Featured
-
-
Article
| Open Access Characterisation of colistin resistance in Gram-negative microbiota of pregnant women and neonates in Nigeria
Characterisation of colistin resistance in Gram-negative microbiota of pregnant women and neonates in Nigeria
Here, the authors report the results of a BARNARDS sub-study identifying a 1% mobile colistin resistance gene (mcr) carriage rate in around 5000 rectal swabs from mothers and neonates across Nigeria, of which 90% were mcr-10 (mostly Enterobacter spp.) and 10% were mcr-1 and mcr9.
- E. A. R. Portal
- , K. Sands
- & O. B. Spiller
-
Article
| Open Access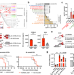 Dependency on host vitamin B12 has shaped Mycobacterium tuberculosis Complex evolution
Dependency on host vitamin B12 has shaped Mycobacterium tuberculosis Complex evolution
Campos-Pardos et al show that the virulence of Mycobacterium tuberculosis is dependent on sufficient uptake of exogenous vitamin B12 from host serum and this phenotype is not conserved in environmental, opportunistic and ancestral lineages.
- Elena Campos-Pardos
- , Santiago Uranga
- & Jesús Gonzalo-Asensio
-
Article
| Open Access Capsules and their traits shape phage susceptibility and plasmid conjugation efficiency
Capsules and their traits shape phage susceptibility and plasmid conjugation efficiency
Bacterial capsules provide protection against the environment, including host immune systems. Authors swap capsule loci in Klebsiella pneumoniae to reveal the role of these sugar coats against plasmid conjugation and phage infection, showing that the serotype is a key player in regulating conjugation rates, and phage susceptibility.
- Matthieu Haudiquet
- , Julie Le Bris
- & Olaya Rendueles
-
Article
| Open Access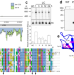 Protein NirP1 regulates nitrite reductase and nitrite excretion in cyanobacteria
Protein NirP1 regulates nitrite reductase and nitrite excretion in cyanobacteria
Some cyanobacteria excrete nitrite when the supply of inorganic carbon is limiting, but the underlying mechanisms are unclear. Here, Kraus et al. identify a conserved protein that interacts with nitrite reductase, thus regulating nitrogen metabolism and promoting nitrite excretion.
- Alexander Kraus
- , Philipp Spät
- & Wolfgang R. Hess
-
Article
| Open Access The plasmidome associated with Gram-negative bloodstream infections: A large-scale observational study using complete plasmid assemblies
The plasmidome associated with Gram-negative bloodstream infections: A large-scale observational study using complete plasmid assemblies
Plasmids carry antimicrobial resistance genes and contribute to the rapid dissemination of resistance. Here, the authors sequence 1,880 complete plasmids from 738 isolates from bloodstream infections, shedding light on the links between plasmid types, bacterial hosts and antimicrobial resistance.
- Samuel Lipworth
- , William Matlock
- & Nicole Stoesser
-
Article
| Open Access Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Duplicated antibiotic resistance genes reveal ongoing selection and horizontal gene transfer in bacteria
Mobile genetic elements can promote the duplication of antibiotic resistance genes which may in turn accelerate the evolution of resistance to new drugs. Here, the authors show that duplicated antibiotic resistance genes are enriched in bacterial isolates from environments associated with rampant antibiotic use.
- Rohan Maddamsetti
- , Yi Yao
- & Lingchong You
-
Article
| Open Access Transgenic expression of cif genes from Wolbachia strain wAlbB recapitulates cytoplasmic incompatibility in Aedes aegypti
Transgenic expression of cif genes from Wolbachia strain wAlbB recapitulates cytoplasmic incompatibility in Aedes aegypti
The Wolbachia cifA and cifB genes generate cytoplasmic incompatibility (CI) in insect hosts but the role of cifA is still debated. Here, the authors report the transgenic recapitulation of CI in the major arbovirus vector Aedes aegypti and provide evidence for cifA inhibiting cifB toxicity in the male germline.
- Cameron J. McNamara
- , Thomas H. Ant
- & Steven P. Sinkins
-
Article
| Open Access Quantitative measurement of antibiotic resistance in Mycobacterium tuberculosis reveals genetic determinants of resistance and susceptibility in a target gene approach
Quantitative measurement of antibiotic resistance in Mycobacterium tuberculosis reveals genetic determinants of resistance and susceptibility in a target gene approach
Molecular diagnostics for tuberculosis have focused on predicting drug susceptibilities in a binary manner (i.e., strains are either susceptible or resistant). Here, CRyPTIC Consortium researchers use whole genome sequencing and a quantitative assay to identify associations between genomic mutations and minimum inhibitory concentrations in over 15,000 Mycobacterium tuberculosis clinical isolates.
- Ivan Barilar
- , Simone Battaglia
- & Baoli Zhu
-
Article
| Open Access CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
CRISPR-Cas-based identification of a sialylated human milk oligosaccharides utilization cluster in the infant gut commensal Bacteroides dorei
Human milk oligosaccharides (HMOs) utilization by Bacteroides species remains poorly understood. Here, the authors describe a single specific gene cluster responsible for sialylated HMOs utilization in a B. dorei natural isolate and prove its functionality in vivo using CRISPR-Cas12a.
- Sivan Kijner
- , Dena Ennis
- & Moran Yassour
-
Article
| Open Access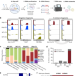 In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
Here the authors report a new approach to profile RNA-RNA interactions in live bacterial cells. The charted RNA interaction networks unveil a key mRNA regulatory hub targeted by twelve small RNAs, including a novel RNA involved in fatty acid metabolism.
- Fang Liu
- , Ziying Chen
- & Yanjie Chao
-
Article
| Open Access Quorum-sensing synthase mutations re-calibrate autoinducer concentrations in clinical isolates of Pseudomonas aeruginosa to enhance pathogenesis
Quorum-sensing synthase mutations re-calibrate autoinducer concentrations in clinical isolates of Pseudomonas aeruginosa to enhance pathogenesis
Simanek et al. discovered variants that arise in the protein responsible for synthesizing a molecule required for bacterial communication, which mediates the progression of virulence in the pathogen Pseudomonas aeruginosa.
- Kayla A. Simanek
- , Megan L. Schumacher
- & Jon E. Paczkowski
-
Article
| Open Access High-resolution temporal profiling of E. coli transcriptional response
High-resolution temporal profiling of E. coli transcriptional response
Understanding how cells dynamically adapt to their environment is important, but temporal information about cellular behaviour is often limited. Here, Miano et al. apply unsupervised machine learning to a dataset describing the activity of over 1,800 promoters in E. coli, measured every 10 minutes, defining three primary stages of promoter activation in response to heavy metal stress.
- Arianna Miano
- , Kevin Rychel
- & Jeff Hasty
-
Article
| Open Access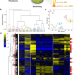 Direct comparison of spatial transcriptional heterogeneity across diverse Bacillus subtilis biofilm communities
Direct comparison of spatial transcriptional heterogeneity across diverse Bacillus subtilis biofilm communities
The bacterium Bacillus subtilis can form various types of surface-associated communities, such as colonies, pellicles and submerged biofilms. Here, Dergham et al. provide a direct comparison of spatial transcriptional heterogeneity across the three types of surface-associated communities, revealing mosaic expression patterns for genes involved in various pathways.
- Yasmine Dergham
- , Dominique Le Coq
- & Romain Briandet
-
Article
| Open Access c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
H-NS is a global regulatory protein that represses expression of many genes in bacteria. Here, Li et al. show that a second messenger, cyclic di-GMP, binds to H-NS and inhibits its binding to DNA, thus relieving H-NS-mediated transcriptional silencing.
- Shuyu Li
- , Qinmeng Liu
- & Lei Zhang
-
Article
| Open Access Clinically relevant antibiotic resistance genes are linked to a limited set of taxa within gut microbiome worldwide
Clinically relevant antibiotic resistance genes are linked to a limited set of taxa within gut microbiome worldwide
Antimicrobial resistance genes (ARGs) in commensal gut bacteria may act as a reservoir for acquisition by pathogens. Here, the authors assess the distribution and transfer potential of ARGs in gut microbiomes and find that clinically important ARGs are taxonomically restricted despite being associated with mobile plasmids
- Peter J. Diebold
- , Matthew W. Rhee
- & Ilana L. Brito
-
Article
| Open Access Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes
Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes
Bacteria of the phylum Planctomycetes display unique cell biology features but are relatively understudied. Here, the authors report a genome-wide analysis of essential gene content in a planctomycete, providing insights into the divergent molecular and cell biology of these organisms.
- Elena Rivas-Marin
- , David Moyano-Palazuelo
- & Damien P. Devos
-
Article
| Open Access Diagnostic and commensal Staphylococcus pseudintermedius genomes reveal niche adaptation through parallel selection of defense mechanisms
Diagnostic and commensal Staphylococcus pseudintermedius genomes reveal niche adaptation through parallel selection of defense mechanisms
Staphylococcus pseudintermedius has a wide host-range in domesticated and wild animals, yet it has also been isolated as an opportunistic pathogen in human wounds. In this work, the authors genotypically analyse S. pseudintermedius isolates from veterinary diagnostic laboratories and medical care centres, alongside household surfaces and inhabitants.
- Sanjam S. Sawhney
- , Rhiannon C. Vargas
- & Gautam Dantas
-
Article
| Open Access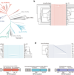 Catalytically inactive long prokaryotic Argonaute systems employ distinct effectors to confer immunity via abortive infection
Catalytically inactive long prokaryotic Argonaute systems employ distinct effectors to confer immunity via abortive infection
Here, Song et al. show that catalytically inactive long prokaryotic Argonaute proteins are equipped with distinct effectors that are activated upon recognition of invading genetic elements to trigger cell death and confer abortive infection immunity.
- Xinmi Song
- , Sheng Lei
- & Wenyuan Han
-
Article
| Open Access Identification and characterization of endo-α-, exo-α-, and exo-β-d-arabinofuranosidases degrading lipoarabinomannan and arabinogalactan of mycobacteria
Identification and characterization of endo-α-, exo-α-, and exo-β-d-arabinofuranosidases degrading lipoarabinomannan and arabinogalactan of mycobacteria
Lipoarabinomannan and arabinogalactan in the mycobacterial cell wall contain d-arabinan core. Here, the authors identify and characterize the molecular structures and mechanisms of four bacterial enzymes that synergistically degrade the alpha- and beta-linkages of d-arabinan.
- Michiko Shimokawa
- , Akihiro Ishiwata
- & Kiyotaka Fujita
-
Article
| Open Access Acquisition, co-option, and duplication of the rtx toxin system and the emergence of virulence in Kingella
Acquisition, co-option, and duplication of the rtx toxin system and the emergence of virulence in Kingella
The bacterial genus Kingella includes pathogenic species that secrete a toxin called RtxA, which is absent in commensal species. Here, Morreale et al. identify key steps in the evolutionary transition from commensal to pathogen, including horizontal gene transfer of the toxin-encoding genes, co-option of an existing secretion system, and gene duplication.
- Daniel P. Morreale
- , Eric A. Porsch
- & Paul J. Planet
-
Article
| Open Access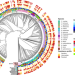 Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Here, Tisza, Dekker, and colleagues perform large scale analysis of genome methylation in the gut commensal and pathogen, Bacteroides fragilis group, revealing immense methyl motif diversity and evidence of widespread methyltransferase exchange among phages.
- Michael J. Tisza
- , Derek D. N. Smith
- & John P. Dekker
-
Article
| Open Access Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Transcription termination can tune bacterial gene expression in response to diverse signals. Here, the authors use several RNA-seq approaches to map RNA ends for the transcriptome of the spirochete Borrelia burgdorferi, providing insights into various modes of transcription termination and identifying potential RNA regulators in this pathogen.
- Emily Petroni
- , Caroline Esnault
- & Philip P. Adams
-
Article
| Open Access Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
The Cox’s Bazar area of Bangladesh has received a large number of Forcibly Displaced Myanmar Nationals. Cholera outbreaks have been detected in the area, and here, the authors perform genomic surveillance of cholera in the refugee and non-refugee population to infer the risk of epidemic spread.
- Alyce Taylor-Brown
- , Mokibul Hassan Afrad
- & Firdausi Qadri
-
Article
| Open Access Evolutionary and functional history of the Escherichia coli K1 capsule
Evolutionary and functional history of the Escherichia coli K1 capsule
Little is known about the distribution, evolution and functions of the K1 capsule at a population level, despite the important role in the pathogenesis of E. coli; authors explore this through the utilisation of over 5,000 clinical isolates in population genomics studies and statistical modelling.
- Sergio Arredondo-Alonso
- , George Blundell-Hunter
- & Alex J. McCarthy
-
Article
| Open Access Engineered hypermutation adapts cyanobacterial photosynthesis to combined high light and high temperature stress
Engineered hypermutation adapts cyanobacterial photosynthesis to combined high light and high temperature stress
Cyanobacteria mutants with improved tolerance to combined high light and high temperature (HLHT) are rarely reported. Here, the authors use a hypermutation system for adaptive laboratory evolution and identify a mutant with improved HLHT tolerance by enhancing expression of shikimate kinase.
- Huili Sun
- , Guodong Luan
- & Xuefeng Lu
-
Article
| Open Access Deep mutational scanning of essential bacterial proteins can guide antibiotic development
Deep mutational scanning of essential bacterial proteins can guide antibiotic development
Deep mutational scanning can be used to investigate protein function and stability. Here, Dewachter et al. use deep mutational scanning on three essential bacterial proteins to study the mutations’ effects in their original genomic context, providing insight into the proteins’ function and their potential as targets for new antibiotic development.
- Liselot Dewachter
- , Aaron N. Brooks
- & Jan Michiels
-
Article
| Open Access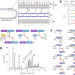 Genome-wide identification of genes required for alternative peptidoglycan cross-linking in Escherichia coli revealed unexpected impacts of β-lactams
Genome-wide identification of genes required for alternative peptidoglycan cross-linking in Escherichia coli revealed unexpected impacts of β-lactams
β-lactam-induced bacterial killing is complex and not fully resolved. Authors carry out a genome-wide analysis, through penicillin-binding protein replacement, to identify genes essential for drug efficacy.
- Henri Voedts
- , Sean P. Kennedy
- & Jean-Emmanuel Hugonnet
-
Article
| Open Access Bifurcation drives the evolution of assembly-line biosynthesis
Bifurcation drives the evolution of assembly-line biosynthesis
Reprogramming biosynthetic assembly-lines is a topic of interest for antibiotics. Here, the authors explore the evolutionary biosynthesis of anti-tubercular wollamides, show gene duplication and neo-functionalisation results in bifurcation allowing for testing of new structures with the ability to recover old structures by gene loss.
- Thomas J. Booth
- , Kenan A. J. Bozhüyük
- & Barrie Wilkinson
-
Article
| Open Access Short- and long-read metagenomics expand individualized structural variations in gut microbiomes
Short- and long-read metagenomics expand individualized structural variations in gut microbiomes
Here, Wang and colleagues combine short and long sequencing reads to characterize structural variations, prophage and CRISPR spacer elements in human gut microbiomes, and reveal functional differences at a finer level of bacterial strains.
- Liang Chen
- , Na Zhao
- & Jun Wang
-
Article
| Open Access The induction of natural competence adapts staphylococcal metabolism to infection
The induction of natural competence adapts staphylococcal metabolism to infection
Orthologs of natural competence genes are conserved in non-competent bacterial species, suggesting they have a role other than in transformation. Here, the authors show that competence induction in Staphylococcus aureus occurs in response to reactive oxygen species and host defenses that compromise bacterial respiration during infection, leading to increased DNA and glucose uptake and glycolytic flux.
- Mar Cordero
- , Julia García-Fernández
- & Daniel Lopez
-
Article
| Open Access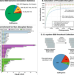 Generation of a Gluconobacter oxydans knockout collection for improved extraction of rare earth elements
Generation of a Gluconobacter oxydans knockout collection for improved extraction of rare earth elements
Bioleaching of rare earth elements using microorganisms offers an environmentally friendly alternative to thermochemical extraction. Here, Schmitz et al. generate a whole-genome knockout collection of mutants for one such microorganism, Gluconobacter oxydans, and identify genes affecting the production of acidic biolixiviant and thus bioleaching efficacy.
- Alexa M. Schmitz
- , Brooke Pian
- & Buz Barstow
-
Article
| Open Access Temporal evolution of master regulator Crp identifies pyrimidines as catabolite modulator factors
Temporal evolution of master regulator Crp identifies pyrimidines as catabolite modulator factors
Microbial evolution often involves transient phenotypes and sequential development of multiple mutations of unclear relevance. Here, the authors show that the evolution of non-growing E. coli cells can be driven by alterations in pyrimidine nucleoside levels associated with colony ageing and/or due to mutations in metabolic or regulatory genes.
- Ida Lauritsen
- , Pernille Ott Frendorf
- & Morten H. H. Nørholm
-
Article
| Open Access Discovery of an ene-reductase for initiating flavone and flavonol catabolism in gut bacteria
Discovery of an ene-reductase for initiating flavone and flavonol catabolism in gut bacteria
Flavonoids are abundant polyphenols in plants but it is not well understood how their metabolism is initiated by microbes in the human gut. Here, the authors identify and characterise an ene-reductase from the gut bacterium, Flavonifractor plautii ATCC 49531 that catalyses the hydrogenation of the C2–C3 double bond of flavones and flavonols and present its crystal structure.
- Gaohua Yang
- , Sen Hong
- & Yang Gu
-
Article
| Open Access Pandemic Vibrio cholerae shuts down site-specific recombination to retain an interbacterial defence mechanism
Pandemic Vibrio cholerae shuts down site-specific recombination to retain an interbacterial defence mechanism
Vibrio cholerae uses a type VI secretion system (T6SS) to kill neighbouring competitors. Here, Santoriello et al. show that a T6SS gene cluster (Aux3) exists as a mobile, prophage-like element in some environmental strains, and as a stable truncated form in pandemic isolates. They propose that Aux3 acquisition increased competitive fitness of pre-pandemic V. cholerae.
- Francis J. Santoriello
- , Lina Michel
- & Stefan Pukatzki
-
Article
| Open Access The seventh pandemic of cholera in Europe revisited by microbial genomics
The seventh pandemic of cholera in Europe revisited by microbial genomics
Since 1970, several cholera outbreaks caused by the “seventh pandemic” (7PET) lineage have been reported in Europe. Here, the authors demonstrate that the outbreaks were caused by repeated introductions of 7PET into Europe, rather than local environmental sources.
- Mihaela Oprea
- , Elisabeth Njamkepo
- & François-Xavier Weill
-
Article
| Open Access Widespread transfer of mobile antibiotic resistance genes within individual gut microbiomes revealed through bacterial Hi-C
Widespread transfer of mobile antibiotic resistance genes within individual gut microbiomes revealed through bacterial Hi-C
Linking antibiotic resistance (AR) in the gut microbiome with their bacterial hosts remains challenging. Here, the authors apply bacterial Hi-C to map mobile genetic elements in metagenomes, and illustrate that genes are present in more diverse taxa in neutropenic patients than healthy subjects.
- Alyssa G. Kent
- , Albert C. Vill
- & Ilana Lauren Brito
-
Article
| Open Access “Gene accordions” cause genotypic and phenotypic heterogeneity in clonal populations of Staphylococcus aureus
“Gene accordions” cause genotypic and phenotypic heterogeneity in clonal populations of Staphylococcus aureus
Gene tandem amplifications can drive bacterial evolution. Here, Belikova et al. identify copy number variations of lipoprotein-encoding genes in Staphylococcus aureus clinical isolates, and show that the loci expand and contract during bacterial growth in vitro and in mice, leading to changes in immunostimulatory capacity.
- Darya Belikova
- , Angelika Jochim
- & Simon Heilbronner
-
Article
| Open Access Sphingolipids produced by gut bacteria enter host metabolic pathways impacting ceramide levels
Sphingolipids produced by gut bacteria enter host metabolic pathways impacting ceramide levels
Ceramides are a type of sphingolipid (SL) that have been shown to play a role in several metabolic disorders. Here, the authors investigate the effect of SL-production by gut Bacteroides on host SL homeostasis and show that microbiome-derived SLs enter host circulation and alter ceramide production.
- Elizabeth L. Johnson
- , Stacey L. Heaver
- & Ruth E. Ley
-
Article
| Open Access Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Antimicrobial resistance (AMR) represents a global health threat. Here, the authors analyse the oral and gut resistomes from metagenomes of diverse populations and find that the oral resistome harbours higher abundance but lower diversity of antimicrobial resistance genes than the gut resistome.
- Victoria R. Carr
- , Elizabeth A. Witherden
- & David L. Moyes
-
Article
| Open Access Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR-Cas spacer acquisition platform
Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR-Cas spacer acquisition platform
Horizontal gene transfer (HGT) in complex bacterial communities has been mostly studied using metagenomic analyses. Here, the authors develop an E. coli CRISPR-Cas spacer acquisition platform that allows real-time recording of HGT events at nucleotide-resolution, identifying diverse DNA transfer events in human clinical fecal samples.
- Christian Munck
- , Ravi U. Sheth
- & Harris H. Wang
-
Article
| Open Access Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis
Here, the authors explore the potential of the 16S gene for discriminating bacterial taxa and show that full-length sequencing combined with appropriate clustering of intragenomic sequence variation can provide accurate representation of bacterial species in microbiome datasets.
- Jethro S. Johnson
- , Daniel J. Spakowicz
- & George M. Weinstock
-
Article
| Open Access A modular effector with a DNase domain and a marker for T6SS substrates
A modular effector with a DNase domain and a marker for T6SS substrates
Bacteria deliver toxic effectors via type VI secretion systems (T6SSs) to dominate competitors. Here, the authors identify a Vibrio antibacterial effector that contains a new DNase toxin domain and a domain of unknown function that can be used as a marker to identify new T6SS effectors.
- Biswanath Jana
- , Chaya M. Fridman
- & Dor Salomon
-
Article
| Open Access Essentiality of sterol synthesis genes in the planctomycete bacterium Gemmata obscuriglobus
Essentiality of sterol synthesis genes in the planctomycete bacterium Gemmata obscuriglobus
Sterols play essential functions in eukaryotic cell membranes, but are produced by few bacterial species. Here, the authors show that they are essential for growth of the planctomycete bacterium Gemmata obscuriglobus.
- Elena Rivas-Marin
- , Sean Stettner
- & Damien P. Devos
-
Article
| Open Access Deterministic processes structure bacterial genetic communities across an urban landscape
Deterministic processes structure bacterial genetic communities across an urban landscape
Disease transmission is particularly complex at the human-livestock-wildlife interface. Here the authors sample E. coli from wild birds near households in Nairobi and show that antimicrobial resistance gene diversity is correlated with human and lifestock density, while virulence gene diversity is correlated with avian species richness.
- J. M. Hassell
- , M. J. Ward
- & E. M. Fèvre
-
Article
| Open Access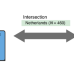 Joint sequencing of human and pathogen genomes reveals the genetics of pneumococcal meningitis
Joint sequencing of human and pathogen genomes reveals the genetics of pneumococcal meningitis
Streptococcus pneumoniae is a causative agent of meningitis and bacteremia. In a combined pathogen and host GWAS, Lees et al. find that host genetic variation is associated with both susceptibility and severity of pneumococcal meningitis, and specific bacterial genetic variation associated with susceptibility.
- John A. Lees
- , Bart Ferwerda
- & Diederik van de Beek
-
Article
| Open Access Stress-induced inactivation of the Staphylococcus aureus purine biosynthesis repressor leads to hypervirulence
Stress-induced inactivation of the Staphylococcus aureus purine biosynthesis repressor leads to hypervirulence
PurR acts as transcriptional repressor of purine biosynthesis genes in some bacterial species. Here, the authors show that purR mutations can arise in Staphylococcus aureus upon exposure to stress, leading to upregulation of fibronectin-binding proteins and increased virulence.
- Mariya I. Goncheva
- , Ronald S. Flannagan
- & David E. Heinrichs
-
Article
| Open Access Identification and characterization of a direct activator of a gene transfer agent
Identification and characterization of a direct activator of a gene transfer agent
Gene transfer agents (GTAs) are ‘domesticated’ bacteriophages that can transfer any genes between bacteria. Here, Paul Fogg identifies a protein that directly regulates transcription of GTA genes and whose expression is in turn controlled by a global cell-cycle regulator and a quorum-sensing regulator.
- Paul C. M. Fogg
-
Article
| Open Access Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Gain of function methods based on gene overexpression are not easily applied to high-throughput screening across different experimental conditions. Here, the authors present Dub-seq, which separates overexpression library characterization from functional screening and uses random DNA barcodes to increase the experimental throughput.
- Vivek K. Mutalik
- , Pavel S. Novichkov
- & Adam P. Arkin
-
Article
| Open Access Proteolysis of histidine kinase VgrS inhibits its autophosphorylation and promotes osmostress resistance in Xanthomonas campestris
Proteolysis of histidine kinase VgrS inhibits its autophosphorylation and promotes osmostress resistance in Xanthomonas campestris
Bacterial histidine kinases (HKs) play key roles in the response to stimuli and are regulated by reversible phosphorylation. Here, the authors show that the activity of a HK in the plant pathogen Xanthomonas campestris is modulated by irreversible, proteolytic modification in response to osmostress.
- Chao-Ying Deng
- , Huan Zhang
- & Wei Qian

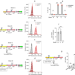 Transcription-driven DNA supercoiling counteracts H-NS-mediated gene silencing in bacterial chromatin
Transcription-driven DNA supercoiling counteracts H-NS-mediated gene silencing in bacterial chromatin