Abstract
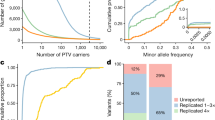
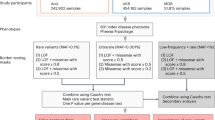
Genome-wide association studies (GWAS) have identified common variants of modest-effect size at hundreds of loci for common autoimmune diseases; however, a substantial fraction of heritability remains unexplained, to which rare variants may contribute1,2. To discover rare variants and test them for association with a phenotype, most studies re-sequence a small initial sample size and then genotype the discovered variants in a larger sample set3,4,5. This approach fails to analyse a large fraction of the rare variants present in the entire sample set. Here we perform simultaneous amplicon-sequencing-based variant discovery and genotyping for coding exons of 25 GWAS risk genes in 41,911 UK residents of white European origin, comprising 24,892 subjects with six autoimmune disease phenotypes and 17,019 controls, and show that rare coding-region variants at known loci have a negligible role in common autoimmune disease susceptibility. These results do not support the rare-variant synthetic genome-wide-association hypothesis6 (in which unobserved rare causal variants lead to association detected at common tag variants). Many known autoimmune disease risk loci contain multiple, independently associated, common and low-frequency variants, and so genes at these loci are a priori stronger candidates for harbouring rare coding-region variants than other genes. Our data indicate that the missing heritability for common autoimmune diseases may not be attributable to the rare coding-region variant portion of the allelic spectrum, but perhaps, as others have proposed, may be a result of many common-variant loci of weak effect7,8,9,10.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Accession codes
Data deposits
Genome data has been deposited at the European Genome–phenome Archive (http://www.ebi.ac.uk/ega/), which is hosted at the EBI, under accession number EGAS00001000476.
References
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009)
Gibson, G. Rare and common variants: twenty arguments. Nature Rev. Genet. 13, 135–145 (2012)
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009)
Rivas, M. A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nature Genet. 43, 1066–1073 (2011)
Momozawa, Y. et al. Resequencing of positional candidates identifies low frequency IL23R coding variants protecting against inflammatory bowel disease. Nature Genet. 43, 43–47 (2011)
Dickson, S. P., Wang, K., Krantz, I., Hakonarson, H. & Goldstein, D. B. Rare variants create synthetic genome-wide associations. PLoS Biol. 8, e1000294 (2010)
Bloom, J. S., Ehrenreich, I. M., Loo, W. T., Lite, T. L. & Kruglyak, L. Finding the sources of missing heritability in a yeast cross. Nature 494, 234–237 (2013)
Stahl, E. A. et al. Bayesian inference analyses of the polygenic architecture of rheumatoid arthritis. Nature Genet. 44, 483–489 (2012)
Park, J. H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nature Genet. 42, 570–575 (2010)
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nature Genet. 42, 565–569 (2010)
Coventry, A. et al. Deep resequencing reveals excess rare recent variants consistent with explosive population growth. Nature Commun. 1, 131 (2010)
Nelson, M. R. et al. An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 people. Science 337, 100–104 (2012)
Tennessen, J. A. et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337, 64–69 (2012)
Fu, W. et al. Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493, 216–220 (2013)
Strange, A. et al. A genome-wide association study identifies new psoriasis susceptibility loci and an interaction between HLA-C and ERAP1. Nature Genet. 42, 985–990 (2010)
Jordan, C. T. et al. Rare and common variants in CARD14, encoding an epidermal regulator of NF-kappaB, in psoriasis. Am. J. Hum. Genet. 90, 796–808 (2012)
Diogo, D. et al. Rare, low-frequency, and common variants in the protein-coding sequence of biological candidate genes from GWASs contribute to risk of rheumatoid arthritis. Am. J. Hum. Genet. 92, 15–27 (2013)
Lesage, S. et al. CARD15/NOD2 mutational analysis and genotype-phenotype correlation in 612 patients with inflammatory bowel disease. Am. J. Hum. Genet. 70, 845–857 (2002)
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nature Genet. 43, 1193–1201 (2011)
Barrett, J. C. et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nature Genet. 41, 703–707 (2009)
Jacob, C. O. et al. Lupus-associated causal mutation in neutrophil cytosolic factor 2 (NCF2) brings unique insights to the structure and function of NADPH oxidase. Proc. Natl Acad. Sci. USA 109, E59–E67 (2012)
Kolaczkowska, E. & Kubes, P. Neutrophil recruitment and function in health and inflammation. Nature Rev. Immunol. 13, 159–175 (2013)
Liu, D. J. & Leal, S. M. Estimating genetic effects and quantifying missing heritability explained by identified rare-variant associations. Am. J. Hum. Genet. 91, 585–596 (2012)
Cooper, J. D. et al. Seven newly identified loci for autoimmune thyroid disease. Hum Mol Genet 21, 5202–5208 (2012)
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012)
Sawcer, S. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011)
Tsoi, L. C. et al. Identification of 15 new psoriasis susceptibility loci highlights the role of innate immunity. Nat Genet 44, 1341–1348 (2012)
Dendrou, C. A. et al. Cell-specific protein phenotypes for the autoimmune locus IL2RA using a genotype-selectable human bioresource. Nat Genet 41, 1011–1015 (2009)
Kong, Y. Btrim: a fast, lightweight adapter and quality trimming program for next-generation sequencing technologies. Genomics 98, 152–153 (2011)
MacArthur, D. G. et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 335, 823–828 (2012)
Acknowledgements
The study was primarily funded by the Medical Research Council (MRC G1001158 to D.A.v.H and V.P.), with further funding from Coeliac UK (to D.A.v.H). We thank C. Wijmenga and G. Trynka for sharing ImmunoChip data, and the International Multiple Sclerosis Genomics Consortium for ImmunoChip data and samples. J.N.B. and R.C.T. are supported by MRC grant G0601387. This research was supported by the National Institutes for Health Research (NIHR) Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust and King’s College London. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. The study was supported by the Cambridge NIHR Biomedical Research Centre. We thank E. Gray and D. Jones (Wellcome Trust Sanger Institute) for sample preparation. We acknowledge use of DNA from The UK Blood Services collection of Common Controls (UKBS-CC collection), funded by the Wellcome Trust grant 076113/C/04/Z and by NIHR programme grant to NHS Blood and Transplant (RP-PG-0310-1002). The collection was established as part of the Wellcome Trust Case Control Consortium (WTCCC). We acknowledge use of DNA from the British 1958 Birth Cohort collection, funded by the UK MRC grant G0000934 and the Wellcome Trust grant 068545/Z/02. We thank nurses and doctors for recruiting autoimmune thyroid disease (AITD) subjects into the AITD National Collection, funded by the Wellcome Trust grant 068181. We acknowledge use of DNA from the Cambridge BioResource. We acknowledge use of DNA from the Juvenile Diabetes Research Foundation (JDRF)/Wellcome Trust Case-Series (GRID), funded by JDRF and the Wellcome Trust (grant references JDRF 4-2001-1008 and WT061858). The subjects were recruited in the UK by D. Dunger and his team with support from the British Society for Paediatric Endocrinology and Diabetes. The samples were prepared and provided by the JDRF/Wellcome Trust Diabetes and Inflammation Laboratory, University of Cambridge, UK. Psoriasis samples used were based on the WTCCC2 GWAS clinical panel, for which we thank D. Burden, C. Griffiths, M. Cork and R. McManus. Finally, we would like to thank all autoimmune disease and control subjects for participating in this study.
Author information
Authors and Affiliations
Contributions
D.A.v.H. designed and led the study. K.A.H. coordinated wet laboratory work, with K.A.H., V.M., N.A.B. and E.W. performing DNA sample preparation, Fluidigm PCR amplification, sample barcoding, MiSeq library validation and Sanger sequencing preparation. HiSeq sequencing was performed by M.M. and E.P. D.A.v.H., V.P. and M.S. performed bioinformatics and statistical analyses. All other authors contributed to diverse aspects of sample collection, phenotyping, DNA preparation, ImmunoChip data production or specific analyses. D.A.v.H. and V.P drafted the manuscript, which all authors reviewed.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Figures 1-3. (PDF 704 kb)
Supplementary Data
This file contains full results (for all tested phenotypes, all MAF, all single variants or pooled gene tests) for the data shown in Supplementary Table 4 and Figure 1. (XLSX 1278 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Hunt, K., Mistry, V., Bockett, N. et al. Negligible impact of rare autoimmune-locus coding-region variants on missing heritability. Nature 498, 232–235 (2013). https://doi.org/10.1038/nature12170
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12170
This article is cited by
-
The genetics of non-monogenic IBD
Human Genetics (2023)
-
Validation of genotype imputation in Southeast Asian populations and the effect of single nucleotide polymorphism annotation on imputation outcome
BMC Medical Genetics (2018)
-
A genomic perspective on HLA evolution
Immunogenetics (2018)
-
A pathway-centric approach to rare variant association analysis
European Journal of Human Genetics (2017)
-
The multiple pathways to autoimmunity
Nature Immunology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.