Featured
-
-
Article
| Open Access Transmission and dynamics of mother-infant gut viruses during pregnancy and early life
Transmission and dynamics of mother-infant gut viruses during pregnancy and early life
Gut ecosystem colonization impacts lifelong health. Here, authors track mother-infant gut viruses over time, reveal feeding’s influence on early viral colonization, and demonstrate the co-transmission of bacteriophages and bacteria from mothers to infants.
- Sanzhima Garmaeva
- , Trishla Sinha
- & Alexandra Zhernakova
-
Article
| Open Access Competition-driven eco-evolutionary feedback reshapes bacteriophage lambda’s fitness landscape and enables speciation
Competition-driven eco-evolutionary feedback reshapes bacteriophage lambda’s fitness landscape and enables speciation
Niche theory is often invoked to explain biodiversity, but it does not explain how species evolve to exploit unique niches. Using a combination of experimental and computational approaches, this study shows that resource competition can deform fitness landscapes, opening new pathways that promote ecological speciation.
- Michael B. Doud
- , Animesh Gupta
- & Justin R. Meyer
-
Article
| Open Access Co-option of a non-retroviral endogenous viral element in planthoppers
Co-option of a non-retroviral endogenous viral element in planthoppers
Non-retroviral endogenous viral elements are widely dispersed in eukaryotic genomes, but their functions remain largely unknown. Here, Huang et al show that one such element in planthoppers has been co-opted and contributes to insect fitness..
- Hai-Jian Huang
- , Yi-Yuan Li
- & Jun-Min Li
-
Article
| Open Access Genomic epidemiology offers high resolution estimates of serial intervals for COVID-19
Genomic epidemiology offers high resolution estimates of serial intervals for COVID-19
The serial interval (time between symptom onset in an infector and infectee) is usually estimated from contact tracing data, but this is not always available. Here, the authors develop a method for estimation of serial intervals using whole genome sequencing data and apply it data from clusters of SARS-CoV-2 in Victoria, Australia.
- Jessica E. Stockdale
- , Kurnia Susvitasari
- & Caroline Colijn
-
Article
| Open Access Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Certain bats species have previously been identified as ancestral sources of coronaviruses that infect humans but there is limited data on the genomic diversity or zoonotic potential of viruses infecting bats in the UK. Here, the authors use deep sequencing and in vitro assays to characterise coronaviruses recovered from 48 bat faecal samples.
- Cedric C. S. Tan
- , Jahcub Trew
- & Vincent Savolainen
-
Article
| Open Access Virus diversity, wildlife-domestic animal circulation and potential zoonotic viruses of small mammals, pangolins and zoo animals
Virus diversity, wildlife-domestic animal circulation and potential zoonotic viruses of small mammals, pangolins and zoo animals
Monitoring the diversity of viruses infecting animals is important for assessing zoonotic risk. Here, the authors use metatranscriptomics to characterise the viromes of small mammals, pangolins, and zoo animals in China to identify potentially zoonotic viruses.
- Xinyuan Cui
- , Kewei Fan
- & Yongyi Shen
-
Article
| Open Access High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection
High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection
Here, by high-resolution SARS-CoV-2 sequencing, genomic and transcriptomic analyses from tissue samples, Normandin et al. investigate viral dynamics in fatal cases of COVID-19, revealing persistent infection in distinct anatomical sites, including the heart and testis.
- Erica Normandin
- , Melissa Rudy
- & Isaac H. Solomon
-
Article
| Open Access Rapid transmission and tight bottlenecks constrain the evolution of highly transmissible SARS-CoV-2 variants
Rapid transmission and tight bottlenecks constrain the evolution of highly transmissible SARS-CoV-2 variants
Here, by sequencing viruses from individuals in multiple households, Bendall et al. find that SARS-CoV-2 transmission bottleneck does not vary between individuals infected with pre-variant lineages and those infected with highly transmissible Alpha, Delta, or Omicron variants, suggesting these tight bottlenecks will limit the spread of new mutations.
- Emily E. Bendall
- , Amy P. Callear
- & Adam S. Lauring
-
Article
| Open Access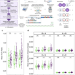 Characterisation of SARS-CoV-2 genomic variation in response to molnupiravir treatment in the AGILE Phase IIa clinical trial
Characterisation of SARS-CoV-2 genomic variation in response to molnupiravir treatment in the AGILE Phase IIa clinical trial
Molnupiravir is an antiviral that forces lethal error catastrophe in SARS-CoV-2 RNAs. Here, the authors confirm the mechanism of action of molnupiravir in humans using samples obtained from the UK’s AGILE phase IIa clinical trial investigating the antiviral efficacy of the drug against SARS-CoV-2. No treatment-associated SARS-CoV-2 mutations were identified.
- I’ah Donovan-Banfield
- , Rebekah Penrice-Randal
- & Thomas Fletcher
-
Article
| Open Access Experimental validation that human microbiome phages use alternative genetic coding
Experimental validation that human microbiome phages use alternative genetic coding
Previous bioinformatic analyses have indicated that bacteriophages can use genetic codes different from those of their host bacteria. Here, Peters et al. use metaproteomics to provide experimental evidence of reassignment of stop codon TAG to glutamine in phages found in the human gut microbiome.
- Samantha L. Peters
- , Adair L. Borges
- & Robert L. Hettich
-
Article
| Open Access Predicting the evolution of the Lassa virus endemic area and population at risk over the next decades
Predicting the evolution of the Lassa virus endemic area and population at risk over the next decades
It is currently unknown how climate and land use changes could affect the endemic area of Lassa virus, a zoonotic pathogen responsible for Lassa fever. Here, the authors show that by 2070, new regions in Africa will likely become ecologically suitable for Lassa virus, drastically increasing the population living in conditions favourable for virus circulation.
- Raphaëlle Klitting
- , Liana E. Kafetzopoulou
- & Simon Dellicour
-
Article
| Open Access Structural characterization of a soil viral auxiliary metabolic gene product – a functional chitosanase
Structural characterization of a soil viral auxiliary metabolic gene product – a functional chitosanase
Metagenomics is revealing auxiliary metabolic genes (AMGs) in soil viral genomes. Here, authors solve the crystal structure for a soil viral AMG product, free and ligand bound, and show the protein can decompose chitin, a common carbon polymer.
- Ruonan Wu
- , Clyde A. Smith
- & Janet K. Jansson
-
Article
| Open Access Genomic epidemiology of Delta SARS-CoV-2 during transition from elimination to suppression in Aotearoa New Zealand
Genomic epidemiology of Delta SARS-CoV-2 during transition from elimination to suppression in Aotearoa New Zealand
Aotearoa New Zealand pursued a COVID-19 elimination strategy until October 2021 when it moved to a suppression strategy. In this genomic surveillance study, the authors describe spread of the virus during the transition between these strategies, with evidence of substantial undetected community transmission.
- Lauren Jelley
- , Jordan Douglas
- & Jemma L. Geoghegan
-
Article
| Open Access Elimination of human rabies in Goa, India through an integrated One Health approach
Elimination of human rabies in Goa, India through an integrated One Health approach
Dog vaccination is an effective rabies prevention measure, but widespread vaccination campaigns are challenging in settings like India with large free-roaming dog populations. Here, the authors describe a One Health campaign in Goa state which led to a large reduction of cases in dogs and elimination in humans.
- A. D. Gibson
- , G. Yale
- & R. J. Mellanby
-
Article
| Open Access Cumulative SARS-CoV-2 mutations and corresponding changes in immunity in an immunocompromised patient indicate viral evolution within the host
Cumulative SARS-CoV-2 mutations and corresponding changes in immunity in an immunocompromised patient indicate viral evolution within the host
Variants of concerns arise from SARS-CoV-2 mutations poise as severe public health threats. Here the authors chronicle SARS-CoV-2 mutations onset and immune parameters in an immunocompromised patient with continuous virus-shedding, thereby hinting potential intra-host viral evolution and escape facilitated by ineffective T cell immunity.
- Sissy Therese Sonnleitner
- , Martina Prelog
- & Gernot Walder
-
Article
| Open Access Genomic assessment of quarantine measures to prevent SARS-CoV-2 importation and transmission
Genomic assessment of quarantine measures to prevent SARS-CoV-2 importation and transmission
Post-international travel quarantine has been widely implemented to mitigate SARS-CoV-2 transmission, but the impacts of such policies are unclear. Here, the authors used linked genomic and contact tracing data to assess the impacts of a 14-day quarantine on return to England in summer 2020.
- Dinesh Aggarwal
- , Andrew J. Page
- & Ewan M. Harrison
-
Article
| Open Access Genomic epidemiology of SARS-CoV-2 in a UK university identifies dynamics of transmission
Genomic epidemiology of SARS-CoV-2 in a UK university identifies dynamics of transmission
In this study, Aggarwal and colleagues perform prospective sequencing of SARS-CoV-2 isolates derived from asymptomatic student screening and symptomatic testing of students and staff at the University of Cambridge. They identify important factors that contributed to within university transmission and onward spread into the wider community.
- Dinesh Aggarwal
- , Ben Warne
- & Ian G. Goodfellow
-
Article
| Open Access SARS-CoV-2 genomes from Saudi Arabia implicate nucleocapsid mutations in host response and increased viral load
SARS-CoV-2 genomes from Saudi Arabia implicate nucleocapsid mutations in host response and increased viral load
In this study, the authors sequence 892 SARS-CoV-2 genomes from Saudi Arabia and describe population dynamics and importations into the country. They identify a nucleocapsid protein mutation associated with increased viral load and host interactions and characterise its role through biochemical analyses.
- Tobias Mourier
- , Muhammad Shuaib
- & Arnab Pain
-
Article
| Open Access Distinct patterns of within-host virus populations between two subgroups of human respiratory syncytial virus
Distinct patterns of within-host virus populations between two subgroups of human respiratory syncytial virus
Respiratory syncytial virus (RSV) is a common infection in children and older adults but little is known about within-host viral population diversity. Here, the authors perform deep sequencing and find that RSV subgroup B exhibited more diversity than subgroup A, with implications for development of therapeutics and vaccines.
- Gu-Lung Lin
- , Simon B. Drysdale
- & Andrew J. Pollard
-
Article
| Open Access Lying in wait: the resurgence of dengue virus after the Zika epidemic in Brazil
Lying in wait: the resurgence of dengue virus after the Zika epidemic in Brazil
Zika and dengue incidence in the Americas declined in 2017–2018, but dengue resurged in 2019 in Brazil. This study uses epidemiological, climatological and genomic data to show that the decline of dengue may be explained by protective immunity from pre-exposure to ZIKV and/or DENV in prior years.
- Anderson Fernandes Brito
- , Lais Ceschini Machado
- & Nathan D. Grubaugh
-
Article
| Open Access Varicella-zoster virus VLT-ORF63 fusion transcript induces broad viral gene expression during reactivation from neuronal latency
Varicella-zoster virus VLT-ORF63 fusion transcript induces broad viral gene expression during reactivation from neuronal latency
Varicella-zoster virus (VZV) establishes lifelong neuronal latency in humans. Here, Ouwendijk and Depledge et al. identify a fusion transcript, VLT-ORF63, which is expressed during lytic and latent infection, and demonstrate a role for the translated fusion protein in induction of lytic gene expression from latent VZV genomes.
- Werner J. D. Ouwendijk
- , Daniel P. Depledge
- & Tomohiko Sadaoka
-
Article
| Open Access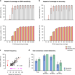 Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis
Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis
Nanopore sequencing (ONT) has been used in SARS-CoV-2 studies, however adoption of ONT for SARS-CoV-2 surveillance has been limited due to common concerns around sequencing accuracy. Here, the authors perform a comprehensive evaluation of ONT analytical performance on 157 matched SARS-CoV-2-positive patient specimens and synthetic RNA controls.
- Rowena A. Bull
- , Thiruni N. Adikari
- & Ira W. Deveson
-
Article
| Open Access No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2
No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2
SARS-CoV-2 has emerged recently and may still adapt to the human host. Here the authors show that none of the so far identified recurrent mutations in SARS-CoV-2 are significantly associated with increased viral transmission.
- Lucy van Dorp
- , Damien Richard
- & François Balloux
-
Article
| Open Access A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
Viruses have been difficult to position in the Tree of Life using phylogenetic methods. This study uses an ancient enzyme multi-subunit RNA polymerase (RNAP) to reveal a novel viral group, the Caudovirales, and to suggest an ancient origin of RNAP in this group.
- Alaina R. Weinheimer
- & Frank O. Aylward
-
Article
| Open Access Tracking the COVID-19 pandemic in Australia using genomics
Tracking the COVID-19 pandemic in Australia using genomics
Genome sequencing can be used to infer pathogen transmission dynamics and inform public health responses. Here, the authors sequence >1,200 SARS-CoV-2 samples from Victoria, Australia and find genomic support for the effectiveness of social restrictions in reducing transmission.
- Torsten Seemann
- , Courtney R. Lane
- & Benjamin P. Howden
-
Article
| Open Access Selective flexible packaging pathways of the segmented genome of influenza A virus
Selective flexible packaging pathways of the segmented genome of influenza A virus
The mechanism underlying packaging of the 8 segments of the influenza virus genome into virions is not well understood. Here, the authors use a multiplexed FISH assay to monitor the 8 segments in parallel in infected cells suggesting bundling routes during the packaging process.
- Ivan Haralampiev
- , Simon Prisner
- & Andreas Herrmann
-
Article
| Open Access Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
CrAss-like phages are bacterial viruses often found in the human gut. Here, Siranosian et al. analyze gut metagenomic data to evaluate the patterns of acquisition, transmission and strain diversity of these phages in mother-infant pairs and in patients undergoing fecal microbiota transplantation.
- Benjamin A. Siranosian
- , Fiona B. Tamburini
- & Ami S. Bhatt
-
Article
| Open Access Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
Incomplete influenza A virus genomes occur frequently but are readily complemented during localized viral spread
The genome of influenza is often incomplete in infected cells, but the implications for infection remain unclear. Here, Jacobs et al. show that an average of 3.6 particles is necessary for productive infection and that coinfection supports efficient complementation within a host but not upon transmission to a new host.
- Nathan T. Jacobs
- , Nina O. Onuoha
- & Anice C. Lowen
-
Article
| Open Access HIV-1 DNA sequence diversity and evolution during acute subtype C infection
HIV-1 DNA sequence diversity and evolution during acute subtype C infection
The dynamics of HIV-1 DNA sequences early after HIV-1 transmission remains poorly characterized. Here, the authors perform a longitudinal evaluation of HIV-1 DNA sequences in subtype C-infected individuals during acute infection, providing a landscape of the nature and evolution of the very early viral genome.
- Guinevere Q. Lee
- , Kavidha Reddy
- & Mathias Lichterfeld
-
Article
| Open Access Tracking HIV-1 recombination to resolve its contribution to HIV-1 evolution in natural infection
Tracking HIV-1 recombination to resolve its contribution to HIV-1 evolution in natural infection
Recombination contributes to HIV evolution in patients, but its identification can be difficult. Here, the authors develop a computational tool called RAPR to track recombination in patients, identify recombination hot spots, and show contribution of recombination to antibody escape.
- Hongshuo Song
- , Elena E. Giorgi
- & Feng Gao
-
Article
| Open Access Major bacterial lineages are essentially devoid of CRISPR-Cas viral defence systems
Major bacterial lineages are essentially devoid of CRISPR-Cas viral defence systems
It is thought that CRISPR-Cas systems, which confer acquired immunity to phage and archaeal viruses, are widespread among bacteria and archaea. Here, Burstein et al.show that entire lineages of uncultivated microorganisms are essentially devoid of CRISPR-Cas systems.
- David Burstein
- , Christine L. Sun
- & Jillian F. Banfield
-
Article
| Open Access The external domains of the HIV-1 envelope are a mutational cold spot
The external domains of the HIV-1 envelope are a mutational cold spot
Mutations allow RNA virus to adapt fast but also entail fitness costs. Geller et al. show that, in HIV-1, mutations occur three times less often in the most external domains of the envelope, and that this is due to changes in RNA sequence context and structure, which control viral and host-encoded mutational mechanisms.
- Ron Geller
- , Pilar Domingo-Calap
- & Rafael Sanjuán
-
Article
| Open Access Structure of Ljungan virus provides insight into genome packaging of this picornavirus
Structure of Ljungan virus provides insight into genome packaging of this picornavirus
The Ljungan virus is a picornavirus that lacks the internal coat protein VP4, and the packaging of its RNA genome is poorly understood. Here, the authors use cryo-electron microscopy to visualize this virus and suggest that it uses a different mechanism to other viruses for encapsidation of its genome.
- Ling Zhu
- , Xiangxi Wang
- & David I. Stuart
-
Article
| Open Access Convergent capture of retroviral superantigens by mammalian herpesviruses
Convergent capture of retroviral superantigens by mammalian herpesviruses
Horizontal gene transfer from retroviruses to mammals is rare between unrelated viruses. Here the authors show the convergent acquisition by herpesviruses of a virulence gene of ancient retroviruses, which occurred at least twice from different donor lineages, to distinct herpesviruses that infect mammals.
- Amr Aswad
- & Aris Katzourakis
-
Article
| Open Access Identification of mammalian-adapting mutations in the polymerase complex of an avian H5N1 influenza virus
Identification of mammalian-adapting mutations in the polymerase complex of an avian H5N1 influenza virus
Understanding the factors that enable some bird flu viruses to infect humans is important for the identification of circulating viruses with higher potential to infect us. Here, Taft et al.identify novel mutations in the polymerase of an avian H5N1 virus that help the virus to replicate in human cells and in mice
- Andrew S. Taft
- , Makoto Ozawa
- & Yoshihiro Kawaoka
-
Article |
 Epistatic interactions between neuraminidase mutations facilitated the emergence of the oseltamivir-resistant H1N1 influenza viruses
Epistatic interactions between neuraminidase mutations facilitated the emergence of the oseltamivir-resistant H1N1 influenza viruses
Understanding influenza evolution is challenging. Here, the authors determine the timing and order of critical amino acid changes that contributed to a world-wide predominance of oseltamivir-resistant H1N1 influenza viruses and show the role of epistasis in the emergence of novel influenza phenotypes.
- Susu Duan
- , Elena A. Govorkova
- & Richard J. Webby
-
Article |
 Specific recognition of the HIV-1 genomic RNA by the Gag precursor
Specific recognition of the HIV-1 genomic RNA by the Gag precursor
The Gag precursor protein recognizes the HIV-1 genomic RNA among hundreds of other viral and cellular RNAs during assembly of viral particles. Here, the authors identify the primary binding site of Gag on the HIV-1 RNA, and show that other RNA regions enhance or inhibit Gag binding.
- Ekram W. Abd El-Wahab
- , Redmond P. Smyth
- & Roland Marquet
-
Article |
 Gene copy number is differentially regulated in a multipartite virus
Gene copy number is differentially regulated in a multipartite virus
The potential evolutionary advantage associated with genome segmentation in multipartite viruses is not well established. Here Sicard et al. demonstrate that genome segmentation can allow a differential regulation of the copy number of each gene in a multipartite plant nanovirus during host infection.
- Anne Sicard
- , Michel Yvon
- & Stéphane Blanc
-
Article |
 Cooperation between different RNA virus genomes produces a new phenotype
Cooperation between different RNA virus genomes produces a new phenotype
RNA viruses are known to rapidly evolve new features through errors in replication and reshuffling of genomic segments. These authors report another strategy used by the measles virus to improve infectivity; the cooperation between wild-type and mutant fusion proteins in the same viral particle.
- Yuta Shirogane
- , Shumpei Watanabe
- & Yusuke Yanagi
-
Article |
 Adaptive mutations in NEP compensate for defective H5N1 RNA replication in cultured human cells
Adaptive mutations in NEP compensate for defective H5N1 RNA replication in cultured human cells
Adaptive mutations in the avian influenza virus permit replication in mammals but how these mutations enable this effect is unclear. In this study, mutations found in the nuclear export protein of human isolates of H5N1 are shown to enhance the replication of viral RNA in human cells in culture.
- Benjamin Mänz
- , Linda Brunotte
- & Martin Schwemmle
-
Article |
 Convergence and coevolution of Hepatitis B virus drug resistance
Convergence and coevolution of Hepatitis B virus drug resistance
Lamivudine treatment of hepatitis B is associated with drug-resistance mutations in the virus’ DNA polymerase. In this study, 11 patients with drug resistance are investigated and the primary mutation in the DNA polymerase shown to be essential but not sufficient for establishing drug resistance.
- Hong Thai
- , David S. Campo
- & Yury Khudyakov

 Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks