Featured
-
-
Article
| Open Access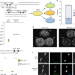 A ‘parameiosis’ drives depolyploidization and homologous recombination in Candida albicans
A ‘parameiosis’ drives depolyploidization and homologous recombination in Candida albicans
Mating of Candida albicans produces tetraploid products that return to the diploid state via a non-meiotic process known as concerted chromosome loss (CCL). Here, Anderson et al. show high recombination rates during CCL and identify factors that are essential for chromosome stability and recombination during CCL.
- Matthew Z. Anderson
- , Gregory J. Thomson
- & Richard J. Bennett
-
Article
| Open Access An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
Invasive non-typhoidal Salmonella (iNTS) infections are dominated by antibiotic resistant isolates of the sequence type (ST) 313. Here, the authors identify the ST313 sublineage II.1 in the Democratic Republic of the Congo exhibiting extensive drug resistance and genetic signatures potentially associated with host adaptation.
- Sandra Van Puyvelde
- , Derek Pickard
- & Stijn Deborggraeve
-
Article
| Open Access Derailing the aspartate pathway of Mycobacterium tuberculosis to eradicate persistent infection
Derailing the aspartate pathway of Mycobacterium tuberculosis to eradicate persistent infection
Amino acid biosynthetic pathways are an attractive alternative to treat chronic infections such as Mycobacterium tuberculosis (Mtb). Here, the authors investigate the metabolic response to disruption of the aspartate pathway in persistent Mtb and identify essential enzymes as potential new targets for drug development.
- Erik J. Hasenoehrl
- , Dannah Rae Sajorda
- & Michael Berney
-
Article
| Open Access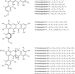 A role for antibiotic biosynthesis monooxygenase domain proteins in fidelity control during aromatic polyketide biosynthesis
A role for antibiotic biosynthesis monooxygenase domain proteins in fidelity control during aromatic polyketide biosynthesis
Formicapyridines are similar to pentacyclic fasamycin and formicamycin aromatic polyketides but with a pyridine moiety. Here the authors rationally mutate the biosynthetic gene cluster to increase production and identify a non-catalytic role for the ABM superfamily of proteins.
- Zhiwei Qin
- , Rebecca Devine
- & Barrie Wilkinson
-
Article
| Open Access A modular effector with a DNase domain and a marker for T6SS substrates
A modular effector with a DNase domain and a marker for T6SS substrates
Bacteria deliver toxic effectors via type VI secretion systems (T6SSs) to dominate competitors. Here, the authors identify a Vibrio antibacterial effector that contains a new DNase toxin domain and a domain of unknown function that can be used as a marker to identify new T6SS effectors.
- Biswanath Jana
- , Chaya M. Fridman
- & Dor Salomon
-
Article
| Open Access Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
The genome of influenza is often incomplete in infected cells, but the implications for infection remain unclear. Here, Jacobs et al. show that an average of 3.6 particles is necessary for productive infection and that coinfection supports efficient complementation within a host but not upon transmission to a new host.
- Nathan T. Jacobs
- , Nina O. Onuoha
- & Anice C. Lowen
-
Article
| Open Access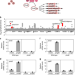 The Salmonella virulence protein MgtC promotes phosphate uptake inside macrophages
The Salmonella virulence protein MgtC promotes phosphate uptake inside macrophages
The virulence factor MgtC is essential for intracellular macrophage survival of Salmonella enterica. Here, the authors show that MgtC targets the PhoB/PhoR regulatory system leading to phosphate uptake inside macrophages and that both phoR mutation and phoB deletion renders Salmonella hypervirulent in mice.
- Soomin Choi
- , Eunna Choi
- & Eun-Jin Lee
-
Article
| Open Access Two-step chromosome segregation in the stalked budding bacterium Hyphomonas neptunium
Two-step chromosome segregation in the stalked budding bacterium Hyphomonas neptunium
In bacteria, DNA replication and segregation are commonly coupled. Here, by investigating the dynamics of these processes in the marine bacterium Hyphomonas neptunium, the authors unravel a two-step chromosomal segregation process reminiscent of eukaryotic mitosis, providing insights into the evolution of bacterial cell organization.
- Alexandra Jung
- , Anne Raßbach
- & Martin Thanbichler
-
Article
| Open Access Genomic epidemiology of syphilis reveals independent emergence of macrolide resistance across multiple circulating lineages
Genomic epidemiology of syphilis reveals independent emergence of macrolide resistance across multiple circulating lineages
Syphilis is caused by the bacterium Treponema pallidum subspecies pallidum (TPA), and incidence has risen recently in many countries. Here, Beale et al. provide whole-genome TPA sequences from 73 clinical samples and show how antimicrobial resistance emerged independently in circulating lineages.
- Mathew A. Beale
- , Michael Marks
- & Nicholas R. Thomson
-
Article
| Open Access Mutation bias and GC content shape antimutator invasions
Mutation bias and GC content shape antimutator invasions
Mutators are expected to re-evolve low mutation rates to reduce deleterious load, but empirical evidence is mixed. Here, the authors show that load can vary across mutators and genetic backgrounds, which their simulations suggest can substantially alter antimutator dynamics.
- Alejandro Couce
- & Olivier Tenaillon
-
Article
| Open Access Direct visualization of a molecular handshake that governs kin recognition and tissue formation in myxobacteria
Direct visualization of a molecular handshake that governs kin recognition and tissue formation in myxobacteria
Many organisms, including the bacterium Myxococcus xanthus, regulate their social life through kin recognition. Here, Cao and Wall show that these bacteria use a polymorphic and fluid cell-surface receptor to recognize and assemble kin cells into a cooperative multicellular community that resembles a tissue.
- Pengbo Cao
- & Daniel Wall
-
Article
| Open Access Essentiality of sterol synthesis genes in the planctomycete bacterium Gemmata obscuriglobus
Essentiality of sterol synthesis genes in the planctomycete bacterium Gemmata obscuriglobus
Sterols play essential functions in eukaryotic cell membranes, but are produced by few bacterial species. Here, the authors show that they are essential for growth of the planctomycete bacterium Gemmata obscuriglobus.
- Elena Rivas-Marin
- , Sean Stettner
- & Damien P. Devos
-
Article
| Open Access HIV-1 DNA sequence diversity and evolution during acute subtype C infection
HIV-1 DNA sequence diversity and evolution during acute subtype C infection
The dynamics of HIV-1 DNA sequences early after HIV-1 transmission remains poorly characterized. Here, the authors perform a longitudinal evaluation of HIV-1 DNA sequences in subtype C-infected individuals during acute infection, providing a landscape of the nature and evolution of the very early viral genome.
- Guinevere Q. Lee
- , Kavidha Reddy
- & Mathias Lichterfeld
-
Article
| Open Access Deterministic processes structure bacterial genetic communities across an urban landscape
Deterministic processes structure bacterial genetic communities across an urban landscape
Disease transmission is particularly complex at the human-livestock-wildlife interface. Here the authors sample E. coli from wild birds near households in Nairobi and show that antimicrobial resistance gene diversity is correlated with human and lifestock density, while virulence gene diversity is correlated with avian species richness.
- J. M. Hassell
- , M. J. Ward
- & E. M. Fèvre
-
Article
| Open Access Genomic signatures of heterokaryosis in the oomycete pathogen Bremia lactucae
Genomic signatures of heterokaryosis in the oomycete pathogen Bremia lactucae
The oomycete Bremia lactucae is a highly variable pathogen that causes lettuce downy mildew. Here, the authors generate a high-quality genome assembly for B. lactucae, detect a high prevalence of heterokaryosis, and investigate its pathogenic consequences.
- Kyle Fletcher
- , Juliana Gil
- & Richard Michelmore
-
Article
| Open Access Hypervirulent Listeria monocytogenes clones’ adaption to mammalian gut accounts for their association with dairy products
Hypervirulent Listeria monocytogenes clones’ adaption to mammalian gut accounts for their association with dairy products
Here, Maury et al. show that hypervirulent Listeria monocytogenes (Lm) clones associated to dairy products exhibit higher adaptation to the mammalian gut environment, while hypovirulent clones persist in food processing environment, suggesting a relationship between Lm pathogenic potential and niche adaptation.
- Mylène M. Maury
- , Hélène Bracq-Dieye
- & Marc Lecuit
-
Article
| Open Access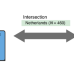 Joint sequencing of human and pathogen genomes reveals the genetics of pneumococcal meningitis
Joint sequencing of human and pathogen genomes reveals the genetics of pneumococcal meningitis
Streptococcus pneumoniae is a causative agent of meningitis and bacteremia. In a combined pathogen and host GWAS, Lees et al. find that host genetic variation is associated with both susceptibility and severity of pneumococcal meningitis, and specific bacterial genetic variation associated with susceptibility.
- John A. Lees
- , Bart Ferwerda
- & Diederik van de Beek
-
Article
| Open Access GWAS for quantitative resistance phenotypes in Mycobacterium tuberculosis reveals resistance genes and regulatory regions
GWAS for quantitative resistance phenotypes in Mycobacterium tuberculosis reveals resistance genes and regulatory regions
Resistance to antibiotics hampers the treatment of infectious diseases such as tuberculosis (TB). Here, Farhat et al. perform genome-wide association testing for minimum inhibitory concentration (MIC) of 12 anti-TB drugs in whole-genome sequenced clinical M. tuberculosis isolates and identify 13 genomic loci.
- Maha R. Farhat
- , Luca Freschi
- & Megan Murray
-
Article
| Open Access A bacterial checkpoint protein for ribosome assembly moonlights as an essential metabolite-proofreading enzyme
A bacterial checkpoint protein for ribosome assembly moonlights as an essential metabolite-proofreading enzyme
Adventitious oxidation of erythrose-4-phosphate generates 4-phosphoerythronate, which is detoxified by metabolite-proofreading phosphatases in eukaryotes. Here, Sachla & Helmann show that a similar function is carried out in Bacillus subtilis by a checkpoint protein involved in ribosome assembly.
- Ankita J. Sachla
- & John D. Helmann
-
Article
| Open Access A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
Certain archaea appear to metabolize fatty acids, but the regulation of these pathways is unclear. Here, Wang et al. provide genetic, functional and structural evidence supporting that a TetR-family transcriptional regulator is involved in regulation of fatty acid metabolism in Sulfolobus acidocaldarius.
- Kun Wang
- , David Sybers
- & Eveline Peeters
-
Article
| Open Access Pervasive function and evidence for selection across standing genetic variation in S. cerevisiae
Pervasive function and evidence for selection across standing genetic variation in S. cerevisiae
Genetic architecture underlies the complexity of heritable traits. Here, the authors perform high-resolution genetic mapping of metabolic traits in S. cerevisiae and show evidence for selection across standing genetic variation.
- Christopher M. Jakobson
- , Richard She
- & Daniel F. Jarosz
-
Article
| Open Access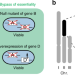 Systematic analysis reveals the prevalence and principles of bypassable gene essentiality
Systematic analysis reveals the prevalence and principles of bypassable gene essentiality
An essential gene may become non-essential when another gene is mutated. Here, the authors investigate this type of digenic interaction, termed ‘bypass of essentiality’, in the fission yeast Schizosaccharomyces pombe, and show that bypassable essential genes are common and share certain features.
- Jun Li
- , Hai-Tao Wang
- & Li-Lin Du
-
Article
| Open Access Stress-induced inactivation of the Staphylococcus aureus purine biosynthesis repressor leads to hypervirulence
Stress-induced inactivation of the Staphylococcus aureus purine biosynthesis repressor leads to hypervirulence
PurR acts as transcriptional repressor of purine biosynthesis genes in some bacterial species. Here, the authors show that purR mutations can arise in Staphylococcus aureus upon exposure to stress, leading to upregulation of fibronectin-binding proteins and increased virulence.
- Mariya I. Goncheva
- , Ronald S. Flannagan
- & David E. Heinrichs
-
Article
| Open Access Identification and characterization of a direct activator of a gene transfer agent
Identification and characterization of a direct activator of a gene transfer agent
Gene transfer agents (GTAs) are ‘domesticated’ bacteriophages that can transfer any genes between bacteria. Here, Paul Fogg identifies a protein that directly regulates transcription of GTA genes and whose expression is in turn controlled by a global cell-cycle regulator and a quorum-sensing regulator.
- Paul C. M. Fogg
-
Article
| Open Access Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Gain of function methods based on gene overexpression are not easily applied to high-throughput screening across different experimental conditions. Here, the authors present Dub-seq, which separates overexpression library characterization from functional screening and uses random DNA barcodes to increase the experimental throughput.
- Vivek K. Mutalik
- , Pavel S. Novichkov
- & Adam P. Arkin
-
Article
| Open Access Genome-wide association analyses of invasive pneumococcal isolates identify a missense bacterial mutation associated with meningitis
Genome-wide association analyses of invasive pneumococcal isolates identify a missense bacterial mutation associated with meningitis
Meningitis is a severe form of invasive pneumococcal disease (IPD). To study the contribution of bacterial genomic variation, here Li et al. perform whole genome sequencing of pneumococcal isolates from IPD patients and identify an association for higher risk of meningitis with a pbp1bA641C variant
- Yuan Li
- , Benjamin J. Metcalf
- & Bernard W. Beall
-
Article
| Open Access Metaepigenomic analysis reveals the unexplored diversity of DNA methylation in an environmental prokaryotic community
Metaepigenomic analysis reveals the unexplored diversity of DNA methylation in an environmental prokaryotic community
Our knowledge of DNA methylation systems in prokaryotes is mostly limited to those of culturable microbes. Here, Hiraoka et al. analyse DNA methylation patterns in metagenomic data from a microbial community, revealing new methylated motifs and experimentally validating the methyltransferases’ specificities.
- Satoshi Hiraoka
- , Yusuke Okazaki
- & Wataru Iwasaki
-
Article
| Open Access RETRACTED ARTICLE: A switch in the poly(dC)/RmlB complex regulates bacterial persister formation
RETRACTED ARTICLE: A switch in the poly(dC)/RmlB complex regulates bacterial persister formation
The mechanisms underlying bacterial persisters formation remain poorly understood. Here, Chen et al. identify a complex formed by extracellular poly(dC) and the binding protein RmlB that controls Pseudomonas aeruginosa persister formation in response to environmental stimuli.
- Xu Chen
- , Gen Li
- & Kouhong Sun
-
Article
| Open Access Antibiotic sensitivity reveals that wall teichoic acids mediate DNA binding during competence in Bacillus subtilis
Antibiotic sensitivity reveals that wall teichoic acids mediate DNA binding during competence in Bacillus subtilis
Natural genetic transformation in bacteria requires DNA binding at the surface of competent cells. Here, Mirouze et al. show that wall teichoic acids are specifically produced or modified during competence in Bacillus subtilis and promote (directly or indirectly) DNA binding at the cell surface.
- Nicolas Mirouze
- , Cécile Ferret
- & Rut Carballido-López
-
Article
| Open Access Proteolysis of histidine kinase VgrS inhibits its autophosphorylation and promotes osmostress resistance in Xanthomonas campestris
Proteolysis of histidine kinase VgrS inhibits its autophosphorylation and promotes osmostress resistance in Xanthomonas campestris
Bacterial histidine kinases (HKs) play key roles in the response to stimuli and are regulated by reversible phosphorylation. Here, the authors show that the activity of a HK in the plant pathogen Xanthomonas campestris is modulated by irreversible, proteolytic modification in response to osmostress.
- Chao-Ying Deng
- , Huan Zhang
- & Wei Qian
-
Article
| Open Access Gene inversion potentiates bacterial evolvability and virulence
Gene inversion potentiates bacterial evolvability and virulence
Head-on replication-transcription collisions occur within genes encoded on the lagging DNA strand. Here, the authors show that a large number of originally co-oriented (leading strand) genes have inverted to the head-on orientation, increasing both gene-specific mutation rates, and the overall evolvability of several bacterial pathogens.
- Christopher N. Merrikh
- & Houra Merrikh
-
Article
| Open Access Machine learning and structural analysis of Mycobacterium tuberculosis pan-genome identifies genetic signatures of antibiotic resistance
Machine learning and structural analysis of Mycobacterium tuberculosis pan-genome identifies genetic signatures of antibiotic resistance
Mycobacterium tuberculosis exhibits complex evolution of antimicrobial resistance (AMR). Here, the authors perform machine learning and structural analysis to identify signatures of AMR evolution to 13 antibiotics.
- Erol S. Kavvas
- , Edward Catoiu
- & Bernhard O. Palsson
-
Article
| Open Access Convergent evolution of complex genomic rearrangements in two fungal meiotic drive elements
Convergent evolution of complex genomic rearrangements in two fungal meiotic drive elements
Meiotic drive elements are selfish genetic elements that mediate a skew in their sexual transmission from parent to offspring. Here, the authors sequence and analyze large and complex genomic regions associated with meiotic drive elements in the fungus Neurospora intermedia.
- Jesper Svedberg
- , Sara Hosseini
- & Hanna Johannesson
-
Article
| Open Access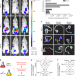 Host-associated niche metabolism controls enteric infection through fine-tuning the regulation of type 3 secretion
Host-associated niche metabolism controls enteric infection through fine-tuning the regulation of type 3 secretion
Infection of mice with Citrobacter rodentium is a common model of infection with attaching-and-effacing pathogens. Here, Connolly et al. analyse the transcriptome of C. rodentium during mouse infection, showing host-induced coordinated upregulation of virulence factors and 1,2-propanediol metabolism.
- James P. R. Connolly
- , Sabrina L. Slater
- & Andrew J. Roe
-
Article
| Open Access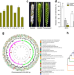 Wheat microbiome bacteria can reduce virulence of a plant pathogenic fungus by altering histone acetylation
Wheat microbiome bacteria can reduce virulence of a plant pathogenic fungus by altering histone acetylation
The molecular mechanisms behind the interactions between bacteria and fungi are largely unclear. Here, Chen et al. show that a compound secreted by bacteria from the wheat head microbiome inhibits growth and virulence of a plant pathogenic fungus by manipulating fungal histone modification.
- Yun Chen
- , Jing Wang
- & Zhonghua Ma
-
Article
| Open Access Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination
Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination
Since the 1980s, hypervirulent clonal-group CG23 serotype K1 Klebsiella pneumoniae has been recognised as a prominent cause of community-acquired liver abscess and other severe infections. Here, the authors investigate the genomic evolutionary history of CG23 and suggest a new reference strain for CG23.
- Margaret M. C. Lam
- , Kelly L. Wyres
- & Kathryn E. Holt
-
Article
| Open Access Evolutionary trade-offs associated with loss of PmrB function in host-adapted Pseudomonas aeruginosa
Evolutionary trade-offs associated with loss of PmrB function in host-adapted Pseudomonas aeruginosa
Mutations in gene pmrB are found in Pseudomonas aeruginosa isolates from cystic fibrosis patients. Here, Bricio-Moreno et al. show in a mouse model of respiratory infection that the mutations enhance bacterial adherence to epithelial cells and resistance to lysozyme, but also increase antibiotic susceptibility.
- Laura Bricio-Moreno
- , Victoria H. Sheridan
- & Daniel R. Neill
-
Article
| Open Access Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance
Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance
Systemic investigation of the bacterial genome is essential for both basic microbiology and bioengineering. Here the authors demonstrate CRISPRi pooled screening as a high-throughput tool for identifying gene and phenotype associations in bacteria.
- Tianmin Wang
- , Changge Guan
- & Xin-Hui Xing
-
Article
| Open Access Anti-phage islands force their target phage to directly mediate island excision and spread
Anti-phage islands force their target phage to directly mediate island excision and spread
Mobile genetic elements called PLEs protect Vibrio cholerae from infection with phage ICP1 by unclear mechanisms. Here, McKitterick and Seed show that a PLE-encoded large serine recombinase exploits an ICP1 protein as a recombination directionality factor to excise this PLE in response to phage infection.
- Amelia C. McKitterick
- & Kimberley D. Seed
-
Article
| Open Access Analysis of 3800-year-old Yersinia pestis genomes suggests Bronze Age origin for bubonic plague
Analysis of 3800-year-old Yersinia pestis genomes suggests Bronze Age origin for bubonic plague
Yersinia pestis has caused infections (plague) in humans since the Early Bronze Age (5000 years ago). Here, Spyrou et al. reconstruct Y. pestis genomes from Late Bronze Age individuals, and find genomic evidence compatible with flea-mediated transmission causing bubonic plague.
- Maria A. Spyrou
- , Rezeda I. Tukhbatova
- & Johannes Krause
-
Article
| Open Access Gene flow contributes to diversification of the major fungal pathogen Candida albicans
Gene flow contributes to diversification of the major fungal pathogen Candida albicans
The fungal pathogen Candida albicans can undergo a parasexual process that may contribute to genetic diversity, but its actual relevance is unclear. Here, Ropars et al. analyse the genomic sequences of 182 C. albicans isolates collected worldwide and find evidence of gene flow and thus parasexuality in nature.
- Jeanne Ropars
- , Corinne Maufrais
- & Christophe d’Enfert
-
Article
| Open Access CRISPR-FRT targets shared sites in a knock-out collection for off-the-shelf genome editing
CRISPR-FRT targets shared sites in a knock-out collection for off-the-shelf genome editing
Genome editing requires precise targeting of loci with specific gRNAs. Here the authors introduce CRISPR-FRT, which targets flippase recognition sites, common in bacterial genetic collections, for fast off-the-shelf genome engineering.
- Toon Swings
- , David C. Marciano
- & Jan Michiels
-
Article
| Open Access Tracking HIV-1 recombination to resolve its contribution to HIV-1 evolution in natural infection
Tracking HIV-1 recombination to resolve its contribution to HIV-1 evolution in natural infection
Recombination contributes to HIV evolution in patients, but its identification can be difficult. Here, the authors develop a computational tool called RAPR to track recombination in patients, identify recombination hot spots, and show contribution of recombination to antibody escape.
- Hongshuo Song
- , Elena E. Giorgi
- & Feng Gao
-
Article
| Open Access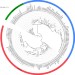 Culturing of female bladder bacteria reveals an interconnected urogenital microbiota
Culturing of female bladder bacteria reveals an interconnected urogenital microbiota
The female bladder seems to harbor a poorly characterized indigenous microbiota. Here, the authors isolate and genome-sequence 149 bacterial strains from catheterized urine of 77 women, generating a culture collection representing two thirds of the bacterial diversity within the samples.
- Krystal Thomas-White
- , Samuel C. Forster
- & Trevor D. Lawley
-
Article
| Open Access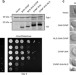 Viral proteins as a potential driver of histone depletion in dinoflagellates
Viral proteins as a potential driver of histone depletion in dinoflagellates
Dinoflagellates are known to use dinoflagellate-viral-nucleoproteins (DVNPs) in place of histones, yet this evolutionary transition is not well understood. Here, Irwin et al. use yeast expressing DVNP to show that DVNP displaces histones and that histone reduction allows cells to cope with DVNP.
- Nicholas A. T. Irwin
- , Benjamin J. E. Martin
- & LeAnn J. Howe
-
Article
| Open Access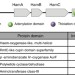 Biosynthesis of fragin is controlled by a novel quorum sensing signal
Biosynthesis of fragin is controlled by a novel quorum sensing signal
Fragin is a diazeniumdiolate metabolite with antifungal activity, produced by some bacteria. Here, Jenul et al. show that metal chelation is the molecular basis of fragin’s antifungal activity, and that a gene cluster directing fragin biosynthesis is also involved in the synthesis of a signal molecule.
- Christian Jenul
- , Simon Sieber
- & Leo Eberl
-
Article
| Open Access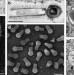 Tailed giant Tupanvirus possesses the most complete translational apparatus of the known virosphere
Tailed giant Tupanvirus possesses the most complete translational apparatus of the known virosphere
Giant viruses are the largest viruses of the known virosphere and their genetic analysis can provide insights into virus evolution. Here, the authors discover Tupanvirus, a unique giant virus that has an unusually long tail and contains the largest translational apparatus of the known virosphere.
- Jônatas Abrahão
- , Lorena Silva
- & Bernard La Scola
-
Article
| Open Access Biochemical mechanisms determine the functional compatibility of heterologous genes
Biochemical mechanisms determine the functional compatibility of heterologous genes
Sequence composition is thought to be a major factor governing the functionality of horizontally transferred genes. In contrast, Porse et al. show that phylogenetic origin, and the type of resistance mechanism, are major factors affecting the functionality of horizontally transferred antibiotic resistance genes.
- Andreas Porse
- , Thea S. Schou
- & Morten O. A. Sommer
-
Article
| Open Access Sensory deprivation in Staphylococcus aureus
Sensory deprivation in Staphylococcus aureus
Bacteria use two-component systems (TCSs) to sense and respond to environmental changes. Here, the authors show that Staphylococcus aureus can survive in the absence of all its 16 TCSs under growth arrest conditions, and each TCS seems to be sufficient to sense and respond to specific environmental clues.
- Maite Villanueva
- , Begoña García
- & Iñigo Lasa

 Phylogeography of the second plague pandemic revealed through analysis of historical Yersinia pestis genomes
Phylogeography of the second plague pandemic revealed through analysis of historical Yersinia pestis genomes