Featured
-
-
Article
| Open Access The UK10K project identifies rare variants in health and disease
The UK10K project identifies rare variants in health and disease
Low read depth sequencing of whole genomes and high read depth exomes of nearly 10,000 extensively phenotyped individuals are combined to help characterize novel sequence variants, generate a highly accurate imputation reference panel and identify novel alleles associated with lipid-related traits; in addition to describing population structure and providing functional annotation of rare and low-frequency variants the authors use the data to estimate the benefits of sequencing for association studies.
- Klaudia Walter
- , Josine L. Min
- & Weihua Zhang
-
Letter |
 NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs
NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs
A newly developed method, NAD captureSeq, has been used to show that bacteria cap the 5′-ends of some RNAs to protect against degradation, much as happens with eukaryotic messenger RNAs, although with a different modification: nicotinamide adenine dinucleotide.
- Hana Cahová
- , Marie-Luise Winz
- & Andres Jäschke
-
Letter |
 Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction
Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction
Exome sequence analysis of nearly 10,000 people was carried out to identify alleles associated with early-onset myocardial infarction; mutations in low-density lipoprotein receptor (LDLR) or apolipoprotein A-V (APOA5) were associated with disease risk, identifying the key roles of low-density lipoprotein cholesterol and metabolism of triglyceride-rich lipoproteins.
- Ron Do
- , Nathan O. Stitziel
- & Sekar Kathiresan
-
Letter |
 Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia
Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia
Somatic TP53 mutations are highly prevalent in therapy-related acute myeloid leukaemia and myelodysplastic syndrome, which arise as complications of cytotoxic chemotherapy or radiotherapy; although it was believed that these TP53 mutations are directly induced by cytotoxic therapy, new data indicate that they predate cytotoxic therapy and that haematopoietic progenitors harbouring these pre-existing mutations may selectively expand after exposure to chemotherapy or radiotherapy.
- Terrence N. Wong
- , Giridharan Ramsingh
- & Richard K. Wilson
-
Article |
 Synaptic, transcriptional and chromatin genes disrupted in autism
Synaptic, transcriptional and chromatin genes disrupted in autism
Whole-exome sequencing in a large autism study identifies over 100 autosomal genes that are likely to affect risk for the disorder; these genes, which show unusual evolutionary constraint against mutations, carry de novo loss-of-function mutations in over 5% of autistic subjects and many function in synaptic, transcriptional and chromatin-remodelling pathways.
- Silvia De Rubeis
- , Xin He
- & Joseph D. Buxbaum
-
Letter |
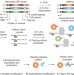 Multiplex single-molecule interaction profiling of DNA-barcoded proteins
Multiplex single-molecule interaction profiling of DNA-barcoded proteins
Single-molecular-interaction-sequencing involves attaching DNA barcodes to proteins, assaying these barcoded proteins en masse in an aqueous solution, followed by immobilization in a polyacrylamide film and amplifying and analysing the barcoding DNAs—the method allows for precise protein quantification and simultaneous interrogation of molecular binding affinity and specificity.
- Liangcai Gu
- , Chao Li
- & George M. Church
-
Article
| Open Access Gibbon genome and the fast karyotype evolution of small apes
Gibbon genome and the fast karyotype evolution of small apes
The genome of the gibbon, a tree-dwelling ape from Asia positioned between Old World monkeys and the great apes, is presented, providing insights into the evolutionary history of gibbon species and their accelerated karyotypes, as well as evidence for selection of genes such as those for forelimb development and connective tissue that may be important for locomotion through trees.
- Lucia Carbone
- , R. Alan Harris
- & Richard A. Gibbs
-
Letter |
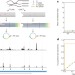 Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
The modification of uridine to pseudouridine is widespread in transfer and ribosomal RNAs but not observed so far in a coding RNA; here a new technique is used to detect this modification on a genome-wide scale, leading to the identification of pseudouridylation in messenger RNAs as well as almost 100 new sites in non-coding RNAs.
- Thomas M. Carlile
- , Maria F. Rojas-Duran
- & Wendy V. Gilbert
-
Letter |
 Genome sequencing identifies major causes of severe intellectual disability
Genome sequencing identifies major causes of severe intellectual disability
Whole-genome sequencing is used to identify genetic alterations in patients with severe intellectual disability for whom all other tests, including array and exome sequencing, returned negative results; de novo single-nucleotide and copy number variations affecting the coding region seem to be a major cause of this disorder.
- Christian Gilissen
- , Jayne Y. Hehir-Kwa
- & Joris A. Veltman
-
Letter |
 Novel somatic and germline mutations in intracranial germ cell tumours
Novel somatic and germline mutations in intracranial germ cell tumours
Intracranial germ cell tumours are rare tumours affecting mainly male adolescents, mainly in Asia; here the authors identify frequent mutations in the KIT/RAS and AKT/mTOR signalling pathways as well as rare germline variants in JMJD1C, suggesting potential therapeutic strategies focusing on the inhibition of KIT/RAS activation and the AKT1/mTOR pathway.
- Linghua Wang
- , Shigeru Yamaguchi
- & Ching C. Lau
-
Article |
 A draft map of the human proteome
A draft map of the human proteome
A draft map of the human proteome is presented here, accounting for over 80% of the annotated protein-coding genes in humans; some novel protein-coding regions, including translated pseudogenes, non-coding RNAs and upstream open reading frames, are identified.
- Min-Sik Kim
- , Sneha M. Pinto
- & Akhilesh Pandey
-
Article |
 Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators
Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators
A study comparing the Y chromosome across mammalian species reveals that selection to maintain the ancestral dosage of homologous X–Y gene pairs preserved a handful of genes on the Y chromosome while the rest were lost; the survival of broadly expressed dosage-sensitive regulators of gene expression suggest that the human Y chromosome is essential for male viability.
- Daniel W. Bellott
- , Jennifer F. Hughes
- & David C. Page
-
Article |
 Poly(A)-tail profiling reveals an embryonic switch in translational control
Poly(A)-tail profiling reveals an embryonic switch in translational control
A new high-throughput sequencing method to determine mRNA poly(A)-tail length enabled studies of individual RNAs across species and developmental stages to investigate the role of poly(A) length in translational regulation; the relationship between poly(A) length and translational efficiency shown in early embryo systems does not occur later in development, a finding that explains different regulatory consequences of microRNAs acting at different developmental times.
- Alexander O. Subtelny
- , Stephen W. Eichhorn
- & David P. Bartel
-
Article |
 De novo mutations in schizophrenia implicate synaptic networks
De novo mutations in schizophrenia implicate synaptic networks
The authors report the largest family-trio exome sequencing study of schizophrenia to date; mutations are overrepresented in genes for glutamatergic synaptic proteins and also genes mutated in autism and intellectual disability, providing insights into aetiological mechanisms and pathopshyisology shared with other neurodevelopmental disorders.
- Menachem Fromer
- , Andrew J. Pocklington
- & Michael C. O’Donovan
-
Article |
 A polygenic burden of rare disruptive mutations in schizophrenia
A polygenic burden of rare disruptive mutations in schizophrenia
Exome sequence analysis of more than 5,000 schizophrenia cases and controls identifies a polygenic burden primarily arising from rare, disruptive mutations distributed across many genes, among which are those encoding voltage-gated calcium ion channels and the signalling complex formed by the ARC protein of the postsynaptic density; as in autism, mutations were also found in homologues of known targets of the fragile X mental retardation protein.
- Shaun M. Purcell
- , Jennifer L. Moran
- & Pamela Sklar
-
Letter |
 Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo
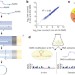
Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo
Understanding how RNA structure influences its function has been hampered by a lack of approaches that can accurately quantify RNA structure in vivo; here, RNA structure is revealed on a global scale and with nucleotide-level resolution, showing that there is less structure within cells than expected from in vitro and in silico analyses.
- Silvi Rouskin
- , Meghan Zubradt
- & Jonathan S. Weissman
-
Letter |
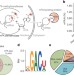 N6-methyladenosine-dependent regulation of messenger RNA stability
N6-methyladenosine-dependent regulation of messenger RNA stability
The mRNAs of higher eukaryotes are extensively modified internally with N6-methyladenosine, but the specific functional role of this modification has been unclear; here this modification on mRNA is shown to be recognized by several proteins, the modification and its recognition serve to regulate the RNA’s lifetime.
- Xiao Wang
- , Zhike Lu
- & Chuan He
-
Article |
 Transcriptome and genome sequencing uncovers functional variation in humans
Transcriptome and genome sequencing uncovers functional variation in humans
Sequencing and deep analysis of mRNA and miRNA from lymphoblastoid cell lines of 462 individuals from the 1000 Genomes Project reveal widespread genetic variation affecting the regulation of most genes, with transcript structure and expression level variation being equally common but genetically largely independent, and the analyses point to putative causal variants for dozens of disease-associated loci.
- Tuuli Lappalainen
- , Michael Sammeth
- & Emmanouil T. Dermitzakis
-
Letter |
 The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line
The haplotype-resolved genome and epigenome of the aneuploid HeLa cancer cell line
Haplotype-resolved whole-genome sequencing of the HeLa CCL-2 strain shows that HeLa is relatively stable in terms of point variation; integration of several data sets reveals strong, haplotype-specific activation of the proto-oncogene MYC by the human papilloma virus type 18 genome, and enables the relationship between gene dosage and expression to be examined.
- Andrew Adey
- , Joshua N. Burton
- & Jay Shendure
-
Article
| Open Access Comprehensive molecular characterization of clear cell renal cell carcinoma
Comprehensive molecular characterization of clear cell renal cell carcinoma
The Cancer Genome Atlas Research Network reports an integrative analysis of more than 400 samples of clear cell renal cell carcinoma based on genomic, DNA methylation, RNA and proteomic characterisation; frequent mutations were identified in the PI(3)K/AKT pathway, suggesting this pathway might be a potential therapeutic target, among the findings is also a demonstration of metabolic remodelling which correlates with tumour stage and severity.
- Chad J. Creighton
- , Margaret Morgan
- & Heidi J. Sofia.
-
Letter |
 Negligible impact of rare autoimmune-locus coding-region variants on missing heritability
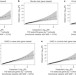
Negligible impact of rare autoimmune-locus coding-region variants on missing heritability
A search for variants in coding exons of 25 genome-wide association study risk genes in a large cohort of autoimmune patients finds that rare coding-region variants at known loci have a negligible role in common autoimmune disease susceptibility, arguing against the previously proposed rare-variant synthetic genome-wide association hypothesis.
- Karen A. Hunt
- , Vanisha Mistry
- & David A. van Heel
-
Letter |
 Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell RNA sequencing is used to investigate the transcriptional response of 18 mouse bone-marrow-derived dendritic cells after lipopolysaccharide stimulation; many highly expressed genes, such as key immune genes and cytokines, show bimodal variation in both transcript abundance and splicing patterns. This variation reflects differences in both cell state and usage of an interferon-driven pathway involving Stat2 and Irf7.
- Alex K. Shalek
- , Rahul Satija
- & Aviv Regev
-
Letter |
 Extensive transcriptional heterogeneity revealed by isoform profiling
Extensive transcriptional heterogeneity revealed by isoform profiling
Variation among RNA transcript isoforms can be generated from alternative start and polyadenylation sites, and results in RNAs and proteins with different properties being generated from the same genomic sequence; here a new method termed transcript isoform sequencing is described in yeast, and the method allows a fuller exploration of transcriptome diversity across the compact yeast genome.
- Vicent Pelechano
- , Wu Wei
- & Lars M. Steinmetz
-
Letter |
 Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA
Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA
A proof of principle study shows that by exome sequencing of cell-free circulating DNA from cancer patient plasma samples, the genomic evolution of metastatic cancers and the acquisition of resistance in response to therapy can be tracked over time.
- Muhammed Murtaza
- , Sarah-Jane Dawson
- & Nitzan Rosenfeld
-
Letter |
 Discrete genetic modules are responsible for complex burrow evolution in Peromyscus mice
Discrete genetic modules are responsible for complex burrow evolution in Peromyscus mice
The complex burrows created by oldfield mice are shown to be governed by genetic modules that each control an aspect of burrow size or shape.
- Jesse N. Weber
- , Brant K. Peterson
- & Hopi E. Hoekstra
-
News |
 Method offers DNA blueprint of a single human cell
Method offers DNA blueprint of a single human cell
Easier cell-to-cell comparisons hold potential insights into cancer and other biological processes.
- Monya Baker
-
News |
 Past 5,000 years prolific for changes to human genome
Past 5,000 years prolific for changes to human genome
High-resolution sequencing study emphasizes importance of rare variants in disease.
- Nidhi Subbaraman
-
Article
| Open Access Analysis of the bread wheat genome using whole-genome shotgun sequencing
Analysis of the bread wheat genome using whole-genome shotgun sequencing
Sequencing of the hexaploid bread wheat genome shows that it is highly dynamic, with significant loss of gene family members on polyploidization and domestication, and an abundance of gene fragments.
- Rachel Brenchley
- , Manuel Spannagl
- & Neil Hall
-
News |
 DNA sequencers stymie superbug spread
DNA sequencers stymie superbug spread
Whole-genome analysis helps identify source of MRSA outbreak on infant ward
- Ewen Callaway
-
Article
| Open Access Analyses of pig genomes provide insight into porcine demography and evolution
Analyses of pig genomes provide insight into porcine demography and evolution
This study presents the assembly and analysis of the genome sequence of a female domestic Duroc pig and a comparison with the genomes of wild and domestic pigs from Europe and Asia; the results shed light on the evolutionary relationship between European and Asian wild boars.
- Martien A. M. Groenen
- , Alan L. Archibald
- & Lawrence B. Schook
-
News Feature |
 Genomics: The single life
Genomics: The single life
Sequencing DNA from individual cells is changing the way that researchers think of humans as a whole.
- Brian Owens
-
Article
| Open Access An integrated map of genetic variation from 1,092 human genomes
An integrated map of genetic variation from 1,092 human genomes
This report from the 1000 Genomes Project describes the genomes of 1,092 individuals from 14 human populations, providing a resource for common and low-frequency variant analysis in individuals from diverse populations; hundreds of rare non-coding variants at conserved sites, such as motif-disrupting changes in transcription-factor-binding sites, can be found in each individual.
- Gil A. McVean
- , David M. Altshuler (Co-Chair)
- & Gil A. McVean
-
News |
 Genome interpreter vies for place in clinical market
Genome interpreter vies for place in clinical market
Launch of system that keeps data local aims to address privacy fears.
- Monya Baker
-
News |
 Two-day test diagnoses genetic diseases in newborns
Two-day test diagnoses genetic diseases in newborns
Platform could be a model for routine whole-genome sequencing in neonatal intensive care.
- Monya Baker
-
News |
 China buys US sequencing firm
China buys US sequencing firm
BGI’s rescue of Complete Genomics will keep a valued technology afloat.
- Monya Baker
-
-
News |
 New DNA analysis shows ancient humans interbred with Denisovans
New DNA analysis shows ancient humans interbred with Denisovans
A new high-coverage DNA sequencing method reconstructs the full genome of Denisovans — relatives to both Neandertals and humans — from genetic fragments in a single finger bone.
- Katherine Harmon
-
Letter |
 Defining the core Arabidopsis thaliana root microbiome
Defining the core Arabidopsis thaliana root microbiome
Sequencing of the Arabidopsis thaliana root microbiome shows that its composition is strongly influenced by location, inside or outside the root, and by soil type.
- Derek S. Lundberg
- , Sarah L. Lebeis
- & Jeffery L. Dangl
-
News |
 Hunter-gatherer genomes a trove of genetic diversity
Hunter-gatherer genomes a trove of genetic diversity
Sequences from African groups offer genetic clues about disease, height and ancient human breeding.
- Ewen Callaway
-
News |
 Contest to sequence centenarians kicks off
Contest to sequence centenarians kicks off
First entrant pins hopes on semiconductor technology.
- Monya Baker
-
Letter |
 Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations
Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations
Medulloblastoma is the most common brain tumour in children; using exome sequencing of tumour samples the authors show that these cancers have low mutation rates and identify 12 significantly mutated genes, among them the gene encoding RNA helicase DDX3X.
- Trevor J. Pugh
- , Shyamal Dilhan Weeraratne
- & Yoon-Jae Cho
-
News & Views |
 Fetal genes in mother's blood
Fetal genes in mother's blood
The genome sequence of a fetus can be inferred from the relative numbers of variants of DNA sequences in a pregnant woman's blood. This advance in non-invasive diagnostics comes with some ramifications. See Article p.320
- Diana W. Bianchi
-
Article
| Open Access Comprehensive molecular characterization of human colon and rectal cancer
Comprehensive molecular characterization of human colon and rectal cancer
The Cancer Genome Atlas consortium reports on their genome-wide characterization of somatic alterations in colorectal cancer; in addition to revealing a remarkably consistent pattern of genomic alteration, with 24 genes being significantly mutated, the study identifies new targets for therapeutic intervention and suggests an important role for MYC-directed transcriptional activation and repression.
- Donna M. Muzny
- , Matthew N. Bainbridge
- & Elizabeth Thomson.
-
News Feature |
 Microbes en masse: The sequencing machine
Microbes en masse: The sequencing machine
Faeces, lizards, keyboards, faces — Rob Knight likes to sequence the microbes on anything and everything. Next, he plans to sequence Earth.
- Virginia Gewin
-
Article
| Open Access Accurate whole-genome sequencing and haplotyping from 10 to 20 human cells
Accurate whole-genome sequencing and haplotyping from 10 to 20 human cells
A new DNA analysis method termed long fragment read technology is described, and the approach is used to determine parental haplotypes and to sequence human genomes cost-effectively and accurately from only 10 to 20 cells.
- Brock A. Peters
- , Bahram G. Kermani
- & Radoje Drmanac
-
Article |
 Non-invasive prenatal measurement of the fetal genome
Non-invasive prenatal measurement of the fetal genome
Prenatal testing usually requires invasive sampling; here molecular counting of parental haplotypes in the maternal plasma allows the fetal genome to be deciphered and molecular counting of individual alleles enables the fetal exome to be captured.
- H. Christina Fan
- , Wei Gu
- & Stephen R. Quake
-
News |
 Correction algorithms extend the reach of genome sequencing
Correction algorithms extend the reach of genome sequencing
Latest sequencers combined with older technology improve accuracy of genome assemblies.
- Monya Baker
-
Letter |
 RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis
RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis
A new method allows the collection of circulating tumour cells (CTCs) despite their rarity; transcriptome sequencing of CTCs could allow identification of pathways involved in metastasis.
- Min Yu
- , David T. Ting
- & Daniel A. Haber
-
Research Highlights |
Sequencing tracks outbreak

 Whole‐genome sequencing identifies EN1 as a determinant of bone density and fracture
Whole‐genome sequencing identifies EN1 as a determinant of bone density and fracture Genomic approaches to studying the human microbiota
Genomic approaches to studying the human microbiota