Featured
-
-
Article
| Open Access Co-option of a non-retroviral endogenous viral element in planthoppers
Co-option of a non-retroviral endogenous viral element in planthoppers
Non-retroviral endogenous viral elements are widely dispersed in eukaryotic genomes, but their functions remain largely unknown. Here, Huang et al show that one such element in planthoppers has been co-opted and contributes to insect fitness..
- Hai-Jian Huang
- , Yi-Yuan Li
- & Jun-Min Li
-
Article
| Open Access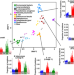 Mutational spectra are associated with bacterial niche
Mutational spectra are associated with bacterial niche
Mutagens and DNA repair defects generate context-specific mutational signatures in cancer cells. Here, Ruis et al. provide evidence of similar processes in bacteria, showing that mutational spectra may be associated with sites of bacterial replication when mutagen exposures differ, and can be used in these cases to infer transmission routes.
- Christopher Ruis
- , Aaron Weimann
- & Julian Parkhill
-
Article
| Open Access Diagnostic and commensal Staphylococcus pseudintermedius genomes reveal niche adaptation through parallel selection of defense mechanisms
Diagnostic and commensal Staphylococcus pseudintermedius genomes reveal niche adaptation through parallel selection of defense mechanisms
Staphylococcus pseudintermedius has a wide host-range in domesticated and wild animals, yet it has also been isolated as an opportunistic pathogen in human wounds. In this work, the authors genotypically analyse S. pseudintermedius isolates from veterinary diagnostic laboratories and medical care centres, alongside household surfaces and inhabitants.
- Sanjam S. Sawhney
- , Rhiannon C. Vargas
- & Gautam Dantas
-
Article
| Open Access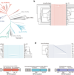 Catalytically inactive long prokaryotic Argonaute systems employ distinct effectors to confer immunity via abortive infection
Catalytically inactive long prokaryotic Argonaute systems employ distinct effectors to confer immunity via abortive infection
Here, Song et al. show that catalytically inactive long prokaryotic Argonaute proteins are equipped with distinct effectors that are activated upon recognition of invading genetic elements to trigger cell death and confer abortive infection immunity.
- Xinmi Song
- , Sheng Lei
- & Wenyuan Han
-
Article
| Open Access Inferring bacterial transmission dynamics using deep sequencing genomic surveillance data
Inferring bacterial transmission dynamics using deep sequencing genomic surveillance data
Studying rare genetic changes that arose as an infectious bacterium spread between lab mice, here the authors show that using the relative abundance of any changes rather than just whether they occurred can more precisely identify who likely infected who.
- Madikay Senghore
- , Hannah Read
- & Siouxsie Wiles
-
Article
| Open Access Origin of fungal hybrids with pathogenic potential from warm seawater environments
Origin of fungal hybrids with pathogenic potential from warm seawater environments
Most clinical isolates of the pathogenic yeast Candida orthopsilosis are hybrids of two parental lineages, only one of which has been identified. Here, del Olmo et al. show that C. orthopsilosis strains isolated from warm seawater are hybrids closely related to clinical isolates, and identify the missing parental lineage, thus providing a more complete view of the genomic evolution of this species.
- Valentina del Olmo
- , Verónica Mixão
- & Toni Gabaldón
-
Article
| Open Access A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
Invasive Salmonella Typhimurium bloodstream infection causes a significant public health burden in sub-Saharan Africa. Here, the authors analyse whole genome sequences of 1,302 S. Typhimurium isolates from Africa and describe its evolution, geographic spread, and antimicrobial resistance characteristics.
- Sandra Van Puyvelde
- , Tessa de Block
- & Octavie Lunguya
-
Article
| Open Access The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
Prophage induction is a fundamental process in the phage life cycle. Here the authors show that the final stage of prophage induction in Staphylococcus aureus is controlled by the ClpX protease, unveiling and unexpected role for ClpX in phage biology.
- Mohammed A. Thabet
- , José R. Penadés
- & Andreas F. Haag
-
Article
| Open Access Identification and characterization of endo-α-, exo-α-, and exo-β-d-arabinofuranosidases degrading lipoarabinomannan and arabinogalactan of mycobacteria
Identification and characterization of endo-α-, exo-α-, and exo-β-d-arabinofuranosidases degrading lipoarabinomannan and arabinogalactan of mycobacteria
Lipoarabinomannan and arabinogalactan in the mycobacterial cell wall contain d-arabinan core. Here, the authors identify and characterize the molecular structures and mechanisms of four bacterial enzymes that synergistically degrade the alpha- and beta-linkages of d-arabinan.
- Michiko Shimokawa
- , Akihiro Ishiwata
- & Kiyotaka Fujita
-
Article
| Open Access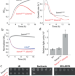 Mechanism of outer membrane destabilization by global reduction of protein content
Mechanism of outer membrane destabilization by global reduction of protein content
The outer membrane (OM) of Gram-negative bacteria is an asymmetric bilayer, with phospholipids in the inner leaflet. Here the authors show that a reduction in OM proteins and the subsequent mislocalization of phospholipids weaken the OM and alter growth rate and cell shape, emphasizing the role of OM proteins in OM stiffness and cell shape.
- Irina V. Mikheyeva
- , Jiawei Sun
- & Thomas J. Silhavy
-
Article
| Open Access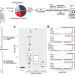 Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
The analysis of microbial genomes from human archaeological samples offers a snapshot of ancient pathogens. Here, Hodgins et al. analyze metagenomic datasets from 38 human archaeological samples and identify bacterial genomic sequences related to modern-day Clostridium tetani, encoding tetanus neurotoxins.
- Harold P. Hodgins
- , Pengsheng Chen
- & Andrew C. Doxey
-
Article
| Open Access Microdiversity of the vaginal microbiome is associated with preterm birth
Microdiversity of the vaginal microbiome is associated with preterm birth
Here, Liao et al. analyze the vaginal microbiome during pregnancy, and find a unique population genetic structure associated with preterm birth, suggesting that evolutionary processes acting on vaginal bacteria may play a role in prematurity.
- Jingqiu Liao
- , Liat Shenhav
- & Tal Korem
-
Article
| Open Access Genomic epidemiology offers high resolution estimates of serial intervals for COVID-19
Genomic epidemiology offers high resolution estimates of serial intervals for COVID-19
The serial interval (time between symptom onset in an infector and infectee) is usually estimated from contact tracing data, but this is not always available. Here, the authors develop a method for estimation of serial intervals using whole genome sequencing data and apply it data from clusters of SARS-CoV-2 in Victoria, Australia.
- Jessica E. Stockdale
- , Kurnia Susvitasari
- & Caroline Colijn
-
Article
| Open Access Genomic dissection of endemic carbapenem resistance reveals metallo-beta-lactamase dissemination through clonal, plasmid and integron transfer
Genomic dissection of endemic carbapenem resistance reveals metallo-beta-lactamase dissemination through clonal, plasmid and integron transfer
Resistance to carbapenems, a class of last-line antibiotics, is a global health threat. This study analysed a two-decade history of carbapenem resistance and identified complex, multi-level (bacterial strain, plasmid, gene) transmission dynamics.
- Nenad Macesic
- , Jane Hawkey
- & Anton Y. Peleg
-
Article
| Open Access Acquisition, co-option, and duplication of the rtx toxin system and the emergence of virulence in Kingella
Acquisition, co-option, and duplication of the rtx toxin system and the emergence of virulence in Kingella
The bacterial genus Kingella includes pathogenic species that secrete a toxin called RtxA, which is absent in commensal species. Here, Morreale et al. identify key steps in the evolutionary transition from commensal to pathogen, including horizontal gene transfer of the toxin-encoding genes, co-option of an existing secretion system, and gene duplication.
- Daniel P. Morreale
- , Eric A. Porsch
- & Paul J. Planet
-
Article
| Open Access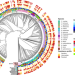 Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Here, Tisza, Dekker, and colleagues perform large scale analysis of genome methylation in the gut commensal and pathogen, Bacteroides fragilis group, revealing immense methyl motif diversity and evidence of widespread methyltransferase exchange among phages.
- Michael J. Tisza
- , Derek D. N. Smith
- & John P. Dekker
-
Article
| Open Access Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Transcription termination can tune bacterial gene expression in response to diverse signals. Here, the authors use several RNA-seq approaches to map RNA ends for the transcriptome of the spirochete Borrelia burgdorferi, providing insights into various modes of transcription termination and identifying potential RNA regulators in this pathogen.
- Emily Petroni
- , Caroline Esnault
- & Philip P. Adams
-
Article
| Open Access Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Genomic screening of 16 UK native bat species through conservationist networks uncovers coronaviruses with zoonotic potential
Certain bats species have previously been identified as ancestral sources of coronaviruses that infect humans but there is limited data on the genomic diversity or zoonotic potential of viruses infecting bats in the UK. Here, the authors use deep sequencing and in vitro assays to characterise coronaviruses recovered from 48 bat faecal samples.
- Cedric C. S. Tan
- , Jahcub Trew
- & Vincent Savolainen
-
Article
| Open Access Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
The Cox’s Bazar area of Bangladesh has received a large number of Forcibly Displaced Myanmar Nationals. Cholera outbreaks have been detected in the area, and here, the authors perform genomic surveillance of cholera in the refugee and non-refugee population to infer the risk of epidemic spread.
- Alyce Taylor-Brown
- , Mokibul Hassan Afrad
- & Firdausi Qadri
-
Article
| Open Access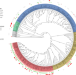 Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
Epistatic interactions between the high pathogenicity island and other iron uptake systems shape Escherichia coli extra-intestinal virulence
The virulence of extra-intestinal pathogenic Escherichia coli is associated with multiple different genes in different lineages. Here, Royer et al. show that the emergence of virulence is associated with acquisition of the siderophore-encoding high-pathogenicity island (HPI), and full virulence is associated with the additional presence of the aer or sit operons.
- Guilhem Royer
- , Olivier Clermont
- & Erick Denamur
-
Article
| Open Access Evolutionary and functional history of the Escherichia coli K1 capsule
Evolutionary and functional history of the Escherichia coli K1 capsule
Little is known about the distribution, evolution and functions of the K1 capsule at a population level, despite the important role in the pathogenesis of E. coli; authors explore this through the utilisation of over 5,000 clinical isolates in population genomics studies and statistical modelling.
- Sergio Arredondo-Alonso
- , George Blundell-Hunter
- & Alex J. McCarthy
-
Article
| Open Access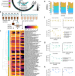 Culturing of a complex gut microbial community in mucin-hydrogel carriers reveals strain- and gene-associated spatial organization
Culturing of a complex gut microbial community in mucin-hydrogel carriers reveals strain- and gene-associated spatial organization
The organization of gut microbes in lumen and mucosa and the microbial genes regulating this organization remain poorly understood. Here, using in vitro cultures incorporating a complex gut microbial community in mucin-hydrogel carriers, the authors show greater richness and strain-specific spatial organization, enabling discovery of associated genes.
- Xiaofan Jin
- , Feiqiao B. Yu
- & Katherine S. Pollard
-
Article
| Open Access A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia
Authors carry out a longitudinal genomic analysis of Salmonella enterica serovar Concord isolates from various geographical locations, to reconstruct population diversity, evolution and antimicrobial resistance distribution.
- Wim L. Cuypers
- , Pieter Meysman
- & Sandra Van Puyvelde
-
Article
| Open Access The divisome but not the elongasome organizes capsule synthesis in Streptococcus pneumoniae
The divisome but not the elongasome organizes capsule synthesis in Streptococcus pneumoniae
The bacterial cell envelope consists of multiple layers, the synthesis of which is coordinated through unclear mechanisms. Here, Nakamoto et al. reveal a mechanism linking the synthesis of capsular polysaccharides and cell wall peptidoglycan in pneumococci.
- Rei Nakamoto
- , Sarp Bamyaci
- & Lok-To Sham
-
Article
| Open Access Virus diversity, wildlife-domestic animal circulation and potential zoonotic viruses of small mammals, pangolins and zoo animals
Virus diversity, wildlife-domestic animal circulation and potential zoonotic viruses of small mammals, pangolins and zoo animals
Monitoring the diversity of viruses infecting animals is important for assessing zoonotic risk. Here, the authors use metatranscriptomics to characterise the viromes of small mammals, pangolins, and zoo animals in China to identify potentially zoonotic viruses.
- Xinyuan Cui
- , Kewei Fan
- & Yongyi Shen
-
Article
| Open Access A host defense peptide mimetic, brilacidin, potentiates caspofungin antifungal activity against human pathogenic fungi
A host defense peptide mimetic, brilacidin, potentiates caspofungin antifungal activity against human pathogenic fungi
Current treatment of fungal infections is threatened by emerging antifungal drug resistance. In this work, the authors explore the synergistic activity of a host defense peptide mimetic, brilacidin, with caspofungin against a panel of fungal strains.
- Thaila Fernanda dos Reis
- , Patrícia Alves de Castro
- & Gustavo H. Goldman
-
Article
| Open Access Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
Associate toxin-antitoxin with CRISPR-Cas to kill multidrug-resistant pathogens
CRISPR-regulated toxin-antitoxin (CreTA), safeguards CRISPR-Cas immune systems. Here the authors characterize a bacterial CreTA and use this to generate a proof-of-concept antimicrobial strategy, ATTACK, which associates TA and CRISPR-Cas to kill multidrug resistant pathogens.
- Rui Wang
- , Xian Shu
- & Ming Li
-
Article
| Open Access Major proliferation of transposable elements shaped the genome of the soybean rust pathogen Phakopsora pachyrhizi
Major proliferation of transposable elements shaped the genome of the soybean rust pathogen Phakopsora pachyrhizi
Asian soybean rust caused by Phakopsora pachyrhizi is an important plant pathogen, but an accurate genome assembly for this fungus has been lacking. This study sequenced three independent P. pachyrhizi isolates and generated reference quality assemblies and genome annotations, representing a critical step for further in-depth studies of this pathogen and the development of new methods of control.
- Yogesh K. Gupta
- , Francismar C. Marcelino-Guimarães
- & H. Peter van Esse
-
Article
| Open Access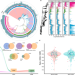 The genomic landscape of reference genomes of cultivated human gut bacteria
The genomic landscape of reference genomes of cultivated human gut bacteria
Here, the authors present an expanded version of the Cultivated Genome Reference (CGR), termed CGR2, a catalog that includes 3324 high-quality draft genomes based on gut bacterial isolates from Chinese individuals, and classifies 527 species from 8 phyla, including 179 previously unidentified species, and provides information of secondary metabolite biosynthetic gene clusters and gut phage-bacteria interactions.
- Xiaoqian Lin
- , Tongyuan Hu
- & Yuanqiang Zou
-
Article
| Open Access The evolution of antibiotic resistance is associated with collateral drug phenotypes in Mycobacterium tuberculosis
The evolution of antibiotic resistance is associated with collateral drug phenotypes in Mycobacterium tuberculosis
Here using drug susceptibility profiling, genomics and evolutionary studies the authors provide strategies to exploit collateral drug responses in Mycobacterium tuberculosis to prevent the emergence of drug resistance.
- Natalie J. E. Waller
- , Chen-Yi Cheung
- & Matthew B. McNeil
-
Article
| Open Access Engineered hypermutation adapts cyanobacterial photosynthesis to combined high light and high temperature stress
Engineered hypermutation adapts cyanobacterial photosynthesis to combined high light and high temperature stress
Cyanobacteria mutants with improved tolerance to combined high light and high temperature (HLHT) are rarely reported. Here, the authors use a hypermutation system for adaptive laboratory evolution and identify a mutant with improved HLHT tolerance by enhancing expression of shikimate kinase.
- Huili Sun
- , Guodong Luan
- & Xuefeng Lu
-
Article
| Open Access Membrane-localized expression, production and assembly of Vibrio parahaemolyticus T3SS2 provides evidence for transertion
Membrane-localized expression, production and assembly of Vibrio parahaemolyticus T3SS2 provides evidence for transertion
It has been proposed that bacterial membrane proteins may be produced via ‘transertion’, or concurrent transcription, translation and membrane insertion from membrane-associated genes. Here, Kaval et al. provide evidence supporting that Vibrio parahaemolyticus uses transertion to assemble a transmembrane complex (type III secretion system) used to inject virulence factors into host cells.
- Karan Gautam Kaval
- , Suneeta Chimalapati
- & Kim Orth
-
Article
| Open Access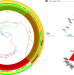 Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Genomic attributes of Vibrio cholerae O1 responsible for 2022 massive cholera outbreak in Bangladesh
Vibrio cholerae has undergone continuous evolution, and differing strains have caused numerous outbreaks. Here, the authors present a genomic study of Vibrio cholerae O1 responsible for a 2022 outbreak in Dhaka.
- Md Mamun Monir
- , Mohammad Tarequl Islam
- & Munirul Alam
-
Article
| Open Access High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection
High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection
Here, by high-resolution SARS-CoV-2 sequencing, genomic and transcriptomic analyses from tissue samples, Normandin et al. investigate viral dynamics in fatal cases of COVID-19, revealing persistent infection in distinct anatomical sites, including the heart and testis.
- Erica Normandin
- , Melissa Rudy
- & Isaac H. Solomon
-
Article
| Open Access Host-microbe co-metabolism via MCAD generates circulating metabolites including hippuric acid
Host-microbe co-metabolism via MCAD generates circulating metabolites including hippuric acid
Here, using a mouse model, the authors report a previously undescribed role for medium-chain acyl-CoA dehydrogenase in host metabolism of gut microbiota metabolites, and show that circulating compounds, including the abundant organic acid hippurate, depend on host-microbe co-metabolism of phenylalanine by Clostridium sporogenes.
- Kali M. Pruss
- , Haoqing Chen
- & Dylan Dodd
-
Article
| Open Access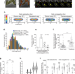 Real-time visualisation of the intracellular dynamics of conjugative plasmid transfer
Real-time visualisation of the intracellular dynamics of conjugative plasmid transfer
Conjugation is a contact-dependent mechanism for the transfer of plasmid DNA between bacterial cells. Here, Couturier et al. use live-cell microscopy to visualise the intracellular dynamics of conjugation in real time, revealing a molecular strategy that allows the sequential production of factors involved in establishing, maintaining and disseminating the plasmid.
- Agathe Couturier
- , Chloé Virolle
- & Christian Lesterlin
-
Article
| Open Access Rapid transmission and tight bottlenecks constrain the evolution of highly transmissible SARS-CoV-2 variants
Rapid transmission and tight bottlenecks constrain the evolution of highly transmissible SARS-CoV-2 variants
Here, by sequencing viruses from individuals in multiple households, Bendall et al. find that SARS-CoV-2 transmission bottleneck does not vary between individuals infected with pre-variant lineages and those infected with highly transmissible Alpha, Delta, or Omicron variants, suggesting these tight bottlenecks will limit the spread of new mutations.
- Emily E. Bendall
- , Amy P. Callear
- & Adam S. Lauring
-
Article
| Open Access Harnessing gut microbes for glycan detection and quantification
Harnessing gut microbes for glycan detection and quantification
Detecting distinct glycans within heterogeneous mixtures is hindered by glycan structural complexity and diversity. Here the authors exploit the ability of gut microbes to sense different glycan structures in order to develop quantitative glycan biosensors by coupling bacterial detection machinery to an optimised luciferase reporter.
- Jennifer L. Modesto
- , Victoria H. Pearce
- & Guy E. Townsend II
-
Article
| Open Access Deep mutational scanning of essential bacterial proteins can guide antibiotic development
Deep mutational scanning of essential bacterial proteins can guide antibiotic development
Deep mutational scanning can be used to investigate protein function and stability. Here, Dewachter et al. use deep mutational scanning on three essential bacterial proteins to study the mutations’ effects in their original genomic context, providing insight into the proteins’ function and their potential as targets for new antibiotic development.
- Liselot Dewachter
- , Aaron N. Brooks
- & Jan Michiels
-
Article
| Open Access Resolving colistin resistance and heteroresistance in Enterobacter species
Resolving colistin resistance and heteroresistance in Enterobacter species
Taxonomical complexity has muddled the classification of clinically relevant Enterobacter species. Authors carry out a genome-based study on clinical isolates to investigate colistin resistance and heteroresistance in Enterobacter.
- Swapnil Prakash Doijad
- , Nicolas Gisch
- & Trinad Chakraborty
-
Article
| Open Access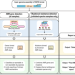 An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance
An ISO-certified genomics workflow for identification and surveillance of antimicrobial resistance
The implementation of genomics for identification and surveillance of antimicrobial resistance (AMR) in clinical laboratories remains challenging. Here, Sherry et al. present a bioinformatics platform for detection of AMR determinants from whole-genome sequencing data, suitable for clinical and public-health microbiology reporting.
- Norelle L. Sherry
- , Kristy A. Horan
- & Torsten Seemann
-
Article
| Open Access Multistep diversification in spatiotemporal bacterial-phage coevolution
Multistep diversification in spatiotemporal bacterial-phage coevolution
Bacteria and their viruses coexist and coevolve in nature, but maintaining them together in the lab is challenging. Here, a spatially structured environment allowed prolonged coevolution, with bacteria and phage diversifying into multiple ecotypes, uncovering gene mechanisms affecting phage-bacteria interactions.
- Einat Shaer Tamar
- & Roy Kishony
-
Article
| Open Access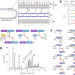 Genome-wide identification of genes required for alternative peptidoglycan cross-linking in Escherichia coli revealed unexpected impacts of β-lactams
Genome-wide identification of genes required for alternative peptidoglycan cross-linking in Escherichia coli revealed unexpected impacts of β-lactams
β-lactam-induced bacterial killing is complex and not fully resolved. Authors carry out a genome-wide analysis, through penicillin-binding protein replacement, to identify genes essential for drug efficacy.
- Henri Voedts
- , Sean P. Kennedy
- & Jean-Emmanuel Hugonnet
-
Article
| Open Access Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Extended-spectrum cephalosporin resistance genes in Escherichia coli have spread worldwide. Here, the authors dissect the emergence and distribution of these genes over time, and across geographic location and host species, to better understand their dynamics and mechanisms of transmission.
- Roxana Zamudio
- , Patrick Boerlin
- & Alison E. Mather
-
Article
| Open Access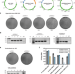 Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Eggerthella lenta is a prominent human gut bacterium implicated in several physiological processes, but its study has remained limited. Here, by developing a genetic toolbox for E. lenta, the authors provide insights into how the bacterium regulates drug and dietary compound metabolism.
- Xueyang Dong
- , Ben G. H. Guthrie
- & Emily P. Balskus
-
Article
| Open Access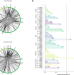 An RNA sponge controls quorum sensing dynamics and biofilm formation in Vibrio cholerae
An RNA sponge controls quorum sensing dynamics and biofilm formation in Vibrio cholerae
Small regulatory RNAs (sRNAs) often act in concert with the RNA-chaperone Hfq to regulate the expression of multiple target transcripts in bacteria. Here, the authors identify Hfq-interacting sRNAs and their targets in the pathogen Vibrio cholerae, including an RNA sponge that binds and inactivates four sRNAs that modulate the quorum sensing pathway.
- Michaela Huber
- , Anne Lippegaus
- & Kai Papenfort
-
Article
| Open Access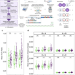 Characterisation of SARS-CoV-2 genomic variation in response to molnupiravir treatment in the AGILE Phase IIa clinical trial
Characterisation of SARS-CoV-2 genomic variation in response to molnupiravir treatment in the AGILE Phase IIa clinical trial
Molnupiravir is an antiviral that forces lethal error catastrophe in SARS-CoV-2 RNAs. Here, the authors confirm the mechanism of action of molnupiravir in humans using samples obtained from the UK’s AGILE phase IIa clinical trial investigating the antiviral efficacy of the drug against SARS-CoV-2. No treatment-associated SARS-CoV-2 mutations were identified.
- I’ah Donovan-Banfield
- , Rebekah Penrice-Randal
- & Thomas Fletcher
-
Article
| Open Access Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Ribosomal RNA genes are abundant in eukaryotic genomes and code for the universal and essential RNA components of the ribosome. This study uncovers high sequence diversity of the genes within a single species and discusses the contribution of selection in the evolution of ribosomal RNA.
- Daniel Sultanov
- & Andreas Hochwagen
-
Article
| Open Access A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
H37Rv is the most widely used Mycobacterium tuberculosis strain, and its genome is the reference sequence for this pathogen. Here, Chitale et al. present a bioinformatic pipeline for accurate assembly of bacterial genome sequences, and use it to provide important updates to the M. tuberculosis reference genome.
- Poonam Chitale
- , Alexander D. Lemenze
- & David Alland

 Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes
Essential gene complement of Planctopirus limnophila from the bacterial phylum Planctomycetes