A)22, 23 (Figure 1). After immobilization of the target PCR products on the sensor chips, beta39, beta(I)1, beta(I)6, and beta(I)110 normal (N-) and mutated (M-) probes were sequentially injected and the bound resonance units (RU) were detected.
Location of beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110 thalassemia mutations, 5′-biotinylated biot-beta-glob-F and beta-glob-R PCR primers, beta39 (grey box), beta(I)1 (white box), beta(I)6, (black box), and beta(I)110 (dashed box) normal and mutated probes within the human beta-globin gene. The underlined sequences represent exonic portions of the beta-globin gene.
Materials and methods
Synthetic Oligonucleotides
The location and nucleotide sequences of the PCR primers and the normal and mutated oligonucleotide probes used in our experiments are reported in Table 1 and Figure 1. The beta-globin region was amplified using the 5′-end biotinylated beta-glob-F (biot-beta-glob-F) and the beta-glob-R primers. These oligonucleotides were purified by high-performance liquid chromatography (HPLC) and purchased from Sigma Genosys (Cambridge, UK).
Polymerase Chain Reaction
In each PCR reaction, 100 ng of human genomic DNA were amplified by Taq DNA polymerase using the beta-glob-F and beta-glob-R primers, amplifying the region of the human beta-globin gene containing sequences corresponding to the beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110 thalassemia mutations. PCR was performed in a final volume of 100 μl, containing 50 mM KCl, 10 mM TRIS-HCl, pH 8.8, 1.5 mM MgCl2, 0.1% Triton® X-100, 33 μM dNTPs, 0.3 μM PCR primers, by using 2 U/reaction of Taq DNA polymerase. The 40 PCR cycles used were as follows: denaturation, 30 s, 95°C; annealing, 20 s, 65°C; and elongation, 20 s, 72°C. The length of the beta-globin PCR product was 232 bp.
Surface Plasmon Resonance
The BIAcore™ 1000 analytical system (Biacore AB, Uppsala, Sweden) was used in all experiments. The running buffer HEPES-buffered saline-EP (HBS-EP), containing 10 mM HEPES, pH 7.4, 0.15 M NaCl, 3 mM EDTA, 0.005% (v/v) Surfactant P20, was from Biacore AB (Uppsala, Sweden). The experiments were conducted at 25°C with a 5 μl/min flow rate. In order to obtain an efficient capture of 5′-biotinylated PCR products onto the sensor chip, the well-documented streptavidin–biotin interaction was employed.24 To this end, the sensor chip SA, precoated with streptavidin, was used. After pretreatment with three 10 μl pulses with 40 mM NaOH–1 M NaCl, three injections of 40 μl of HBS-EP, 0.5 M NaCl, containing PCR products were administered in different flow cells. Hybridization was carried out by injecting 20 μl of HBS-EP buffer containing normal beta-N and mutated beta-M oligonucleotide probes. After hybridization, the sensor chips were regenerated by performing a 5 μl pulse of 50 mM NaOH.8, 10
Sensorgrams were analyzed with the BIAevaluation™ 2.1 software.6 Blank subtractions were performed in all the experiments reported.
Sequencing of PCR Products
PCR products obtained using, as target, genomic DNA from normal, heterozygous, or homozygous subjects were purified with Microcon®-30 (Millipore Corporation, Bedford, MA, USA) and sequenced using the ABI PRISM™ BigDye™ Terminator Cycle Sequencing Ready Reaction Kit and the ABI PRISM™ 377 DNA sequencer (PE Applied Biosystems, Foster City, CA, USA).
Computer-assisted Prediction of Secondary Structures of Single-Stranded PCR Products
Secondary structures of single-stranded PCR products carrying both normal and mutated beta-globin gene sequences were determined using the MFOLD software (version 3.1) developed by Zuker et al25 and Mathews et al.26 The analysis was performed at a temperature of 25°C and at 0.15 M NaCl.
Statistical Analysis
All the data were normally distributed and presented as mean±s.d. Statistical differences between groups were compared using one-way ANOVA (ANalyses Of VAriance between groups) software. Statistical significance was assumed at P<0.05.
Results
Experimental Strategy and Expected Secondary Structures of Target PCR Products
For diagnostic purposes, we first produced PCR products containing all the sequences informative for beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110 thalassemia mutations using, as target, genomic DNA from normal and heterozygous subjects or homozygous beta-thalassemia patients. Two double-heterozygous beta°39/beta°IVSI-1 patients were also analyzed. Figure 1 shows the location of the 5′-biotinylated biot-beta-glob-F and the beta-glob-R PCR primers within the human beta-globin gene. Table 1 indicates the nucleotide sequences of all the primers and probes used in this study.
Double-stranded target beta-globin gene PCR products were produced using an excess of the beta-glob-R primer with respect to the biotinylated biot-beta-glob-F primer. This was performed in order to minimize the presence of the unincorporated biotinylated PCR primer in the PCR mixture. The quality of the obtained PCR products was checked by agarose gel electrophoresis and direct DNA sequencing (data not shown).
After immobilization of PCR products, the four normal probes and the probes recognizing the four thalassemia mutations were sequentially injected as outlined in Figure 2a, and the RU bound determined.19, 20 Figure 2(b–f) shows the expected secondary structures of target PCR products without mutations (b) or carrying the indicated thalassemia mutations (c–f). The secondary structures were generated using the MFOLD software (version 3.1) developed by Zuker et al25 and Mathews et al.26 The analysis was performed at a temperature of 25°C and at 0.15 M NaCl, used in all the BIA experiments described in the present paper. The data obtained demonstrate that the biotinylated PCR products are, as expected, able to generate secondary structures. As clearly appreciable, and underlined by the arrows, portions of the sequences corresponding to the beta-thalassemia mutations considered are, at least in theory, involved in secondary structures. Therefore, we expect that the different probes employed could behave differently with respect to hybridization efficiency and stability of the generated hybrids.15 This cannot be avoided and could introduce a difficulty in the SPR-based diagnosis. In any case, we expect that the probes recognizing beta-thalassemia mutations will either be more efficient than the normal probes in hybridizing with full matched target sequences (for instance, in hybridizing with PCR products from homozygous thalassemia patients), or will generate more stable hybrids, as reported by our group for SPR-based analysis of single-point mutations.8, 10 Furthermore, the same probe (specific for either normal or mutated sequence) might behave differently if injected in flow cells carrying PCR products with different mutations and, therefore, different secondary structures.
(a) SPR-based format employed for molecular diagnosis. After immobilization of PCR products, normal and mutated probes were sequentially injected and the RU bound were determined. (b–f) Expected secondary structures of target PCR products without mutations (b) or carrying beta°39 (c), beta°IVSI-1 (d), beta+IVSI-6 (e), and beta+IVSI-110 (f) thalassemia mutations. The secondary structures were generated using the MFOLD software (version 3.1) developed by Zuker et al25 and Mathews et al.26 The analysis was performed at a temperature of 25°C and at 0.15 M NaCl.
Immobilization of Target PCR Product and Hybridization Analysis
Before injection on SA sensor chips, the final PCR products were purified by Microcon®-30. Figure 3a shows a representative example of the immobilization of biotinylated biot-beta-glob-F/beta-glob-R PCR products on an SA sensor chip. As it is evident, the binding kinetics are slow (Figure 3a, segment ‘a’ of the sensorgram); after 40 μl pulse of the biotinylated PCR product (approximately 1 μg/40 μl), no plateau levels of RU are reached. In any case, as expected, the regeneration step with buffer containing 40 mM NaOH, 1 M NaCl (segment ‘c’ of the sensorgram) induces a decrease in the RUfin to about 1/2, the RUres being still present on the flow cell due to the ssPCR product stably immobilized on the sensor chip (arrowed in Figure 3a). Repeated injections of biotinylated PCR product were performed in order to reach saturation of streptavidin binding sites of the flow cells. We usually obtained 1000–2000 RU of target PCR products immobilized on the sensor chip.
(a) Injection on an SA sensor chip of a biotinylated PCR product obtained using genomic DNA from a homozygous beta°IVSI-1 patient as target. Segment ‘a’ of the sensorgram: association; segment ‘b’ of the sensorgram: dissociation obtained injecting HBS-EP buffer; and segment ‘c’ of the sensorgram: regeneration step with buffer containing 40 mM NaOH, 1 M NaCl. (b) Hybridization of the M-beta(I)1 probe (solid line) to immobilized PCR and identification of RUi, RUfin, and RUres values. Dotted line: blank sensorgram obtained after injection of HBS-EP/H2O without oligonucleotide probes. (c) Final sensorgram obtained after subtraction of the blank sensorgram (dotted line of panel b) to the hybridization sensorgram (solid line of panel b).
Figure 3b shows the hybridization of the M-beta(I)1 probe to immobilized PCR product from a homozygous subject carrying this mutation, and shows the definitions of RUi, RUfin, and RUres values. The binding (solid line of b) is fast, but the RUfin obtained are low (in the representative example shown in b, 57 RUfin and 45 RUres are obtained). In this case, background subtraction of blank samples (dotted line of b) (HBS-EP/H2O without oligonucleotide probes) are required to take into consideration possible bulk effects that can affect the sensorgram. Subtraction of the blank sensorgram (dotted line of b) to the hybridization sensorgram (solid line of b) gives the final sensorgram shown in Figure 3c, leading to RUfin and RUres values of 54±2 and 39±1.5, respectively, in three consecutive independent injections. This procedure was employed for all the hybridization experiments reported in this paper.
Hybridization between Normal and Mutated ODN Probes and Biotinylated PCR Products Immobilized on an SA Sensor Chip
Figure 4(a–d) shows the binding efficiency of the normal (dotted lines) and mutated (solid lines) ODN probes corresponding to the beta°39 (a), beta°IVSI-1 (b), beta+IVSI-6 (c), and beta+IVSI-110 (d) thalassemia mutations to biotinylated PCR products from a normal subject.
The results demonstrate that the beta39, beta(I)1, beta(I)6, and beta(I)110 normal and mutated probes are all suitable to define the PCR product immobilized as a ‘normal’ product lacking mutations. This conclusion is drawn by looking to (RUfin−RUi) values for the beta°39 (a), beta°IVSI-1 (b) and beta+IVSI-110 (d) thalassemia mutations. For these mutations, the (RUres−RUi) values are also informative. By contrast, analysis of (RUfin−RUi) values are not informative for the beta+IVSI-6 mutation (c), since hybridization is obtained with both N-beta(I)6 and M-beta(I)6 probes. In this specific case, the analysis of (RUres–RUi) values is necessary for discriminating normal from mutated sequences.
In any case, the data of Figure 4(a–d) indicate that injections of normal and mutated probes to a single large PCR product containing the sites for beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110 thalassemia mutations give informative results for all the mutations when the (RUres−RUi) values are analyzed.
Therefore, we performed sequential injections of the same four normal and mutated ODN probes on immobilized PCR products from heterozygous subjects or homozygous beta-thalassemia patients. In addition, double heterozygous beta°39/beta°IVSI-1 patients were also analyzed.
Figure 4(e–n) shows representative examples obtained. In the first example, M-beta39, N-beta39, M-beta(I)1, and N-beta(I)1 probes were injected into a flow cell carrying a PCR product from a homozygous beta°39 patient (e and f). In the second example, the same probes were injected into a sensor chip carrying a PCR product from a homozygous beta°IVSI-1 patient (g and h).
As expected, only M-beta39 (e) and N-beta(I)1 (f) probes hybridize to the flow cell carrying a PCR product from a homozygous beta°39 patient; accordingly, only N-beta39 (g) and M-beta(I)1 (h) probes hybridize to the flow cell carrying a PCR product from a homozygous beta°IVSI-1 patient.
In the third representative example shown in Figure 4 (see panels i–n), analysis was conducted on a flow cell carrying a PCR product from a double-heterozygous beta°39/beta°IVSI-1 patient. In this case, the probes able to hybridize are M-beta39 and N-beta39 (i), M-beta(I)1 and N-beta(I)1 (l), N-beta(I)6 (m, dotted line), and N-beta(I)110 (n, dotted line). Accordingly, M-beta(I)6 (m, solid line) and M-beta(I)110 (n, solid line) probes do not hybridize.
The SPR-based BIA Protocol is Suitable for Molecular Diagnosis of the Beta°39, Beta°IVSI-1, Beta+IVSI-6 and Beta+IVSI-110 Thalassemia Mutations
In order to verify whether the experimental protocol described is suitable for reproducible molecular diagnosis of all the four thalassemia mutations analyzed, we determined the ‘beta-thalassemia index’ (beta-THAL index) as the value (RUres−RUi) (N)/(RUres−RUi)(M), where (RUres−RUi)(N) are the values obtained with the normal N-beta probes and (RUres−RUi)(M) values (in any case corrected to values ≥1) are those obtained with the M-beta probes recognizing the thalassemia mutations. The beta-THAL index was always found to be high (>2.5 for beta°39, >8 for beta°IVSI-1, >2 for beta+IVSI-6, and >5 for beta+IVSI-110) when PCR products from normal subjects were employed. On the contrary, this value was found to be intermediate when PCR products from heterozygous subjects were used. Finally, the beta-THAL index was always found to be lower than 0.6 (<0.5 for beta°39, <0.55 for beta°IVSI-1, <0.16 for beta+IVSI-6, and <0.22 for beta+IVSI-110) when PCR products from homozygous thalassemia patients were immobilized on the SA sensor chip. These data are analytically shown in Figure 5 and summarized in Table 2. The results obtained allow to conclude that the employed normal and mutated probes are useful for real-time SPR-based molecular diagnosis of beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110.
Statistical analyses demonstrate that all the differences between the beta-THAL index of normal and heterozygous subjects and homozygous thalassemia patients were significant in all cases (Figure 5).
Discussion
Recent papers demonstrated the possible use of BIA by SPR and biosensor technologies as an excellent approach for the molecular diagnosis of mutations involved in hereditary diseases. 8, 9, 10 However, to our knowledge, no report was published on the parallel diagnosis of multiple mutations. This is a very important feature of diagnosis protocols, since in most cases, hereditary diseases involving point mutations are highly heterogeneous. Moreover, double-heterozygous subjects are often present in the population.
In this paper, we employed an SPR-based BIA format for the analysis of multiple mutations within large PCR products immobilized on a sensor chip. In this format, the target PCR product is immobilized on the sensor chip, and then probes for detecting different mutations (in our case beta°39, beta°IVSI-1, beta+IVSI-6, and beta+IVSI-110 thalassemia mutations) are sequentially injected (Figure 2a).
The major problem we encountered is that during the association phase carried out with HBS-EP at 25°C, discrimination between mismatched and full matched probes/DNA hybrids is not readily and reproducibly observed with all the beta39, beta(I)1, beta(I)6, and beta(I)110 probes by analyzing the association phases of the obtained sensorgrams. However, (RUres−RUi) values were always found to be suitable for diagnostic purposes.
In this study, a total of ten normal and seven heterozygous subjects, and six homozygous thalassemia patients were considered (details are shown in Table 2) for a total of 11, 30, 29, and 9 determinations using beta39, beta(I)1, beta(I)6, and beta(I)110 probes, respectively. We emphasize that unexpected beta-THAL index values were never found in our analyses, indicating that this is a very reproducible and informative technique.
As we have studied relatively few samples, this paper should be considered as a pilot study; our data, however, firmly demonstrate the ability of SPR-based BIA of detecting multiple point mutations of the beta-globin gene by real-time monitoring of hybridization between oligonucleotide probes and target biotinylated PCR products generated from genomic DNA of normal and heterozygous subjects and homozygous beta-thalassemia patients.
The results reported in the present paper represent, in our opinion, a novel finding, since the procedure described to our knowledge is the first reported on the use of SPR-based BIA and biosensor technology for multiple detections of point mutations. The method is a real-time, very fast procedure of great interest for molecular preimplantation diagnosis, in order to discriminate homozygous thalassemic embryos from heterozygous and normal embryos. In this case, the speed of the analysis is critical in order to minimize the production of embryos to be successfully tested and implanted.27 Other advantages of the methodology described in the present paper are: (a) that it is a nonradioactive methodology and (b) that gel electrophoresis and/or dot-spot analysis are not required.
Three issues should be, in our opinion, investigated in the near future: the first is related to the efficiency of hybridization and amount of RU obtained for single hybridization. The development of modified probes able to generate higher RU shifts following hybridization is highly recommended. The second is to verify whether the approach described here could be applied to entire genes. A very interesting possibility is to perform a large PCR amplifying the entire beta-globin gene and to immobilize this large PCR product on the sensor chip, being able to perform molecular diagnosis for all the known mutations causing beta-thalassemia. We do not know what the limits of this technology are with respect to the size of the PCR product to be immobilized; this point deserves great attention in the future in order to propose this method for detection of point mutations affecting large-sized genes. A third issue to be investigated is the possibility of applying the described approach to other genetic diseases caused by point mutations, such as, for example, hemochromatosis and cystic fibrosis.
References
Johnsson U, Fagerstam L, Ivarsson B, et al. Real-time biospecific interaction analysis using surface plasmon resonance and a sensor chip technology. BioTechniques 1991;11:620–627.
Vadgama P, Crump PW . Biosensors: recent trends, a review. Analyst 1992;117:1657–1670.
Malmqvist M . Biospecific interactions analysis using biosensor technology. Nature 1993;361:186–187.
Wood SJ . DNA–DNA hybridization in real time using BIAcore. Microchemical J 1993;47:330–337.
Bianchi N, Rutigliano C, Tomassetti M, et al. Biosensor technology and surface plasmon resonance for real-time detection of HIV-1 genomic sequences amplified by polymerase chain reaction. Clin Diagn Virol 1997;8:199–208.
Nilsson P, Persson B, Larsson A, et al. Detection of mutations in PCR products from clinical samples by surface plasmon resonance. J Mol Recogn 1997;10:7–17.
Wang J, Rivas G, Cai X . Detection of point mutation in the p53 gene using peptide nucleic acid biosensor. Anal Chim Acta 1997;344:111–118.
Feriotto G, Lucci M, Bianchi N, et al. Detection of the ΔF508 (F508del) mutation of the cystic fibrosis gene by surface plasmon resonance and biosensor technology. Hum Mutat 1999;13:390–400.
Feriotto G, Corradini R, Sforza S, et al. Peptide nucleic acids and biosensor technology for real-time detection of the cystic fibrosis W1282X mutation by surface plasmon resonance. Lab Invest 2001;81:1415–1427.
Feriotto G, Ferlini A, Ravani A, et al. Biosensor technology for real-time detection of the cystic fibrosis W1282X mutation in CFTR. Hum Mutat 2001;18:70–81.
Saiki RK, Scharf S, Faloona F, et al. Enzymatic amplification of β-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 1985;230:1350–1354.
Nielsen PE, Egholm M, Berg RH, et al. Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 1991; 254:1497–1500.
Hyrup B, Nielsen PE . Peptide nucleic acids (PNA): synthesis, properties and potential applications. Bioorg Med Chem 1996;4:5–23.
Nielsen PE, Egholm M . Peptide Nucleic Acids: Protocols and Applications. Horizon Scientific Press: Norfolk, England, 1999.
Feriotto G, Gambari R . Surface plasmon resonance based biosensor technology for real-time detection of PCR products. In: Weissensteiner T, Griffin HG, Griffin A (eds). PCR Technology: Current Innovations. CRC Press: Boca Raton, FL, 2003, pp 141–154.
Chehab FF, Honig GR, Kan YW . Spontaneous mutation in beta-thalassaemia producing the same nucleotide substitution as that in a common hereditary form. Lancet 1986;1:3–5.
Rosatelli MC, Dozy A, Faa V, et al. Molecular characterization of beta-thalassemia in the Sardinian population. Am J Hum Genet 1992;50:422–426.
Feriotto G, Breveglieri G, Gardenghi S, et al. Surface plasmon resonance and biosensor technology for real-time molecular diagnosis of β°39 thalassemia mutation. Mol Diagn 2004, in press.
Feriotto G, Borgatti M, Mischiati C, et al. Biosensor technology and surface plasmon resonance for real-time detection of genetically modified Roundup Ready soybean gene sequences. J Agric Food Chem 2002;50:955–962.
Feriotto G, Gardenghi S, Bianchi N, et al. Quantitation of Bt-176 maize genomic sequences by surface plasmon resonance-based biospecific interaction analysis of multiplex polymerase chain Reaction (PCR). J Agric Food Chem 2003;51:4640–4646.
Orkin SH, Kazazian Jr HH, Antonarakis SE, et al. Linkage of beta-thalassaemia mutations and beta-globin gene polymorphisms with DNA polymorphisms in human beta-globin gene cluster. Nature 1982;296:627–631.
Spritz RA, Jagadeeswaran P, Choudary PV, et al. Base substitution in an intervening sequence of a beta-plus-thalassemic human globin gene. Proc Natl Acad Sci 1981;78:2455–2459.
Westaway D, Williamson R . An intron nucleotide sequence variant in a cloned beta-plus-thalassaemia globin gene. Nucleic Acids Res 1981;9:1777–1788.
Leblond-Francillard M, Dreyfus M, Rougeon F . Isolation of DNA–protein complexes based on streptavidin and biotin interaction. Eur J Biochem 1987;166:351–355.
Zuker M, Mathews DH, Turner DH . Algorithms and thermodynamics for RNA secondary structure prediction: a practical guide. In: Barciszewski J, Clark BFC (eds). RNA Biochemistry and Biotechnology. NATO ASI Series. Kluwer Academic Publishers: Dordrecht, 1999, pp. 11–43.
Mathews DH, Sabina J, Zuker M, et al. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J Mol Biol 1999;288:911–940.
Ao A, Ray P, Harper J . Clinical experience with preimplantation genetic diagnosis of cystic fibrosis (delta F508). Prenat Diagn 1996;16:137–142.
Acknowledgements
This research was supported by the Ministero della Sanità, Italy, by COFIN-2002, by FIRB-2001, and by Associazione Veneta per la Lotta alla Talassemia (AVLT). SG was supported by a fellowship from the Associazione per la Lotta alla Talassemia di Ferrara.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Feriotto, G., Breveglieri, G., Finotti, A. et al. Real-time multiplex analysis of four beta-thalassemia mutations employing surface plasmon resonance and biosensor technology. Lab Invest 84, 796–803 (2004). https://doi.org/10.1038/labinvest.3700106
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700106
Keywords
- biosensors
- beta-thalassemia
- molecular diagnosis
- polymerase chain reaction
- real-time assays
- surface plasmon resonance
Keywords
This article is cited by
-
Encapsulation of eukaryotic cells in alginate microparticles: cell signaling by TNF-alpha through capsular structure of cystic fibrosis cells
Journal of Cell Communication and Signaling (2011)
-
Label-free detection of DNA mutations by SPR: application to the early detection of inherited breast cancer
Analytical and Bioanalytical Chemistry (2009)
-
Plasmonic DNA: Towards Genetic Diagnosis Chips
Plasmonics (2007)
-
Inside Lab Invest
Laboratory Investigation (2004)