Featured
-
-
Article
| Open Access GWAS reveals determinants of mobilization rate and dynamics of an active endogenous retrovirus of cattle
GWAS reveals determinants of mobilization rate and dynamics of an active endogenous retrovirus of cattle
Endogenous retroviruses constitute 5–10% of mammalian genome space. This study characterize the bovine ERVK[2-1- LTR] clade showing that its activity varies between individuals as a function of the number of inherited autonomous elements, yet that most de novo insertions are non-autonomous elements lacking functional genes.
- Lijing Tang
- , Benjamin Swedlund
- & Carole Charlier
-
Article
| Open Access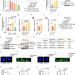 Nuclear cGAS restricts L1 retrotransposition by promoting TRIM41-mediated ORF2p ubiquitination and degradation
Nuclear cGAS restricts L1 retrotransposition by promoting TRIM41-mediated ORF2p ubiquitination and degradation
Zhen and colleagues show that nuclear cGAS represses L1 retrotransposition to stabilize the genome by enhancing the interaction between ORF2p and the E3 ligase TRIM41 upon DNA damage, which leads to the ubiquitination and degradation of ORF2p.
- Zhengyi Zhen
- , Yu Chen
- & Zhiyong Mao
-
Article
| Open Access Downregulation of transposable elements extends lifespan in Caenorhabditis elegans
Downregulation of transposable elements extends lifespan in Caenorhabditis elegans
Transposable elements in somatic cells become increasingly mobile during ageing. Here, the authors show that in the nematode Caenorhabditis elegans, downregulation of transposable elements extends lifespan, and that their increases with age are coupled with progressively growing N6-adenine methylation in these genetic loci.
- Ádám Sturm
- , Éva Saskői
- & Tibor Vellai
-
Article
| Open Access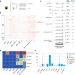 Extrachromosomal circular DNA and structural variants highlight genome instability in Arabidopsis epigenetic mutants
Extrachromosomal circular DNA and structural variants highlight genome instability in Arabidopsis epigenetic mutants
Epigenetic control of extrachromosomal circular DNA (eccDNA) compartment and the relationships between eccDNA and plant genome stability remain unclear. Here, the authors investigate eccDNA and structural variations in Arabidopsis epigenetic mutants to reveal the eccDNA repertoire and its impact on genome stability.
- Panpan Zhang
- , Assane Mbodj
- & Marie Mirouze
-
Article
| Open Access IS21 family transposase cleaved donor complex traps two right-handed superhelical crossings
IS21 family transposase cleaved donor complex traps two right-handed superhelical crossings
Spínola-Amilibia et al. present the cryo-EM structure of the IS21 transposase in complex with the donor DNA and show that IstA recognizes the transposon ends with a highly intertwined configuration to facilitate the strand-transfer reaction.
- Mercedes Spínola-Amilibia
- , Lidia Araújo-Bazán
- & Ernesto Arias-Palomo
-
Article
| Open Access Multivalent interactions essential for lentiviral integrase function
Multivalent interactions essential for lentiviral integrase function
The authors determined high-resolution cryo-EM structures of the lentiviral intasome — the nucleoprotein complex that inserts viral DNA into a host chromosome — and show that the architecture comprising 16 integrase subunits is critical for its function.
- Allison Ballandras-Colas
- , Vidya Chivukula
- & Peter Cherepanov
-
Article
| Open Access A point mutation in HIV-1 integrase redirects proviral integration into centromeric repeats
A point mutation in HIV-1 integrase redirects proviral integration into centromeric repeats
HIV-1 integration sites are biased towards actively transcribed genes, likely mediated by binding of the viral integrase (IN) protein to host factors. Here, Winans et al. show that the K258R point mutation in IN eredirects viral DNA integration to the centromeres of host chromosomes, which may affect HIV latency.
- Shelby Winans
- , Hyun Jae Yu
- & Stephen P. Goff
-
Article
| Open Access Structural basis of Ty3 retrotransposon integration at RNA Polymerase III-transcribed genes
Structural basis of Ty3 retrotransposon integration at RNA Polymerase III-transcribed genes
Ty3 retrotransposon integrates with an exquisite specificity upstream of RNA Polymerase III-transcribed genes, such as transfer RNAs. Here the authors resolve a cryo-EM structure of an active Ty3 intasome in complex with a TFIIIB-bound tRNA promoter, shedding light into the molecular determinants of harmless retrotransposition.
- Guillermo Abascal-Palacios
- , Laura Jochem
- & Alessandro Vannini
-
Article
| Open Access Structure of a Ty1 restriction factor reveals the molecular basis of transposition copy number control
Structure of a Ty1 restriction factor reveals the molecular basis of transposition copy number control
In Saccharomyces cerevisiae, unchecked proliferation of Ty1 retrotransposons is controlled by the process of copy number control (CNC), which requires the p22/p18 protein, translated from an internal transcript within the Ty1 GAG gene. Here, the authors present the 2.8 Å crystal structure of a minimal p18 from Ty1-Gag that is able to restrict Ty1 transposition and identify two dimer interfaces in p18, whose roles were probed by mutagenesis both in vitro and in vivo. As p22/p18 contains only one of two conserved domains required for retroelement Gag assembly, they propose that p22/p18-Gag interactions block the Ty1 virus-like particle assembly pathway, resulting in defective particles incapable of supporting retrotransposition.
- Matthew A. Cottee
- , Sean L. Beckwith
- & Ian A. Taylor
-
Article
| Open Access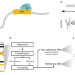 Cas9 targeted enrichment of mobile elements using nanopore sequencing
Cas9 targeted enrichment of mobile elements using nanopore sequencing
Mobile element insertions (MEIs) are a source of repetitive genetic variation and can lead to genetic disorders. Here the authors use Cas9-targeted nanopore sequencing to efficiently saturate enrichment for known and non-reference MEIs.
- Torrin L. McDonald
- , Weichen Zhou
- & Alan P. Boyle
-
Article
| Open Access L1 retrotransposons exploit RNA m6A modification as an evolutionary driving force
L1 retrotransposons exploit RNA m6A modification as an evolutionary driving force
L1 is a group of active retrotransposons in humans. Here the authors show that m6A modifications on L1 RNA increase translation efficiency and retrotransposition in human cells. M6A motifs are more enriched in evolutionary young L1s.
- Sung-Yeon Hwang
- , Hyunchul Jung
- & Kwangseog Ahn
-
Article
| Open Access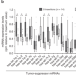 The tumor suppressor microRNA let-7 inhibits human LINE-1 retrotransposition
The tumor suppressor microRNA let-7 inhibits human LINE-1 retrotransposition
Human Long INterspersed Element class 1 (LINE-1) elements are expressed and mobilized in many types of cancer, contributing to malignancy. Here the authors show that the tumor suppressor microRNA let-7 targets the LINE-1 mRNA and reduces LINE-1 mobilization.
- Pablo Tristán-Ramos
- , Alejandro Rubio-Roldan
- & Sara R. Heras
-
Article
| Open Access Structural basis of seamless excision and specific targeting by piggyBac transposase
Structural basis of seamless excision and specific targeting by piggyBac transposase
PiggyBac is a transposon used in genome engineering that does not leave excision footprints. Here the authors determine the structures of two complexes in which the piggyBac transposase is bound to DNA representing different steps of the transposition reaction, providing a basis for how the transposition reaction proceeds.
- Qiujia Chen
- , Wentian Luo
- & Fred Dyda
-
Article
| Open Access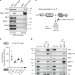 The Mi-2 nucleosome remodeler and the Rpd3 histone deacetylase are involved in piRNA-guided heterochromatin formation
The Mi-2 nucleosome remodeler and the Rpd3 histone deacetylase are involved in piRNA-guided heterochromatin formation
In S. pombe, small non-coding RNA mediates heterochromatin formation by recruiting the nucleosome remodeling and histone deacetylase complex. Here, the authors show that fly nucleosome remodeler Mi-2 and histone deacetylase Rpd3 are involved in piRNA-dependent transcriptional silencing of transposable elements.
- Bruno Mugat
- , Simon Nicot
- & Séverine Chambeyron
-
Article
| Open Access Maximizing the ovarian reserve in mice by evading LINE-1 genotoxicity
Maximizing the ovarian reserve in mice by evading LINE-1 genotoxicity
Mammals lose up to 80% of their finite oocyte supply during fetal development. Here the authors interrogate mechanisms of fetal oocyte attrition in mice, driven by the simultaneous upregulation of LINE-1 retrotransposon activity and inhibit these mechanisms to increase the functional ovarian reserve.
- Marla E. Tharp
- , Safia Malki
- & Alex Bortvin
-
Article
| Open Access Decoding the 5′ nucleotide bias of PIWI-interacting RNAs
Decoding the 5′ nucleotide bias of PIWI-interacting RNAs
PIWI-interacting RNAs (piRNAs) are regulatory RNAs that bind to PIWI proteins to control transposons and maintain genome integrity. Here the authors characterized their binding specificity and reveal the 5′ nucleotide bias of the Drosophila Piwi protein, through mutation of its specificity loop.
- Chad B. Stein
- , Pavol Genzor
- & Astrid D. Haase
-
Article
| Open Access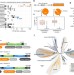 Recurrent acquisition of cytosine methyltransferases into eukaryotic retrotransposons
Recurrent acquisition of cytosine methyltransferases into eukaryotic retrotransposons
Cytosine methyltransferases (DNMTs) often silence transposons in eukaryotic genomes. Here the authors describe the recurrent acquisition of DNMTs by transposons from two distantly-related eukaryotes and suggest that methylation of CG dinucleotides by transposon DNMTs could modify the host epigenome in dinoflagellates.
- Alex de Mendoza
- , Amandine Bonnet
- & Ryan Lister
-
Article
| Open Access Diverse genetic error modes constrain large-scale bio-based production
Diverse genetic error modes constrain large-scale bio-based production
The declining performance of scale-up bioreactor cultures is commonly attributed to phenotypic and physical heterogeneities. Here, the authors reveal multiple recurring intra-pathway error modes that limit engineered E. coli mevalonic acid production over time- and industrial-scale fermentations.
- Peter Rugbjerg
- , Nils Myling-Petersen
- & Morten O. A. Sommer
-
Review Article
| Open Access The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity
The emergence of piRNAs against transposon invasion to preserve mammalian genome integrity
Transposable elements can be activated during germ cell maturation, potentially leading to genome instability and rewiring of the genetic circuitry. In this review, the authors discuss how the piRNA machinery suppresses these elements to ensure accurate spermatogenesis.
- Christina Ernst
- , Duncan T. Odom
- & Claudia Kutter
-
Article
| Open Access A somatic piRNA pathway in the Drosophila fat body ensures metabolic homeostasis and normal lifespan
A somatic piRNA pathway in the Drosophila fat body ensures metabolic homeostasis and normal lifespan
The Piwi-interacting RNA (piRNA) pathway is known to suppress transposable elements in gonadal tissues. Here the authors provide evidence for a functional piRNA pathway in the somatic cells of theDrosophilafat body with roles in metabolism, immunological function and overall health.
- Brian C. Jones
- , Jason G. Wood
- & Stephen L. Helfand
-
Article
| Open Access A Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes
A Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes
Helitron elements are proposed rolling-circle transposons in eukaryotic genomes, but experimental evidence for their transposition has been lacking. Here, Grabundzija et al. reconstruct an active Helitron from bats which they name Helraiser, and characterize its mechanism of transposition in cell-free reactions and in human cell cultures in vitro.
- Ivana Grabundzija
- , Simon A. Messing
- & Zoltán Ivics
-
Article
| Open Access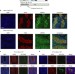 MORC1 represses transposable elements in the mouse male germline
MORC1 represses transposable elements in the mouse male germline
The Microrchidia (Morc) family of GHKL ATPases are important repressors of transposons and other DNA-methylated and silent genes in A. thaliana. Here, the authors show that MORC1 is responsible for repression and methylation of specific classes of transposons in the mouse male germline.
- William A. Pastor
- , Hume Stroud
- & Steven E. Jacobsen
-
Article |
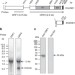 Recombinant SINEs are formed at high frequency during induced retrotransposition in vivo
Recombinant SINEs are formed at high frequency during induced retrotransposition in vivo
SINEs are retrotransposons that insert exact copies of themselves into genomes. Using a marked copy of a SINE, Yadavet al. show that the sequences of newly transposed SINEs are a combination of marked and existing SINEs, suggesting a mechanism for the formation of mosaic SINEs.
- Vijay Pal Yadav
- , Prabhat Kumar Mandal
- & Sudha Bhattacharya

 Condensin-mediated restriction of retrotransposable elements facilitates brain development in Drosophila melanogaster
Condensin-mediated restriction of retrotransposable elements facilitates brain development in Drosophila melanogaster