Article
|
Open Access
Featured
-
-
Article
| Open Access Long non-coding RNA Linc-RAM enhances myogenic differentiation by interacting with MyoD
Long non-coding RNA Linc-RAM enhances myogenic differentiation by interacting with MyoD
Long non-coding RNAs are important regulators of many diverse biological processes. Here the authors describe Linc-RAM, which regulates myogenesis by binding MyoD and promoting the assembly of the MyoD–Baf60c–Brg1 complex at target genes.
- Xiaohua Yu
- , Yong Zhang
- & Dahai Zhu
-
Article
| Open Access Structural basis for the recognition and degradation of host TRIM proteins by Salmonella effector SopA
Structural basis for the recognition and degradation of host TRIM proteins by Salmonella effector SopA
The HECT-like E3 ligase SopA inSalmonellahas been suggested to activate host RING ligases TRIM56 and TRIM65. Here, the authors use mass spectrometry, crystal structures and biochemistry to examine the interactions between these proteins in detail.
- Evgenij Fiskin
- , Sagar Bhogaraju
- & Ivan Dikic
-
Article
| Open Access Structural basis of human PCNA sliding on DNA
Structural basis of human PCNA sliding on DNA
DNA sliding clamps are ring-shaped proteins that encircle DNA and harbour polymerases and other factors that promote processive DNA replication. Here the authors use X-ray crystallography, NMR and MD simulations to propose a model for a PCNA sliding mechanism that relies on short-lived polar interactions.
- Matteo De March
- , Nekane Merino
- & Alfredo De Biasio
-
Article
| Open Access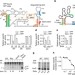 Transcriptional pausing at the translation start site operates as a critical checkpoint for riboswitch regulation
Transcriptional pausing at the translation start site operates as a critical checkpoint for riboswitch regulation
Riboswitches are non-coding RNA elements that detect metabolites and control expression by regulating mRNA levels or translation. Here, the authors provide evidence that theE. coli thiC riboswitch has a pause site in the translation initiation region that acts as a checkpoint for thiCexpression.
- Adrien Chauvier
- , Frédéric Picard-Jean
- & Daniel A. Lafontaine
-
Article
| Open Access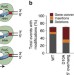 Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus
Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus
CRISPR-Cas9 has rapidly become a common molecular biology tool for modifying genomes and has been modified to generate single-strand nicks as well as double-strand breaks. Here the authors explore the DNA repair pathways activated by the different variants of Cas9.
- Anne Bothmer
- , Tanushree Phadke
- & Cecilia Cotta-Ramusino
-
Article
| Open Access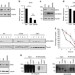 Praja1 E3 ubiquitin ligase promotes skeletal myogenesis through degradation of EZH2 upon p38α activation
Praja1 E3 ubiquitin ligase promotes skeletal myogenesis through degradation of EZH2 upon p38α activation
In skeletal muscle progenitors, EZH2 maintains myogenic genes in a repressed state, but during differentiation its levels are reduced via unknown mechanisms. Here the authors show that during myogenesis, p38α kinase phosphorylates EZH2 and targets it for degradation by the ubiquitin ligase PRAJA1.
- Silvia Consalvi
- , Arianna Brancaccio
- & Daniela Palacios
-
Article
| Open Access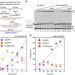 Oxidized nucleotide insertion by pol β confounds ligation during base excision repair
Oxidized nucleotide insertion by pol β confounds ligation during base excision repair
Oxidative stress in cells leads to the oxidations of DNA precursors. Here the authors show that these oxidized precursors can be incorporatedin vivoduring base excision repair, leading to DNA breaks and cytotoxicity.
- Melike Çağlayan
- , Julie K. Horton
- & Samuel H. Wilson
-
Article
| Open Access Structure of the cohesin loader Scc2
Structure of the cohesin loader Scc2
The cohesin complex maintains genome integrity by ensuring correct sister-chromatid segregation during mitosis and meiosis. Here, Chaoet al. present a pseudo-atomic model of the full-length Scc2–Scc4 cohesin loader complex and reveal key Scc2 surfaces crucial for cohesin loading.
- William C. H. Chao
- , Yasuto Murayama
- & Martin R. Singleton
-
Article
| Open Access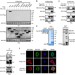 c-Src phosphorylation and activation of hexokinase promotes tumorigenesis and metastasis
c-Src phosphorylation and activation of hexokinase promotes tumorigenesis and metastasis
The protein tyrosine kinase c-Src is a renowned proto-oncogene with pleiotropic effects. Here, the authors show that c-Src induces the metabolic reprogramming of cancer cells by phosphorylating hexokinases HK1 and HK2, which in turns lead to increased HK catalytic activity and consequent enhancement of glycolysis.
- Jia Zhang
- , Suili Wang
- & Qinxi Li
-
Article
| Open Access PAXX promotes KU accumulation at DNA breaks and is essential for end-joining in XLF-deficient mice
PAXX promotes KU accumulation at DNA breaks and is essential for end-joining in XLF-deficient mice
Non-homologous end-joining is the key pathway for repairing double-stranded DNA breaks in mammalian cells. Here the authors show that PAXX promotes the accumulation of KU at DNA breaks and is essential for end-joining in cells lacking XLF.
- Xiangyu Liu
- , Zhengping Shao
- & Shan Zha
-
Article
| Open Access Cell division orientation is coupled to cell–cell adhesion by the E-cadherin/LGN complex
Cell division orientation is coupled to cell–cell adhesion by the E-cadherin/LGN complex
Cell–cell adhesion and oriented cell division play key roles in tissue architecture, but how they are coordinated is not known. Here, the authors show that E-cadherin interacts with LGN, and thereby provides a cortical cue that serves to stabilize cortical attachment of astral microtubules at cell–cell adhesions, thus orienting the mitotic spindle.
- Martijn Gloerich
- , Julie M. Bianchini
- & W. James Nelson
-
Article
| Open Access ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
Nonsense-mediated mRNA decay (NMD) is a quality control pathway that recognizes and degrades transcripts harbouring nonsense mutations. Here the authors show that the ATPase activity of UPF1 mediates functional interactions between the NMD machinery and ribosomes required for efficient ribosome release at premature termination codons.
- Lucas D. Serdar
- , DaJuan L. Whiteside
- & Kristian E. Baker
-
Article
| Open Access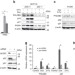 Pleckstrin homology domain-containing protein PHLDB3 supports cancer growth via a negative feedback loop involving p53
Pleckstrin homology domain-containing protein PHLDB3 supports cancer growth via a negative feedback loop involving p53
p53 is an oncosuppressor regulating several genes at the transcriptional level. Here, the authors identify a negative feedback loop between PHLDB3 and p53; PHLDB3 is a transcriptional target of p53 which facilitates MDM2-mediated p53 ubiquitination and degradation, impacting on tumorigenesis.
- Tengfei Chao
- , Xiang Zhou
- & Hua Lu
-
Article
| Open Access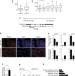 SNHG5 promotes colorectal cancer cell survival by counteracting STAU1-mediated mRNA destabilization
SNHG5 promotes colorectal cancer cell survival by counteracting STAU1-mediated mRNA destabilization
Several lncRNAs have been linked to cancer. Here, the authors identify SNHG5 as a long non-coding RNA promoting proliferation and survival of colorectal cancer cells by protecting specific mRNAs from STAU1-mediated degradation.
- Nkerorema Djodji Damas
- , Michela Marcatti
- & Anders H. Lund
-
Article
| Open Access A somatic piRNA pathway in the Drosophila fat body ensures metabolic homeostasis and normal lifespan
A somatic piRNA pathway in the Drosophila fat body ensures metabolic homeostasis and normal lifespan
The Piwi-interacting RNA (piRNA) pathway is known to suppress transposable elements in gonadal tissues. Here the authors provide evidence for a functional piRNA pathway in the somatic cells of theDrosophilafat body with roles in metabolism, immunological function and overall health.
- Brian C. Jones
- , Jason G. Wood
- & Stephen L. Helfand
-
Article
| Open Access Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
mRNA surveillance is essential to maintain homeostasis in eukaryotes and is activated by mRNAs lacking a stop codon. Here the authors describe a high resolution cryo-EM structure of a nonstop complex that shows how arrested ribosome recognition is achieved during Dom34-mediated mRNA surveillance.
- Tarek Hilal
- , Hiroshi Yamamoto
- & Christian M.T. Spahn
-
Article
| Open Access A bromodomain–DNA interaction facilitates acetylation-dependent bivalent nucleosome recognition by the BET protein BRDT
A bromodomain–DNA interaction facilitates acetylation-dependent bivalent nucleosome recognition by the BET protein BRDT
Many chromatin modifying proteins, including BRDT, contain bromodomains, which are known to interact with nucleosomes. Here, the authors find that BRDT interacts with nucleosomes via only one of its two bromodomains, and that the interaction involves contacts with DNA as well as acetylated histones.
- Thomas C. R. Miller
- , Bernd Simon
- & Christoph W. Müller
-
Article
| Open Access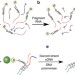 Global repositioning of transcription start sites in a plant-fermenting bacterium
Global repositioning of transcription start sites in a plant-fermenting bacterium
Bacteria may respond to a change in environment by using alternative transcriptional start sites. Here, the authors use a novel genome-wide capture and reverse transcription method to find substrate-specific start sites for hundreds of genes at single base resolution inClostridium phytofermentans.
- Magali Boutard
- , Laurence Ettwiller
- & Andrew C. Tolonen
-
Article
| Open Access Profiling DNA damage response following mitotic perturbations
Profiling DNA damage response following mitotic perturbations
DNA damage arising from replication stress is well studied, but the effect of mitotic errors on genome integrity is less understood. Here the authors knock down 47 mitotic regulators and record how they impact on DNA breakage events, providing a resource for future studies on the relation between cell division and genome integrity.
- Ronni S. Pedersen
- , Gopal Karemore
- & Claudia Lukas
-
Article
| Open Access TRBP ensures efficient Dicer processing of precursor microRNA in RNA-crowded environments
TRBP ensures efficient Dicer processing of precursor microRNA in RNA-crowded environments
The RNA binding protein TRBP is a component of the Dicer complex but its role in microRNA biogenesis remains poorly understood. Here the authors use a crowded RNA environment and single-molecule imaging to show that TRBP acts as a gatekeeper to prevent Dicer engagement with pre miRNA-like substrates.
- Mohamed Fareh
- , Kyu-Hyeon Yeom
- & Chirlmin Joo
-
Article
| Open Access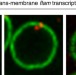 Export of piRNA precursors by EJC triggers assembly of cytoplasmic Yb-body in Drosophila
Export of piRNA precursors by EJC triggers assembly of cytoplasmic Yb-body in Drosophila
In Drosophila ovarian follicle cells, piRNAs generated from RNA precursors are processed in cytoplasmic Yb-bodies. The authors identify the exportin and the exon junction complexes as required to transfer precursors to cytoplasm. They also show that Yb-body formation requires piRNA precursor export.
- Cynthia Dennis
- , Emilie Brasset
- & Chantal Vaury
-
Article
| Open Access Chromatin-remodelling factor Brg1 regulates myocardial proliferation and regeneration in zebrafish
Chromatin-remodelling factor Brg1 regulates myocardial proliferation and regeneration in zebrafish
The adult zebrafish heart is capable of regeneration but the molecular mechanisms are poorly understood. Here the authors show that chromatin remodeling factor Brg1 represses cyclin-dependent kinase inhibitors to promote myocardial regeneration.
- Chenglu Xiao
- , Lu Gao
- & Jing-Wei Xiong
-
Article
| Open Access Transcriptional bursting is intrinsically caused by interplay between RNA polymerases on DNA
Transcriptional bursting is intrinsically caused by interplay between RNA polymerases on DNA
Transcriptional bursting is a potential source of cell-to-cell variability but the molecular mechanisms are unclear. Here the authors use single molecule imaging to analyse the kinetics of bursting on DNA and observe that bursting is an intrinsic property of RNA polymerases on DNA.
- Keisuke Fujita
- , Mitsuhiro Iwaki
- & Toshio Yanagida
-
Article
| Open Access WRN regulates pathway choice between classical and alternative non-homologous end joining
WRN regulates pathway choice between classical and alternative non-homologous end joining
Werner Syndrome is an accelerated aging disorder marked by genome instability, large deletions and telomere fusions, hallmarks of aberrant DNA repair. Here the authors report a role for the WRN helicase in regulating the choice between classical and alternative non-homologous end-joning.
- Raghavendra A. Shamanna
- , Huiming Lu
- & Vilhelm A. Bohr
-
Article
| Open Access Targeting the Notch-regulated non-coding RNA TUG1 for glioma treatment
Targeting the Notch-regulated non-coding RNA TUG1 for glioma treatment
Self-renewal of cancer stem cells can contribute to glioma progression. Here, the authors show that Notch1 activation in glioma stem cells induces expression of the lncRNATUG1, which promotes self-renewal through the repression of differentiation genes, and that targeting TUG1 represses glioma growth in vivo.
- Keisuke Katsushima
- , Atsushi Natsume
- & Yutaka Kondo
-
Article
| Open Access Interrogating the degradation pathways of unstable mRNAs with XRN1-resistant sequences
Interrogating the degradation pathways of unstable mRNAs with XRN1-resistant sequences
Degradation of messenger RNA is a key regulatory step in controlling eukaryotic gene expression. Here the authors present xrFrag, a molecular tool to interrogate the extent and directionality of mRNA turnover by the detection of stabilized decay intermediates produced by several common decay pathways.
- Volker Boehm
- , Jennifer V. Gerbracht
- & Niels H. Gehring
-
Article
| Open Access Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
The correct folding of proteins often requires the intervention molecular chaperones, which can occur co-translationally. Here the authors identify elements of yeast Ssb (Hsp70) that mediate ribosomal binding, and suggest a mechanism that directs efficient interaction of Ssb with the nascent chain.
- Marie A. Hanebuth
- , Roman Kityk
- & Elke Deuerling
-
Article
| Open Access LncBRM initiates YAP1 signalling activation to drive self-renewal of liver cancer stem cells
LncBRM initiates YAP1 signalling activation to drive self-renewal of liver cancer stem cells
Liver cancer stem cells (CSCs) may contribute to the high rate of recurrence of hepatocellular carcinoma. Here, the authors show that the long coding RNA, LcnBRM, regulates the self-renewal of liver CSCs and tumour initiation through binding to BAF complex thereby activating YAP1.
- Pingping Zhu
- , Yanying Wang
- & Zusen Fan
-
Article
| Open Access Structure of the RBM7–ZCCHC8 core of the NEXT complex reveals connections to splicing factors
Structure of the RBM7–ZCCHC8 core of the NEXT complex reveals connections to splicing factors
RBM7 and ZCCHC8 are two core subunits of the Nuclear Exosome Targeting complex, which regulates the degradation of selected non-coding RNAs in human cells. Here, the authors use structural and biochemical methods to show how ZCCHC8 recruits RBM7 in the complex, leaving the RNA binding site accessible and revealing possible implications for splicing.
- Sebastian Falk
- , Ksenia Finogenova
- & Elena Conti
-
Article
| Open Access Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2
Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2
The Polycomb repressive complexes PRC1 and PRC2 play a central role in developmental regulation of the genome in multicellular organisms. Here the authors describe how the PRC2 cofactor Jarid2 mediates the recruitment of the PRC2 complex to chromatin via interaction with H2AK119u1.
- Sarah Cooper
- , Anne Grijzenhout
- & Neil Brockdorff
-
Article
| Open Access Modulation of mRNA and lncRNA expression dynamics by the Set2–Rpd3S pathway
Modulation of mRNA and lncRNA expression dynamics by the Set2–Rpd3S pathway
H3K36 methylation by Set2 targets Rpd3S histone deacetylase to transcribed mRNA genes, repressing internal cryptic promoters and modulating elongation. Here, the authors provide evidence that the Set2-Rpd3S pathway also regulates dynamic expression of mRNAs and lncRNAs.
- Ji Hyun Kim
- , Bo Bae Lee
- & TaeSoo Kim
-
Article
| Open Access Selective suppression of antisense transcription by Set2-mediated H3K36 methylation
Selective suppression of antisense transcription by Set2-mediated H3K36 methylation
Maintenance of chromatin structure in coding regions is partially dependent on transcription, with histone methyltransferase Set2 playing a role in this process. Here, the authors provide evidence that Set2 regulates repression of a specific set of antisense RNAs embedded within the coding genes.
- Swaminathan Venkatesh
- , Hua Li
- & Jerry L. Workman
-
Article
| Open Access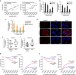 A feed-forward loop between lncARSR and YAP activity promotes expansion of renal tumour-initiating cells
A feed-forward loop between lncARSR and YAP activity promotes expansion of renal tumour-initiating cells
Renal tumour-initiating cells (T-ICs) contribute to tumour initiation and progression. Here, the authors show that lncARSR regulates TICs by blocking LATS1-induced YAP phosphorylation facilitating YAP nuclear translocation, which promotes lncARSR transcription, thus forming a feed-forward circuit to promote TIC expansion.
- Le Qu
- , Zhenjie Wu
- & Linhui Wang
-
Article
| Open Access Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
In yeast, the heterodimeric ribosome-associated complex (RAC) acts in concert with the Hsp70 protein Ssb, forming a unique chaperone triad. Here the authors use structural and biochemical approaches to shed light on how translation and folding are coupled in eukaryotes.
- Andrea Gumiero
- , Charlotte Conz
- & Irmgard Sinning
-
Article
| Open Access Multiple myeloma risk variant at 7p15.3 creates an IRF4-binding site and interferes with CDCA7L expression
Multiple myeloma risk variant at 7p15.3 creates an IRF4-binding site and interferes with CDCA7L expression
Genome wide association studies have identified multiple risk loci for multiple myeloma. Here, the authors show that the expression of CDCA7Lis associated with patient survival and expression of the gene is influenced by a risk variant at 7p15.3, which creates a transcription factor binding site for IRF4.
- Ni Li
- , David C. Johnson
- & Richard S. Houlston
-
Article
| Open Access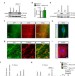 The Robo4 cytoplasmic domain is dispensable for vascular permeability and neovascularization
The Robo4 cytoplasmic domain is dispensable for vascular permeability and neovascularization
Robo4 is a transmembrane protein that regulates vascular permeability. Zhang et al. now reveal the mechanism of Robo4 action and show that Robo4 and UncB are required for VEGF-mediated regulation of vascular barrier by suppressing VEGF-induced phosphorylation of its receptor Vegfr2 on Y949.
- Feng Zhang
- , Claudia Prahst
- & Anne Eichmann
-
Article
| Open Access Repression of RNA polymerase by the archaeo-viral regulator ORF145/RIP
Repression of RNA polymerase by the archaeo-viral regulator ORF145/RIP
How archaeal viruses perturb host transcription machinery is poorly understood. Here, the authors provide evidence that the archaeo-viral transcription factor ORF145/RIP targets host RNA polymerase, repressing its activity.
- Carol Sheppard
- , Fabian Blombach
- & Finn Werner
-
Article
| Open Access Unlocking sperm chromatin at fertilization requires a dedicated egg thioredoxin in Drosophila
Unlocking sperm chromatin at fertilization requires a dedicated egg thioredoxin in Drosophila
In many animals, sperm DNA compaction is achieved by disulfide bonds between the sperm nuclear proteins that replace the histones. Here, the authors provide evidence that Drosophilamaternal thioredoxin Deadhead is required to unlock the sperm chromatin upon fertilization.
- Samantha Tirmarche
- , Shuhei Kimura
- & Benjamin Loppin
-
Article
| Open Access The GCN5-CITED2-PKA signalling module controls hepatic glucose metabolism through a cAMP-induced substrate switch
The GCN5-CITED2-PKA signalling module controls hepatic glucose metabolism through a cAMP-induced substrate switch
GCN5 inhibits hepatic gluconeogenesis through acetylation of PGC-1α. Here the authors show that GCN5 also activates hepatic gluconeogenesis by acetylating histone H3K9, and that the affinity of GCN5 for its different substrates is regulated via phosphorylation at S275 by PKA in a CITED2-dependent manner.
- Mashito Sakai
- , Tomoko Tujimura-Hayakawa
- & Michihiro Matsumoto
-
Article
| Open Access CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
Epithelial-to-mesenchymal transition is a key process in tumorigenesis but little is known about the molecular mechanism regulating such process at the translational level. Here, the authors identify a subset of mRNAs important for this process that are specifically modulated by the RNA-binding protein CELF1.
- Arindam Chaudhury
- , Shebna Cheema
- & Joel R. Neilson
-
Article
| Open Access BRCA1-regulated RRM2 expression protects glioblastoma cells from endogenous replication stress and promotes tumorigenicity
BRCA1-regulated RRM2 expression protects glioblastoma cells from endogenous replication stress and promotes tumorigenicity
BRCA1 loss can result in collapse of replication forks into DNA double strand breaks that can contribute to malignant transformation. Here, the authors find that BRCA1 promotes the expression of RRM2 protecting glioblastoma cells from replication stress, DNA damage and apoptosis.
- Rikke D. Rasmussen
- , Madhavsai K. Gajjar
- & Petra Hamerlik
-
Article
| Open Access Selective removal of deletion-bearing mitochondrial DNA in heteroplasmic Drosophila
Selective removal of deletion-bearing mitochondrial DNA in heteroplasmic Drosophila
Heteroplasmy, in which mutant and wild-type mitochondrial DNA (mtDNA) coexist in a cell, can result in diseases. Here the authors generate transgenic flies with heteroplasmic mtDNA in flight muscles, and show that stimulation of autophagy, or a decrease in mitofusin, promotes clearance of mutant mtDNA.
- Nikolay P. Kandul
- , Ting Zhang
- & Ming Guo
-
Article
| Open Access Epigenetic engineering reveals a balance between histone modifications and transcription in kinetochore maintenance
Epigenetic engineering reveals a balance between histone modifications and transcription in kinetochore maintenance
Centromeres are centrochromatin domains with CENP-A and H3 nucleosomes carrying transcription-associated modifications. Here the authors target synthetic modules to the centromeres to show that transcription plus histone modifications are required for CENP-A assembly and centrochromatin maintenance.
- Oscar Molina
- , Giulia Vargiu
- & William C. Earnshaw
-
Article
| Open Access Reader domain specificity and lysine demethylase-4 family function
Reader domain specificity and lysine demethylase-4 family function
KDM4 histone demethylases target specific chromatin regions by a mechanism that is not fully characterised. Here, the authors identify trimethyl-lysine histone-binding preferences for closely related KDM4 double tudor domains and use structural and biochemical information to examine the molecular details of this interaction.
- Zhangli Su
- , Fengbin Wang
- & John M. Denu
-
Article
| Open Access tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
Mycobacteria can adapt to the stress of human infection by entering a dormant state. Here the authors show that hypoxia-induced dormancy in M. bovisBCG involves the reprogramming of tRNA wobble modifications and copy numbers, coupled with biased use of synonymous codons in survival genes.
- Yok Hian Chionh
- , Megan McBee
- & Peter C. Dedon
-
Article
| Open Access DciA is an ancestral replicative helicase operator essential for bacterial replication initiation
DciA is an ancestral replicative helicase operator essential for bacterial replication initiation
DNA replication requires the loading of the replicative helicase onto the DNA molecule; in bacteria this was believed to be solely accomplished by DnaC and DnaI. Here the authors identify DciA as an ancestral and still widely distributed replicative helicase loader.
- Pierre Brézellec
- , Isabelle Vallet-Gely
- & Jean-Luc Ferat
-
Article
| Open Access Contracting CAG/CTG repeats using the CRISPR-Cas9 nickase
Contracting CAG/CTG repeats using the CRISPR-Cas9 nickase
The expansion of trinucleotide repeats has been linked to several neurodegenerative disorders. Here, the authors show that the CRISPR-Cas9 nuclease induces both expansions and contractions of the repeat region, whereas the nickase leads predominantly to contractions.
- Cinzia Cinesi
- , Lorène Aeschbach
- & Vincent Dion
-
Article
| Open Access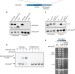 Chaperone addiction of toxin–antitoxin systems
Chaperone addiction of toxin–antitoxin systems
Some bacterial toxin-antitoxin systems consist of a labile antitoxin that inhibits a toxin, and a chaperone that stabilizes the antitoxin. Here, Bordes et al. identify a sequence within the antitoxin to which the chaperone binds and which can be transferred to other proteins to make them chaperone-dependent.
- Patricia Bordes
- , Ambre Julie Sala
- & Pierre Genevaux
-
Article
| Open Access AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
AKAP95 regulates splicing through scaffolding RNAs and RNA processing factors
The chromatin-associated protein AKAP95 is known for its chromatin-related functions including enhancing transcription. Here the authors show that AKAP95 interacts with the splicing regulatory factors as well as RNAs to regulate the inclusion of exons and pre-mRNA splicing.
- Jing Hu
- , Alireza Khodadadi-Jamayran
- & Hao Jiang
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Sox17 drives functional engraftment of endothelium converted from non-vascular cells
Sox17 drives functional engraftment of endothelium converted from non-vascular cells