Featured
-
-
Article
| Open Access Regulation by the RNA-binding protein Unkempt at its effector interface
Regulation by the RNA-binding protein Unkempt at its effector interface
How RNA-binding proteins (RBPs) regulate gene expression via effectors of RNA processing is unclear. Here, the authors dissect the effector interface of an essential RBP, Unkempt, and investigate its contribution to translational control in cells.
- Kriti Shah
- , Shiyang He
- & Jernej Murn
-
Article
| Open Access Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
This study analyzes the embryonic replacement of maternally contributed mRNA with new mRNA in single cells and shows dynamic spatio-temporal regulation of maternal mRNA decay and cell-type specific retention within the earliest specified cell types in zebrafish embryos.
- Lior Fishman
- , Avani Modak
- & Michal Rabani
-
Article
| Open Access Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing
Loss-of-function mutation in PRMT9 causes abnormal synapse development by dysregulation of RNA alternative splicing
Mutations in protein arginine methyltransferase 9 (PRMT9) are linked to intellectual disability. Here, the authors show that mutant PRMT9 fails to methylate its primary substrate SF3B2, causing aberrant RNA splicing and abnormal synapse development.
- Lei Shen
- , Xiaokuang Ma
- & Yanzhong Yang
-
Article
| Open Access A rapid inducible RNA decay system reveals fast mRNA decay in P-bodies
A rapid inducible RNA decay system reveals fast mRNA decay in P-bodies
Studying RNA decay remains a challenging task. Here, the authors present a technology that enables inducible rapid degradation of targeted mRNAs. Visualizing mRNA decay dynamics unveils insights into P-body function in RNA metabolism.
- Lauren A. Blake
- , Leslie Watkins
- & Bin Wu
-
Article
| Open Access Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Here the authors uncovered CNOT3, a subunit of the CCR4-NOT complex, as an essential modulator of translation in leukemia. The work pointed to the potential of targeting the posttranscriptional circuitry via CNOT3 as a therapeutic vulnerability in acute myeloid leukemia.
- Maryam Ghashghaei
- , Yilin Liu
- & Ly P. Vu
-
Article
| Open Access Toll/interleukin-1 receptor (TIR) domain-containing proteins have NAD-RNA decapping activity
Toll/interleukin-1 receptor (TIR) domain-containing proteins have NAD-RNA decapping activity
Toll/interleukin-1 receptor domain-containing proteins can catabolize NAD+. Here, Wang et al show that these proteins can also function as NAD-RNA decapping enzymes by releasing the NAM moiety from the NAD-RNA, resulting in the regulation of gene expression.
- Xufeng Wang
- , Dongli Yu
- & Xuemei Chen
-
Article
| Open Access Specificity, synergy, and mechanisms of splice-modifying drugs
Specificity, synergy, and mechanisms of splice-modifying drugs
Two small-molecule drugs, risdiplam and branaplam, have been developed for treating spinal muscular atrophy. Here the authors develop quantitative modeling methods for the sequence-specific and concentration-dependent effects of these and other splice-modifying drugs.
- Yuma Ishigami
- , Mandy S. Wong
- & Justin B. Kinney
-
Article
| Open Access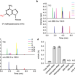 C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
Duan et al. demonstrate that the m2A modification is ubiquitous in plants and tRNA m2A37 promotes a relaxed conformation of tRNA, enhancing translation efficiency by facilitating decoding of tandem m2A-tRNA-dependent codons.
- Hong-Chao Duan
- , Chi Zhang
- & Guifang Jia
-
Article
| Open Access Multiplexed screening reveals how cancer-specific alternative polyadenylation shapes tumor growth in vivo
Multiplexed screening reveals how cancer-specific alternative polyadenylation shapes tumor growth in vivo
Dysregulation of alternative polyadenylation (APA) is associated with poor prognosis in cancer but its functional role is less clear. Here, the authors develop a CRISPR-Cas9- based screen to determine the effects of different APA events on melanoma growth in mouse models.
- Austin M. Gabel
- , Andrea E. Belleville
- & Robert K. Bradley
-
Article
| Open Access Expanded palette of RNA base editors for comprehensive RBP-RNA interactome studies
Expanded palette of RNA base editors for comprehensive RBP-RNA interactome studies
RNA base-editors are often used in methods for RNA binding protein (RBP) target discovery. Here the authors present a new RBP target discovery method, PRINTER, and suggest optimal RNA base-editors for dual-RBP studies, emphasizing the importance of matching rBEs’ editing biases with RBPs’ binding preferences.
- Hugo C. Medina-Munoz
- , Eric Kofman
- & Gene W. Yeo
-
Article
| Open Access 2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
Here the authors captured the structure of human telomerase H/ACA ribonucleoprotein (RNP) by cryo-EM. The structure rationalizes telomere-disorder disease mutations and reveals insights into the mechanism of pseudouridylation by eukaryotic H/ACA RNPs.
- George E. Ghanim
- , Zala Sekne
- & Thi Hoang Duong Nguyen
-
Article
| Open Access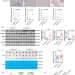 LOX-1 acts as an N6-methyladenosine-regulated receptor for Helicobacter pylori by binding to the bacterial catalase
LOX-1 acts as an N6-methyladenosine-regulated receptor for Helicobacter pylori by binding to the bacterial catalase
N6-methyladenosine (m6A) modification of mRNA regulates gene expression in eukaryotes. Here, Zeng et al. show that m6A modification of mRNAs contributes to protection against the pathogen Helicobacter pylori by downregulating a host protein that acts as receptor for the pathogen.
- Judeng Zeng
- , Chuan Xie
- & William K. K. Wu
-
Article
| Open Access Structural differences between the closely related RNA helicases, UAP56 and URH49, fashion distinct functional apo-complexes
Structural differences between the closely related RNA helicases, UAP56 and URH49, fashion distinct functional apo-complexes
UAP56 is an important factor in the TREX complex, which is responsible for mRNA export. Here the authors show that the closely related RNA helicases, UAP56 and URH49, exhibit different three-dimensional structures due to one amino acid change. Accordingly, they form distinct apo-complexes and function in the nuclear export of specific target mRNAs.
- Ken-ichi Fujita
- , Misa Ito
- & Seiji Masuda
-
Article
| Open Access PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
R-loop accumulation at transcription sites poses a persistent threat to genome integrity. Here the authors demonstrate a role for PSIP1/LEDGF protein in reducing R-loop levels at the site of transcription and preventing transcription replication conflict to maintain genome integrity.
- Sundarraj Jayakumar
- , Manthan Patel
- & Madapura M. Pradeepa
-
Article
| Open Access hnRNP A1 dysfunction alters RNA splicing and drives neurodegeneration in multiple sclerosis (MS)
hnRNP A1 dysfunction alters RNA splicing and drives neurodegeneration in multiple sclerosis (MS)
HnRNP A1 dysfunction is associated with neurodegeneration in multiple sclerosis (MS). Herein, advanced RNA sequencing and CLIPseq of MS brains and relevant models demonstrated that hnRNP A1 binding of target RNAs and RNA splicing were altered, precipitating neurodegeneration.
- Hannah E. Salapa
- , Patricia A. Thibault
- & Michael C. Levin
-
Article
| Open Access Role of UPF1-LIN28A interaction during early differentiation of pluripotent stem cells
Role of UPF1-LIN28A interaction during early differentiation of pluripotent stem cells
UPF1 and LIN28A are RNA-binding proteins involved in post-transcriptional regulation and cell differentiation. Here, authors report that they interact with each other via specific domains and regulate ectodermal specialization of human pluripotent stem cells.
- Seungwon Jung
- , Seung Hwan Ko
- & Jungwook Hwang
-
Article
| Open Access RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
The role of alternative splicing in pancreatic cancer (PDAC) development remains to be explored. Here, RBFOX2 is shown to regulate exon splicing events in transcripts encoding proteins involved in cytoskeletal remodelling programs and its downregulation promotes PDAC progression and liver metastasis.
- Michelle Maurin
- , Mohammadreza Ranjouri
- & Karen M. Mann
-
Article
| Open Access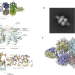 Acetylation regulates the oligomerization state and activity of RNase J, the Helicobacter pylori major ribonuclease
Acetylation regulates the oligomerization state and activity of RNase J, the Helicobacter pylori major ribonuclease
Here the authors find that RNase J, the major ribonuclease of the gastric pathogen Helicobacter pylori is post-translationally modified by acetylation. They show that acetylation can control RNase J activity.
- Alejandro Tejada-Arranz
- , Aleksei Lulla
- & Hilde De Reuse
-
Article
| Open Access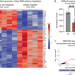 Modulation of insulin secretion by RBFOX2-mediated alternative splicing
Modulation of insulin secretion by RBFOX2-mediated alternative splicing
Insulin secretion from the pancreatic beta cell is a tightly regulated process that is vital for maintaining blood glucose homeostasis. Here, the authors show that the RNA binding protein RBFOX2 is a regulator of insulin secretion through the alternative splicing of genes required for insulin granule docking and exocytosis.
- Nicole D. Moss
- , Kristen L. Wells
- & Lori Sussel
-
Article
| Open Access Identification of NAD-RNA species and ADPR-RNA decapping in Archaea
Identification of NAD-RNA species and ADPR-RNA decapping in Archaea
NAD serves as a 5′-terminal cap for bacterial and eukaryotic transcripts, and can be degraded at high temperatures to generate ADP-ribose (ADPR). Here, Gomes-Filho et al. identify NAD-RNAs in thermophilic and mesophilic archaea and provide insights into NAD- and ADPR-mediated turnover of RNAs in these organisms.
- José Vicente Gomes-Filho
- , Ruth Breuer
- & Lennart Randau
-
Article
| Open Access MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Methylation is the dominant modification in mRNA and occurs at a variety of sites. Here, Hartstock et al. show that a clickable analogue of the key cosubstrate S-adenosyl-L-methionine (SAM) can be produced in cells, allowing for identification and mapping of different methylated nucleosides in mRNA.
- Katja Hartstock
- , Nadine A. Kueck
- & Andrea Rentmeister
-
Article
| Open Access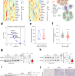 Spliceosome component Usp39 contributes to hepatic lipid homeostasis through the regulation of autophagy
Spliceosome component Usp39 contributes to hepatic lipid homeostasis through the regulation of autophagy
Non-alcoholic fatty liver disease affects 25% of people worldwide. Here the authors report that spliceosome component Usp39 deletion in mice leads to spontaneous steatosis and impaired autophagy through the regulation of alternative splicing.
- Donghai Cui
- , Zixiang Wang
- & Zhaojian Liu
-
Article
| Open Access Phase separation of BuGZ regulates gut regeneration and aging through interaction with m6A regulators
Phase separation of BuGZ regulates gut regeneration and aging through interaction with m6A regulators
Phase separation serves to compartmentalize and concentrate cellular components to facilitate essential physiological processes. Here, the authors elucidate the role and mechanism of BuGZ-mediated phase separation in the context of gut regeneration and aging.
- Qiaoqiao Zhang
- , Kai Deng
- & Hao Jiang
-
Article
| Open Access The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
Sm protein binding to pre-snRNA is a key step in snRNP biogenesis catalyzed by the SMN complex. Here, the authors show that pre-snRNAs adopt compact structures incompatible with Sm protein binding and that Gemin3 and 4 are required for pre-snRNA rearrangement to allow Sm protein interaction.
- Josef Pánek
- , Adriana Roithová
- & David Staněk
-
Article
| Open Access ALKBH5 controls the meiosis-coupled mRNA clearance in oocytes by removing the N 6-methyladenosine methylation
ALKBH5 controls the meiosis-coupled mRNA clearance in oocytes by removing the N 6-methyladenosine methylation
N6-methyladenosine (m6A) maintains maternal RNA stability in oocytes. Here, the authors identify demethylase ALKBH5 as a key determinant of oocyte quality and unveil the facilitating role of ALKBH5-mediated m6A removal in maternal RNA decay.
- Long Bai
- , Yu Xiang
- & Yimin Zhu
-
Article
| Open Access YTHDF2 facilitates aggresome formation via UPF1 in an m6A-independent manner
YTHDF2 facilitates aggresome formation via UPF1 in an m6A-independent manner
YTHDF2 has been extensively studied as an m6A-related RNA metabolism. Here, the authors show that YTHDF2 also contributes to protein homeostasis in an m6A-independent manner by promoting the formation of aggresomes through its interaction with UPF1.
- Hyun Jung Hwang
- , Tae Lim Park
- & Yoon Ki Kim
-
Article
| Open Access Recognition and cleavage mechanism of intron-containing pre-tRNA by human TSEN endonuclease complex
Recognition and cleavage mechanism of intron-containing pre-tRNA by human TSEN endonuclease complex
tRNA splicing is universal. Here the authors report the structure of an active human tRNA splicing endonuclease complex bound to an intron-containing pre-tRNA, which unveils eukaryotic tRNA processing and links archaeal and eukaryotic TSEN evolution.
- Ling Yuan
- , Yaoyao Han
- & Yadong Sun
-
Article
| Open Access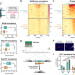 A R-loop sensing pathway mediates the relocation of transcribed genes to nuclear pore complexes
A R-loop sensing pathway mediates the relocation of transcribed genes to nuclear pore complexes
Here the authors report that DNA:RNA hybrid-containing R-loop structures are sensed by the ssDNA-binding protein RPA, triggering their relocation to nuclear pore complexes and attenuating transcription-associated genetic instability.
- Arianna Penzo
- , Marion Dubarry
- & Benoit Palancade
-
Article
| Open Access Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Genome-wide probing of eukaryotic nascent RNA structure elucidates cotranscriptional folding and its antimutagenic effect
Here, the authors present eSPET-seq a method to measure cotranscriptional RNA folding in eukaryotes. Further analysis reveals an antimutagenic effect of nascent RNA folding and contribution to the variability of local mutation rates across the yeast genome.
- Gongwang Yu
- , Yao Liu
- & Jian-Rong Yang
-
Article
| Open Access The master energy homeostasis regulator PGC-1α exhibits an mRNA nuclear export function
The master energy homeostasis regulator PGC-1α exhibits an mRNA nuclear export function
PGC-1α is a master regulator activating the transcription of key genes controlling the cell’s energy production. Here the authors show that PGC-1α has a function in the NXF1-dependent nuclear export of mRNAs.
- Simeon R. Mihaylov
- , Lydia M. Castelli
- & Guillaume M. Hautbergue
-
Article
| Open Access Molecular basis of RNA-binding and autoregulation by the cancer-associated splicing factor RBM39
Molecular basis of RNA-binding and autoregulation by the cancer-associated splicing factor RBM39
RBM39 is an essential splicing factor for several cancer cells. Here, the authors described how RBM39 selects RNA at atomic level and autoregulates at the mRNA level by selecting the 3’-splice site of a poison exon.
- Sébastien Campagne
- , Daniel Jutzi
- & Frédéric H-T. Allain
-
Article
| Open Access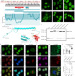 Pharmacological perturbation of the phase-separating protein SMNDC1
Pharmacological perturbation of the phase-separating protein SMNDC1
SMNDC1 is a splicing factor that binds arginine methylation with its Tudor domain. Here, the authors study the protein’s phase-separating behavior and develop small-molecule Tudor domain inhibitors that perturb SMNDC1 function.
- Lennart Enders
- , Marton Siklos
- & Stefan Kubicek
-
Article
| Open Access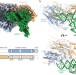 Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Aminoacyl-tRNA synthetases catalyze the ligation of amino acids to their cognate tRNAs. Here the authors report the cryo-EM structure of a human mitochondrial seryl-tRNA synthetase•mtRNASer complex showing how strong mutation pressure on mtRNA genes drove a rewiring of intermolecular recognition rules.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access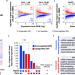 Elevated pre-mRNA 3′ end processing activity in cancer cells renders vulnerability to inhibition of cleavage and polyadenylation
Elevated pre-mRNA 3′ end processing activity in cancer cells renders vulnerability to inhibition of cleavage and polyadenylation
Cancer cells with elevated 3’ end processing activities are vulnerable to CPAi because of transcriptomic disturbances caused by alternative polyadenylation and transcriptional readthrough as well as DNA damages when cells are proliferative.
- Yange Cui
- , Luyang Wang
- & Bin Tian
-
Article
| Open Access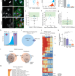 Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3
Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3
The authors show that loss-of-interaction with the nuclear importer, TNPO3, causes cytoplasmic mislocalization of RBM20 variants linked to severe cases of dilated cardiomyopathy. Restoring their nuclear localization alleviates the disease phenotype.
- Julia Kornienko
- , Marta Rodríguez-Martínez
- & Lars M. Steinmetz
-
Article
| Open Access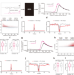 Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
Exon-intron boundary inhibits m6A deposition, enabling m6A distribution hallmark, longer mRNA half-life and flexible protein coding
m6A mRNA modification is not typically found near splice junctions in mRNAs. Here the authors show exon-intron boundary inhibits m6A deposition at ~100 nt region nearby splice site, enabling m6A distribution hallmark, more stable mRNA and flexible protein coding.
- Zhiyuan Luo
- , Qilian Ma
- & Shengdong Ke
-
Article
| Open Access Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
The neuronal RNA-binding protein Ptbp2 is known to regulate neuronal differentiation by modulating alternative splicing. Here, the authors reveal an additional role of cytosolic Ptbp2, which regulates axon growth by fine-tuning the mRNA transport and local synthesis of an RNA-binding protein hnRNP R.
- Saeede Salehi
- , Abdolhossein Zare
- & Michael Sendtner
-
Article
| Open Access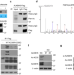 K235 acetylation couples with PSPC1 to regulate the m6A demethylation activity of ALKBH5 and tumorigenesis
K235 acetylation couples with PSPC1 to regulate the m6A demethylation activity of ALKBH5 and tumorigenesis
Deregulation of N6-methyladenosine (m6A) modification can contribute to the pathogenesis of cancers. Here the authors show that m6A demethylase ALKBH5 is acetylated at K235 by acetyltransferase KAT8 and interacts with RNA-binding protein PSCP1 to enhance m6A demethylation and promote tumorigenesis.
- Xiao-Lan Zhang
- , Xin-Hui Chen
- & Guang-Rong Yan
-
Article
| Open Access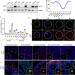 Maternal NAT10 orchestrates oocyte meiotic cell-cycle progression and maturation in mice
Maternal NAT10 orchestrates oocyte meiotic cell-cycle progression and maturation in mice
Generation of mature oocytes requires tight regulation of a discontinuous meiotic cell cycle. Here they show that the acetyltransferase Nat10 mediates modification of RNAs targeted for degradation and find that this process is essential for female oocyte meiosis and maturation.
- Xue Jiang
- , Yu Cheng
- & Jianqiang Bao
-
Article
| Open Access Splicing activates transcription from weak promoters upstream of alternative exons
Splicing activates transcription from weak promoters upstream of alternative exons
Few therapeutic strategies are able to upregulate gene expression. Here, the authors developed an approach to activate expression of human genes through small molecules and antisense oligonucleotides that modulate splicing.
- Maritere Uriostegui-Arcos
- , Steven T. Mick
- & Ana Fiszbein
-
Article
| Open Access Muscleblind-like proteins use modular domains to localize RNAs by riding kinesins and docking to membranes
Muscleblind-like proteins use modular domains to localize RNAs by riding kinesins and docking to membranes
RNA localization is mediated by kinesin motors and anchoring. However, mechanisms underlying specificity are unclear. Here, the authors find that MBNL protein’s zinc fingers prefer specific kinesins, and its unstructured tail mediates membrane anchoring.
- Ryan P. Hildebrandt
- , Kathryn R. Moss
- & Eric T. Wang
-
Article
| Open Access PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
Tracking protein-RNA interaction across cell types is challenging. Here, Ruan et al develop a dual-deaminase method called PIE-Seq, where protein targets are marked by both C-to-U and A-to-I RNA base editors, and apply it to 25 human RNA-binding proteins.
- Xiangbin Ruan
- , Kaining Hu
- & Xiaochang Zhang
-
Article
| Open Access LIS1 RNA-binding orchestrates the mechanosensitive properties of embryonic stem cells in AGO2-dependent and independent ways
LIS1 RNA-binding orchestrates the mechanosensitive properties of embryonic stem cells in AGO2-dependent and independent ways
LIS1 protein is important for brain development and stem cells’ survival. Here the authors show that LIS1 binds RNA and interact with RNA-binding proteins regulating the physical properties of mouse embryonic stem cells.
- Aditya Kshirsagar
- , Svetlana Maslov Doroshev
- & Orly Reiner
-
Article
| Open Access Rio1 downregulates centromeric RNA levels to promote the timely assembly of structurally fit kinetochores
Rio1 downregulates centromeric RNA levels to promote the timely assembly of structurally fit kinetochores
Kinetochores assemble on centromeres via histone H3 variant CENP-A and low levels of centromere transcripts (cenRNAs). Here the authors show the Rio1 kinase limits cenRNA production by reducing RNAPII accessibility and promotes cenRNA degradation by the 5’− 3’exoribonuclease Rat1.
- Ksenia Smurova
- , Michela Damizia
- & Peter De Wulf
-
Article
| Open Access The conserved RNA-binding protein Seb1 promotes cotranscriptional ribosomal RNA processing by controlling RNA polymerase I progression
The conserved RNA-binding protein Seb1 promotes cotranscriptional ribosomal RNA processing by controlling RNA polymerase I progression
Ribosome biogenesis is influenced by the rate of RNAPI progression. Yet, mechanisms that control RNAPI elongation have remained elusive. Here, the authors show that the conserved protein Seb1 promotes cotranscriptional rRNA processing by controlling RNAPI progression.
- Maxime Duval
- , Carlo Yague-Sanz
- & François Bachand
-
Article
| Open Access Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Dawicki-McKenna and Felix et al comprehensively map binding and alternative splicing by PTBP2 in human brain and neurons, thus identifying splice switching therapeutic strategies for the neurodevelopmental disorder associated gene SYNGAP1.
- Jennine M. Dawicki-McKenna
- , Alex J. Felix
- & Benjamin L. Prosser
-
Article
| Open Access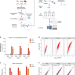 The RNA-binding protein landscapes differ between mammalian organs and cultured cells
The RNA-binding protein landscapes differ between mammalian organs and cultured cells
Characterization of RNA-binding proteins (RBPs) in tissues has been hampered by technical constraints. Here, the authors describe ex vivo eRIC, a method for global profiling of RBPs active in mammalian organs, and report comprehensive RBP atlases from mouse brain, kidney and liver.
- Joel I. Perez-Perri
- , Dunja Ferring-Appel
- & Matthias W. Hentze
-
Comment
| Open Access The problem of selection bias in studies of pre-mRNA splicing
The problem of selection bias in studies of pre-mRNA splicing
In this comment, the authors discuss the potentially widespread problem of selection bias in drawing biological conclusions from RNA sequencing data.
- Zachary W. Dwyer
- & Jeffrey A. Pleiss
-
Article
| Open Access The m6A reader YTHDC1 and the RNA helicase DDX5 control the production of rhabdomyosarcoma-enriched circRNAs
The m6A reader YTHDC1 and the RNA helicase DDX5 control the production of rhabdomyosarcoma-enriched circRNAs
Rhabdomyosarcoma (RMS) is the most diffused soft tissue sarcoma in children and adolescents. Herein, the authors identify the m6A machinery and the RNA helicase DDX5 as factors responsible for the increase of a subset of circRNAs in RMS, providing protein and RNA candidates for the study of its tumorigenicity.
- Dario Dattilo
- , Gaia Di Timoteo
- & Irene Bozzoni

 Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data
Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data