Featured
-
-
Article
| Open Access Spatial atlas of the mouse central nervous system at molecular resolution
Spatial atlas of the mouse central nervous system at molecular resolution
In situ spatial transcriptomic analysis of more than 1 million cells are used to create a 200-nm-resolution spatial molecular atlas of the adult mouse central nervous system and identify previously unknown tissue architectures.
- Hailing Shi
- , Yichun He
- & Xiao Wang
-
Article
| Open Access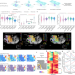 Spatial epigenome–transcriptome co-profiling of mammalian tissues
Spatial epigenome–transcriptome co-profiling of mammalian tissues
The authors present two technologies for spatially resolved, genome-wide, joint profiling of the epigenome and transcriptome by cosequencing chromatin accessibility and gene expression, or histone modifications and gene expression on the same tissue section at near-single-cell resolution.
- Di Zhang
- , Yanxiang Deng
- & Rong Fan
-
Article
| Open Access Broad transcriptomic dysregulation occurs across the cerebral cortex in ASD
Broad transcriptomic dysregulation occurs across the cerebral cortex in ASD
RNA sequencing reveals widespread transcriptomic changes across the cerebral cortex in autism spectrum disorder, including primary sensory regions, in addition to association regions, as well as an attenuation of regional identity.
- Michael J. Gandal
- , Jillian R. Haney
- & Daniel H. Geschwind
-
Article
| Open Access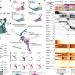 Inferring and perturbing cell fate regulomes in human brain organoids
Inferring and perturbing cell fate regulomes in human brain organoids
A multi-omic atlas of brain organoid development facilitates the inference of an underlying gene regulatory network using the newly developed Pando framework and shows—in conjunction with perturbation experiments—that GLI3 controls forebrain fate establishment through interaction with HES4/5 regulomes.
- Jonas Simon Fleck
- , Sophie Martina Johanna Jansen
- & Barbara Treutlein
-
Article |
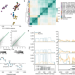 Transcriptome variation in human tissues revealed by long-read sequencing
Transcriptome variation in human tissues revealed by long-read sequencing
To understand the contribution of variants to transcript expression regulation, long-read transcriptome data are generated from the GTEx resource, and a new software package to perform allele-specific analysis is developed.
- Dafni A. Glinos
- , Garrett Garborcauskas
- & Beryl B. Cummings
-
Article
| Open Access Local and systemic responses to SARS-CoV-2 infection in children and adults
Local and systemic responses to SARS-CoV-2 infection in children and adults
Mechanisms explaining the milder clinical syndrome that is observed in children with SARS-CoV-2 infection.
- Masahiro Yoshida
- , Kaylee B. Worlock
- & Kerstin B. Meyer
-
Article |
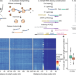 Single-cell Ribo-seq reveals cell cycle-dependent translational pausing
Single-cell Ribo-seq reveals cell cycle-dependent translational pausing
Highly sensitive ribosome profiling of single cells at single-codon resolution enables identification of distinct cell cycle-dependent translational dynamic states in individual cells.
- Michael VanInsberghe
- , Jeroen van den Berg
- & Alexander van Oudenaarden
-
Article |
 The kinetic landscape of an RNA-binding protein in cells
The kinetic landscape of an RNA-binding protein in cells
Time-resolved RNA–protein cross-linking with a pulsed femtosecond ultraviolet laser, followed by immunoprecipitation and high-throughput sequencing, enables the determination of binding and dissociation kinetics of the RNA-binding protein DAZL within cells.
- Deepak Sharma
- , Leah L. Zagore
- & Eckhard Jankowsky
-
Article |
 Physiological blood–brain transport is impaired with age by a shift in transcytosis
Physiological blood–brain transport is impaired with age by a shift in transcytosis
Tagging and tracking the blood plasma proteome as a discovery tool reveals widespread endogenous transport of proteins into the healthy brain and the pharmacologically modifiable mechanisms by which the brain endothelium regulates this process with age.
- Andrew C. Yang
- , Marc Y. Stevens
- & Tony Wyss-Coray
-
Article |
 Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping
Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping
A method termed ac4C-seq is introduced for the transcriptome-wide mapping of the RNA modification N4-acetylcytidine, revealing widespread temperature-dependent acetylation that facilitates thermoadaptation in hyperthermophilic archaea.
- Aldema Sas-Chen
- , Justin M. Thomas
- & Schraga Schwartz
-
Article |
 Single-cell and spatial transcriptomics reveal somitogenesis in gastruloids
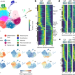
Single-cell and spatial transcriptomics reveal somitogenesis in gastruloids
Single-cell RNA sequencing and spatial transcriptomics reveal that the somitogenesis clock is active in mouse gastruloids, which can be induced to generate somites with the correct rostral–caudal patterning.
- Susanne C. van den Brink
- , Anna Alemany
- & Alexander van Oudenaarden
-
Article |
 Decoding human fetal liver haematopoiesis
Decoding human fetal liver haematopoiesis
Single-cell transcriptomic profiling of fetal liver, skin, kidney and yolk sac reveals the differentiation trajectories of human haematopoietic stem cells and multipotent progenitors, which are validated to produce an integrated map of fetal liver haematopoiesis.
- Dorin-Mirel Popescu
- , Rachel A. Botting
- & Muzlifah Haniffa
-
Letter |
 Single-cell analysis of cardiogenesis reveals basis for organ-level developmental defects
Single-cell analysis of cardiogenesis reveals basis for organ-level developmental defects
Single-cell RNA-sequencing analysis reveals functions of lineage-specifying transcription factors underlying congenital defects in heart development.
- T. Yvanka de Soysa
- , Sanjeev S. Ranade
- & Deepak Srivastava
-
Letter |
 scSLAM-seq reveals core features of transcription dynamics in single cells
scSLAM-seq reveals core features of transcription dynamics in single cells
A technique known as scSLAM-seq that combines single-cell RNA sequencing with metabolic RNA labelling and nucleoside conversion is used to study the onset of cytomegalovirus infection in single mouse fibroblasts.
- Florian Erhard
- , Marisa A. P. Baptista
- & Lars Dölken
-
Article |
 Charting cellular identity during human in vitro β-cell differentiation
Charting cellular identity during human in vitro β-cell differentiation
Single-cell transcriptional profiling of in vitro human pancreatic β-cell differentiation reveals progenitor and terminal fates, produces a detailed time course of endocrine induction and underpins a lineage model.
- Adrian Veres
- , Aubrey L. Faust
- & Douglas A. Melton
-
Letter
| Open Access The impact of rare variation on gene expression across tissues
The impact of rare variation on gene expression across tissues
The authors show that rare genetic variants contribute to large gene expression changes across diverse human tissues and provide an integrative method for interpretation of rare variants in individual genomes.
- Xin Li
- , Yungil Kim
- & Stephen B. Montgomery
-
Letter |
 Zika virus evolution and spread in the Americas
Zika virus evolution and spread in the Americas
One hundred and ten Zika virus genomes from ten countries and territories involved in the Zika virus epidemic reveal rapid expansion of the epidemic within Brazil and multiple introductions to other regions.
- Hayden C. Metsky
- , Christian B. Matranga
- & Pardis C. Sabeti
-
Letter |
 Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism
Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism
Gene expression analysis in brain tissue from individuals with and without autism spectrum disorder (ASD) suggests that the transcription factor SOX5 contributes to an ASD-associated reduction in transcriptional differences between brain areas and indicates that common transcriptomic changes occur in different forms of ASD.
- Neelroop N. Parikshak
- , Vivek Swarup
- & Daniel H. Geschwind
-
Letter |
 Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
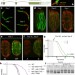
Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
Precursor mRNA splicing homeostasis is a biomarker and predictor of life expectancy in Caenorhabditis elegans and defects in global pre-mRNA splicing associated with age are reduced by dietary restriction via splicing factor 1.
- Caroline Heintz
- , Thomas K. Doktor
- & William B. Mair
-
Letter |
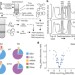 Dynamics of ribosome scanning and recycling revealed by translation complex profiling
Dynamics of ribosome scanning and recycling revealed by translation complex profiling
A translation complex sequencing approach has been developed enabling intermediates of all mRNA-associated processes of translation to be isolated and localized across the transcriptome; the results support longstanding models of initiation and termination and offer new mechanistic insights.
- Stuart K. Archer
- , Nikolay E. Shirokikh
- & Thomas Preiss
-
Letter |
 Tumour-specific proline vulnerability uncovered by differential ribosome codon reading
Tumour-specific proline vulnerability uncovered by differential ribosome codon reading
Tumours can require certain amino acids for their proliferation, and the diricore method described here helps to identify such restrictive amino acids; using this method in kidney cancer tissue and breast carcinoma cells, the authors observe an association between proline deficiency and upregulation of PYCR1, an enzyme required for proline synthesis.
- Fabricio Loayza-Puch
- , Koos Rooijers
- & Reuven Agami
-
Letter |
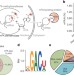 N6-methyladenosine-dependent regulation of messenger RNA stability
N6-methyladenosine-dependent regulation of messenger RNA stability
The mRNAs of higher eukaryotes are extensively modified internally with N6-methyladenosine, but the specific functional role of this modification has been unclear; here this modification on mRNA is shown to be recognized by several proteins, the modification and its recognition serve to regulate the RNA’s lifetime.
- Xiao Wang
- , Zhike Lu
- & Chuan He
-
Letter |
 Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells
Single-cell RNA sequencing is used to investigate the transcriptional response of 18 mouse bone-marrow-derived dendritic cells after lipopolysaccharide stimulation; many highly expressed genes, such as key immune genes and cytokines, show bimodal variation in both transcript abundance and splicing patterns. This variation reflects differences in both cell state and usage of an interferon-driven pathway involving Stat2 and Irf7.
- Alex K. Shalek
- , Rahul Satija
- & Aviv Regev
-
Letter |
 RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis
RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis
A new method allows the collection of circulating tumour cells (CTCs) despite their rarity; transcriptome sequencing of CTCs could allow identification of pathways involved in metastasis.
- Min Yu
- , David T. Ting
- & Daniel A. Haber
-
News |
 RNA studies under fire
RNA studies under fire
High-profile results challenged over statistical analysis of sequence data.
- Erika Check Hayden
-
Article |
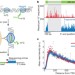 Nascent transcript sequencing visualizes transcription at nucleotide resolution
Nascent transcript sequencing visualizes transcription at nucleotide resolution
A novel technique called native elongating transcript sequencing (NET-seq) is described, which can quantify transcription with single nucleotide resolution. It is based on sequencing nascent transcripts associated with RNA polymerase II that are captured directly from live cells, and is used to gain insights into polymerase pausing and backtracking and the directionality of transcription.
- L. Stirling Churchman
- & Jonathan S. Weissman
-
Letter |
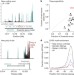 Understanding mechanisms underlying human gene expression variation with RNA sequencing
Understanding mechanisms underlying human gene expression variation with RNA sequencing
There is much interest in understanding the genetic mechanisms that underlie individual variations in gene expression. Here, RNA sequencing has been used to study gene expression in lymphoblastoid cell lines derived from Nigerian individuals for whom extensive genotype information is known. Numerous genetic determinants of variation in gene expression were identified, including variation in transcription, splicing and allele-specific expression.
- Joseph K. Pickrell
- , John C. Marioni
- & Jonathan K. Pritchard
-
Letter |
 Transcriptome genetics using second generation sequencing in a Caucasian population
Transcriptome genetics using second generation sequencing in a Caucasian population
Here, sequencing has been used to characterize the mRNA fraction of the transcriptome in Caucasian individuals, to provide a fine-scale view of transcriptomes and to identify genetic variants that affect alternative splicing. Measuring allele-specific expression identified rare expression quantitative trait loci (eQTLs) and allelic differences in transcript structure, revealing new properties of genetic effects on the transcriptome.
- Stephen B. Montgomery
- , Micha Sammeth
- & Emmanouil T. Dermitzakis
-
Article |
 The primary transcriptome of the major human pathogen Helicobacter pylori
The primary transcriptome of the major human pathogen Helicobacter pylori
The transcriptome of Helicobacter pylori, an important human pathogen involved in gastric ulcers and cancer, is presented. The approach establishes a model for mapping and annotating the primary transcriptomes of many living species.
- Cynthia M. Sharma
- , Steve Hoffmann
- & Jörg Vogel
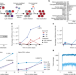
 Latent human herpesvirus 6 is reactivated in CAR T cells
Latent human herpesvirus 6 is reactivated in CAR T cells