Featured
-
-
Article
| Open Access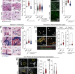 Host response during unresolved urinary tract infection alters female mammary tissue homeostasis through collagen deposition and TIMP1
Host response during unresolved urinary tract infection alters female mammary tissue homeostasis through collagen deposition and TIMP1
Urinary tract infections (UTIs) can elicit systemic host-responses. Here the authors report that, in a mouse model, unresolved UTI is associated with alterations of the mammary tissue, including collagen deposition and hyperplasia.
- Samantha Henry
- , Steven Macauley Lewis
- & Camila Oresco dos Santos
-
Article
| Open Access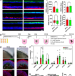 PRPF8-mediated dysregulation of hBrr2 helicase disrupts human spliceosome kinetics and 5´-splice-site selection causing tissue-specific defects
PRPF8-mediated dysregulation of hBrr2 helicase disrupts human spliceosome kinetics and 5´-splice-site selection causing tissue-specific defects
PRPF8 is a hotspot for mutations causing retinitis pigmentosa-type 13. Here the authors generated PRPF8 patient-specific retinal cells, demonstrating an important role for this splicing factor in spliceosome kinetics and 5’ splice site selection.
- Robert Atkinson
- , Maria Georgiou
- & Majlinda Lako
-
Article
| Open Access Single-cell and spatial RNA sequencing reveal the spatiotemporal trajectories of fruit senescence
Single-cell and spatial RNA sequencing reveal the spatiotemporal trajectories of fruit senescence
Fruit senescence is a complex physiological process. Here, the authors construct a single-cell expression atlas of pitaya pericarp pitaya to provide a spatiotemporal perspective of the dynamic process of plant senescence.
- Xin Li
- , Bairu Li
- & Robert Henry
-
Article
| Open Access Profiling the colonic mucosal response to fecal microbiota transplantation identifies a role for GBP5 in colitis in humans and mice
Profiling the colonic mucosal response to fecal microbiota transplantation identifies a role for GBP5 in colitis in humans and mice
Faecal microbiota transplantation (FMT) can be used to treat established colitis. Here the authors profile transcriptional changes in humans after FMT and how this relates to colitis remission identifying a role for GBP5, and this protein is validated in a loss-of-function mouse model.
- Laurence D. W. Luu
- , Abhimanu Pandey
- & Nadeem O. Kaakoush
-
Article
| Open Access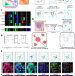 Mapping human tissues with highly multiplexed RNA in situ hybridization
Mapping human tissues with highly multiplexed RNA in situ hybridization
Application of multiplexed RNA in situ mapping techniques to human tissues remains challenging. Here, the authors report DART-FISH, a padlock probe-based technology capable of profiling large numbers of genes in centimetre-sized human tissue sections.
- Kian Kalhor
- , Chien-Ju Chen
- & Kun Zhang
-
Article
| Open Access A universal molecular control for DNA, mRNA and protein expression
A universal molecular control for DNA, mRNA and protein expression
Multi-omics analyses powerfully combine gene expression and translation, however no available controls can be used across these techniques. Here the authors develop pREF, a universal control construct designed for use in DNA, RNA and protein analyses.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Exemestane plus everolimus and palbociclib in metastatic breast cancer: clinical response and genomic/transcriptomic determinants of resistance in a phase I/II trial
Exemestane plus everolimus and palbociclib in metastatic breast cancer: clinical response and genomic/transcriptomic determinants of resistance in a phase I/II trial
Intrinsic and acquired resistances to CDK4/6 inhibitors have been described in patients with breast cancer. Here the authors report the results from a phase I/II clinical trial of the aromatase inhibitor exemestane plus everolimus (mTOR inhibitor) and palbociclib (CDK4/6i) in patients with metastatic breast cancer, assessing safety, clinical efficacy, as well as genomic and transcriptomic determinants of resistance.
- Jorge Gómez Tejeda Zañudo
- , Romualdo Barroso-Sousa
- & Nikhil Wagle
-
Article
| Open Access Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Leveraging the fact that eukaryotic genomes are organized into gene modules, FISHnCHIPs images multiple co-expressed genes simultaneously for sensitive and high throughput profiling of gene programs and cell types in tissues.
- Xinrui Zhou
- , Wan Yi Seow
- & Kok Hao Chen
-
Article
| Open Access Charting cellular differentiation trajectories with Ricci flow
Charting cellular differentiation trajectories with Ricci flow
When stem cells develop into tissues intracellular signalling is rewired, errors in this process lead to cancer. Here, authors applied tools from differential geometry made by Albert Einstein’s General Relativity to understand and predict biological network rewiring in health and disease.
- Anthony Baptista
- , Ben D. MacArthur
- & Christopher R. S. Banerji
-
Article
| Open Access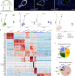 Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
This study uses single-cell transcriptomics to examine how lung cells respond to targeted damage. The authors employ genetically modified mouse models and cell sorting to enrich for rare, actively dividing cells, revealing cell types/states and alternative differentiation paths.
- Leila R. Martins
- , Lina Sieverling
- & Claudia Scholl
-
Article
| Open Access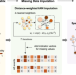 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access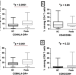 Evidence for immune activation in pathogenesis of the HLA class II associated disease, podoconiosis
Evidence for immune activation in pathogenesis of the HLA class II associated disease, podoconiosis
Podoconiosis is triggered by long term barefoot exposure to volcanic red clay soil. Here, Negash et al characterise the immune profile of podoconiosis patients to show this disease is associated with high levels of immune activation and inflammation.
- Mikias Negash
- , Menberework Chanyalew
- & Melanie J. Newport
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Article
| Open Access High resolution spatial profiling of kidney injury and repair using RNA hybridization-based in situ sequencing
High resolution spatial profiling of kidney injury and repair using RNA hybridization-based in situ sequencing
Advancements in spatial transcriptomics technologies have enabled the analysis of gene expression at cellular resolution in situ. The authors applied direct RNA hybridization-based in situ sequencing (dRNA HybISS) and developed a computational tool, CellScopes, to study gene expression in mouse kidneys, identifying cellular changes and interactions during injury and repair.
- Haojia Wu
- , Eryn E. Dixon
- & Benjamin D. Humphreys
-
Article
| Open Access Tracing back primed resistance in cancer via sister cells
Tracing back primed resistance in cancer via sister cells
Transcriptional cell states can drive treatment resistance in cancer. Here, the authors develop ReSisTrace to predict cell states that are primed to resist ovarian cancer treatment and validate their findings using small molecule inhibitors.
- Jun Dai
- , Shuyu Zheng
- & Anna Vähärautio
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access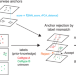 Semi-supervised integration of single-cell transcriptomics data
Semi-supervised integration of single-cell transcriptomics data
Batch effects hinder multi-sample single-cell data analyses. Here, authors present STACAS, a scalable single-cell RNA-seq data integration tool that uses prior cell type knowledge to preserve biological variability, demonstrating robustness to noisy input cell type labels.
- Massimo Andreatta
- , Léonard Hérault
- & Santiago J. Carmona
-
Article
| Open Access Trajectory inference across multiple conditions with condiments
Trajectory inference across multiple conditions with condiments
scRNA-Seq has enabled the study of dynamic systems such as response to a drug at the individual cell and gene levels. Here the authors introduce a framework to interpret differences at the trajectory, cell populations, and individual gene levels.
- Hector Roux de Bézieux
- , Koen Van den Berge
- & Sandrine Dudoit
-
Article
| Open Access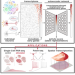 Using deep learning to quantify neuronal activation from single-cell and spatial transcriptomic data
Using deep learning to quantify neuronal activation from single-cell and spatial transcriptomic data
Neuronal activity is associated with transcriptional changes. Here, the authors present a deep learning model that integrates single cell transcriptomic signals to estimate neuronal activation.
- Ethan Bahl
- , Snehajyoti Chatterjee
- & Jacob J. Michaelson
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access A full-body transcription factor expression atlas with completely resolved cell identities in C. elegans
A full-body transcription factor expression atlas with completely resolved cell identities in C. elegans
Invariant cell lineage in C. elegans enables the analysis of transcriptional regulatory mechanisms controlling the fate of each cell at spatiotemporal resolution. Here, the authors develop a tool automating C. elegans cell identification and create an expression atlas of 620 transcription factors.
- Yongbin Li
- , Siyu Chen
- & Xiao Liu
-
Article
| Open Access Silica-associated proteins from hexactinellid sponges support an alternative evolutionary scenario for biomineralization in Porifera
Silica-associated proteins from hexactinellid sponges support an alternative evolutionary scenario for biomineralization in Porifera
Sponges, being early-diverging metazoans and the only animals to develop extensive skeletons of silica, have potential to inform about the evolutionary steps of metazoan traits, including biomineralization. Here, the authors characterize two proteins associated with the hexactinellid sponge silica.
- Katsuhiko Shimizu
- , Michika Nishi
- & Manuel Maldonado
-
Article
| Open Access The single-cell transcriptomic atlas and RORA-mediated 3D epigenomic remodeling in driving corneal epithelial differentiation
The single-cell transcriptomic atlas and RORA-mediated 3D epigenomic remodeling in driving corneal epithelial differentiation
Ocular homeostasis and vision depend on the differentiation of limbal stem/progenitor cells into corneal epithelial cells. Here, the authors report transcriptional dynamics and RORA-mediated epigenetic remodeling underlying the differentiation of human corneal epithelium.
- Mingsen Li
- , Huizhen Guo
- & Hong Ouyang
-
Article
| Open Access Systems-based identification of the Hippo pathway for promoting fibrotic mesenchymal differentiation in systemic sclerosis
Systems-based identification of the Hippo pathway for promoting fibrotic mesenchymal differentiation in systemic sclerosis
Systemic sclerosis (SSc) is an autoimmune disease causing skin fibrosis and organ inflammation. Here the authors generate and analyze SSc skin single cell RNA sequencing data to propose contributions from both myofibroblasts and endothelial-to-mesenchymal -transitioning cells (EndoMT) to skin fibrosis, and to implicate the involvement of Hippo signaling pathways.
- Feiyang Ma
- , Pei-Suen Tsou
- & Johann E. Gudjonsson
-
Article
| Open Access Unraveling the causal genes and transcriptomic determinants of human telomere length
Unraveling the causal genes and transcriptomic determinants of human telomere length
Variation in human telomere length has been well studied, but most previous studies have used adult telomere length. Here, the authors explore the genetic basis of telomere length in the placenta and find suggestive causal genes modulating human telomere length.
- Ying Chang
- , Yao Zhou
- & Dandan Huang
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access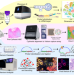 High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis
High resolution mapping of the tumor microenvironment using integrated single-cell, spatial and in situ analysis
The integration of single-cell and spatial data can provide a more comprehensive picture of the network of cells within the tumour microenvironment. Here the authors use a combination of single-cell and spatial technologies including 10x Xenium to characterise serial formalin-fixed, paraffin-embedded human breast cancer sections.
- Amanda Janesick
- , Robert Shelansky
- & Sarah E. B. Taylor
-
Article
| Open Access Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Whole-genome sequencing reveals the molecular implications of the stepwise progression of lung adenocarcinoma
Current sequencing technologies can shed light on the stepwise progression of lung adenocarcinoma. Here, the authors characterize tumor progression in lung adenocarcinomas from an early stage using short and long read whole-genome sequencing, bulk and spatial transcriptomics, and epigenomics.
- Yasuhiko Haga
- , Yoshitaka Sakamoto
- & Ayako Suzuki
-
Article
| Open Access Dissecting the basis for differential substrate specificity of ADAR1 and ADAR2
Dissecting the basis for differential substrate specificity of ADAR1 and ADAR2
Human ADAR1 and ADAR2 edit millions of adenosines transcriptome-wide, altering RNA structure. Here the authors show that variations in RNA binding domains influence site-specific editing, enhancing ADAR2-targeted therapeutics.
- Marlon S. Zambrano-Mila
- , Monika Witzenberger
- & Schraga Schwartz
-
Article
| Open Access Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Intra-cell line heterogeneity remains to be characterized. Here, the use of single multi-omics on a large panel of human cell lines identifies copy number variation, epigenetic variation and extrachromosomal DNA distribution as the main contributors to intra-cell line heterogeneity.
- Qionghua Zhu
- , Xin Zhao
- & Liang Wu
-
Article
| Open Access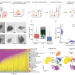 Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Experimentally preserving tissue microenvironments remains challenging for single-cell sequencing methods. Here, the authors introduce fragment-sequencing, a method that can preserve three-dimensional microenvironments in single-cell RNA-seq, thus allowing the reconstruction of spatial tissue niches.
- Kristina Handler
- , Karsten Bach
- & Andreas E. Moor
-
Article
| Open Access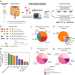 Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Long-read single-cell RNA sequencing is capable of detecting isoform-level gene expression and genomic alterations such as mutations and gene fusions, thereby providing cell-specific genotype-phenotype information. Here, the authors use long-read scRNA-seq on metastatic ovarian cancer samples and detect cell-type specific isoforms and gene fusions that may otherwise be misclassified in short-read data.
- Arthur Dondi
- , Ulrike Lischetti
- & Niko Beerenwinkel
-
Article
| Open Access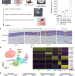 Vector-borne Trypanosoma brucei parasites develop in artificial human skin and persist as skin tissue forms
Vector-borne Trypanosoma brucei parasites develop in artificial human skin and persist as skin tissue forms
Here, authors show that tsetse-fly transmitted trypanosomes rapidly differentiate within human skin equivalents, eventually entering a reversible quiescent stage and conclude that these skin tissue forms may contribute to long-term parasite infections in asymptomatic individuals.
- Christian Reuter
- , Laura Hauf
- & Markus Engstler
-
Article
| Open Access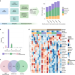 Distinct transcriptomic profiles in children prior to the appearance of type 1 diabetes-linked islet autoantibodies and following enterovirus infection
Distinct transcriptomic profiles in children prior to the appearance of type 1 diabetes-linked islet autoantibodies and following enterovirus infection
Although type-1 diabetes has a clear genetic component, not all children who are at risk eventually develop autoimmunity, suggesting the existence of environmental triggers. In this longitudinal transcriptomic study, the authors find that children who later develop autoimmunity have a distinct profile before the appearance of autoantibodies and may have impaired responses to enterovirus infection.
- Jake Lin
- , Elaheh Moradi
- & Matti Nykter
-
Article
| Open Access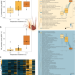 Experimental mining plumes and ocean warming trigger stress in a deep pelagic jellyfish
Experimental mining plumes and ocean warming trigger stress in a deep pelagic jellyfish
The deep ocean is increasingly subjected to human-induced environmental change, but little is known about species-specific responses to stressors, including those from deep sea mining. This study shows that elevated temperatures and simulated sediment plumes cause physiological stress in a cosmopolitan deep-sea jellyfish, confirming the detrimental impact of seabed mining.
- Vanessa I. Stenvers
- , Helena Hauss
- & Henk-Jan T. Hoving
-
Article
| Open Access The transcriptional and phenotypic characteristics that define alveolar macrophage subsets in acute hypoxemic respiratory failure
The transcriptional and phenotypic characteristics that define alveolar macrophage subsets in acute hypoxemic respiratory failure
Acute hypoxemic respiratory failure (AHRF) and the associated lung immune cell features are not well understood. Here the authors use CITE-Seq to analyse the transcriptomic and phenotypic profile of lung and blood cells from a longitudinal cohort of patients with AHRF to identify gene signatures and cell surface proteins associated with disease severity.
- Eric D. Morrell
- , Sarah E. Holton
- & Carmen Mikacenic
-
Article
| Open Access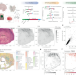 High-sensitive spatially resolved T cell receptor sequencing with SPTCR-seq
High-sensitive spatially resolved T cell receptor sequencing with SPTCR-seq
Understanding T cell behaviour in cancers is vital for improving immunotherapies. Here, the authors present spatially resolved T cell receptor sequencing (SPTCR-seq), a technology that annotates T cell receptors within the tumour ecosystem.
- Jasim Kada Benotmane
- , Jan Kueckelhaus
- & Dieter Henrik Heiland
-
Article
| Open Access A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Pseudotime analysis is prevalent in single-cell RNA-seq, but it remains challenging to perform it across multiple samples and experimental conditions. Here, the authors develop Lamian, a computational framework for multi-sample pseudotime analysis that adjusts for biological and technical variation to detect gene program changes along cell trajectories and across conditions.
- Wenpin Hou
- , Zhicheng Ji
- & Hongkai Ji
-
Article
| Open Access EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
The study reveals limitations in widely used RNA-seq aligners, which create 'phantom' introns in reference databases. The authors introduce EASTR, a computational tool that not only enhances alignment accuracy but also uncovers existing annotation errors. This improvement bolsters the dependability of subsequent RNA-seq analyses.
- Ida Shinder
- , Richard Hu
- & Mihaela Pertea
-
Article
| Open Access Spatial transcriptomics uncover sucrose post-phloem transport during maize kernel development
Spatial transcriptomics uncover sucrose post-phloem transport during maize kernel development
Maize kernels have long intrigued researchers due to their complex structure. Through microscopic sectioning and spatial transcriptomics, the authors observed the spatial distribution of RNA through electronic RNA in situ hybridization maps and discovered how storage accumulation occurs.
- Yuxin Fu
- , Wenxin Xiao
- & Wenqin Wang
-
Article
| Open Access Dissecting the human leptomeninges at single-cell resolution
Dissecting the human leptomeninges at single-cell resolution
The meninges protect the central nervous system at the brain border, and its dysfunction can lead to neural inflammation and cell damage. Here, the authors uncover the gene signatures of diverse cell types in the aged human leptomeninges and highlight their changes in Alzheimer’s Disease.
- Nicola A. Kearns
- , Artemis Iatrou
- & Yanling Wang
-
Article
| Open Access Isoform-resolved transcriptome of the human preimplantation embryo
Isoform-resolved transcriptome of the human preimplantation embryo
Human embryo development involves extensive transcriptional remodeling. In this study, the authors apply long- and short-read RNA-Seq to profile the transcriptomes of 73 human preimplantation embryos spanning zygotic to blastocyst stages, identifying tens of thousands of additional isoforms transcribed from both known and unannotated gene loci.
- Denis Torre
- , Nancy J. Francoeur
- & Robert Sebra
-
Article
| Open Access eQTL mapping in fetal-like pancreatic progenitor cells reveals early developmental insights into diabetes risk
eQTL mapping in fetal-like pancreatic progenitor cells reveals early developmental insights into diabetes risk
Fetal development plays an important role in defining adult diabetes risk. Here, authors identified a genetic link between fetal pancreatic gene expression, obesity, and diabetes risk through eQTL mapping of iPSC-derived pancreatic progenitor cells.
- Jennifer P. Nguyen
- , Timothy D. Arthur
- & Kelly A. Frazer
-
Article
| Open Access Profiling of the Helicobacter pylori redox switch HP1021 regulon using a multi-omics approach
Profiling of the Helicobacter pylori redox switch HP1021 regulon using a multi-omics approach
Helicobacter pylori adapted to the harsh conditions of the human stomach using a handful of regulatory proteins. Here, the authors identify H. pylori processes controlled by the HP1021 response regulator under optimal growth and oxidative stress.
- Mateusz Noszka
- , Agnieszka Strzałka
- & Anna Zawilak-Pawlik
-
Article
| Open Access DNMT and HDAC inhibition induces immunogenic neoantigens from human endogenous retroviral element-derived transcripts
DNMT and HDAC inhibition induces immunogenic neoantigens from human endogenous retroviral element-derived transcripts
Epigenetic therapies are known to synergize with immunotherapies through the de-repression of endogenous retroviral element (ERV)-encoded promoters. Here the authors identify treatment-induced neoantigens and validate their ability to induce T cell response and anti-tumor effects in vitro and in patient samples.
- Ashish Goyal
- , Jens Bauer
- & Christoph Plass
-
Article
| Open Access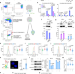 Phagocytosis-initiated tumor hybrid cells acquire a c-Myc-mediated quasi-polarization state for immunoevasion and distant dissemination
Phagocytosis-initiated tumor hybrid cells acquire a c-Myc-mediated quasi-polarization state for immunoevasion and distant dissemination
The CD47/SIRPα axis is well known to mediate immune escape by promoting cancer resistance to phagocytosis. Here the authors show that low CD47-expressing prostate cancer cells still allow phagocytosis but the process is incomplete leading to the formation of macrophage:tumor hybrid cells with immune evasive and pro-metastatic properties.
- Chih-Wei Chou
- , Chia-Nung Hung
- & Tim Hui-Ming Huang
-
Article
| Open Access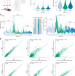 Miniature spatial transcriptomics for studying parasite-endosymbiont relationships at the micro scale
Miniature spatial transcriptomics for studying parasite-endosymbiont relationships at the micro scale
The filarial worm Brugia malayi has evolved a mutualistic association with the endosymbiotic bacteria Wolbachia. Here, Sounart et al describe a spatial transcriptomic technique that can spatially resolve these miniature specimens.
- Hailey Sounart
- , Denis Voronin
- & Stefania Giacomello
-
Article
| Open Access Species-specific metabolic reprogramming in human and mouse microglia during inflammatory pathway induction
Species-specific metabolic reprogramming in human and mouse microglia during inflammatory pathway induction
The innate immune cells undergo metabolic reprogramming upon inflammation. Here, the authors report that both mouse and human microglia display a metabolic reprogramming in the presence of a TLR4 activation, however species-specific enzymes are responsible for this process.
- Angélica María Sabogal-Guáqueta
- , Alejandro Marmolejo-Garza
- & Amalia Dolga
-
Article
| Open Access Unaltered hepatic wound healing response in male rats with ancestral liver injury
Unaltered hepatic wound healing response in male rats with ancestral liver injury
How much the environment influences inherited adaptive traits is debated and challenging to demonstrate in mammals. Here the authors performed a multigeneration study that failed to morphologically replicate enhanced wound healing response following ancestral liver injury in rats. However, heritable transcriptional effects suggest transmission at the molecular level, albeit of unclear functional relevance.
- Johanna Beil
- , Juliane Perner
- & Rémi Terranova

 hoxc12/c13 as key regulators for rebooting the developmental program in Xenopus limb regeneration
hoxc12/c13 as key regulators for rebooting the developmental program in Xenopus limb regeneration