Article
|
Open Access
Featured
-
-
Article
| Open Access Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Some polygenic risk score (PRS) methods for predicting genetic risk for common diseases require an external individual-level dataset for parameter tuning, posing privacy-related concerns. Here, the authors present an empirical Bayes method that tunes PRS models using only summary statistics from the training data.
- Wei Jiang
- , Ling Chen
- & Hongyu Zhao
-
Article
| Open Access Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Accurate and high-throughput sequencing methods for proteins are lacking. Here the authors report Spectralis which improves de novo peptide sequencing using a convolutional layer that connects peaks in spectra spaced by amino acid masses, fragment ion series classification and a peptide-spectrum match confidence score.
- Daniela Klaproth-Andrade
- , Johannes Hingerl
- & Julien Gagneur
-
Article
| Open Access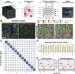 MAPS: pathologist-level cell type annotation from tissue images through machine learning
MAPS: pathologist-level cell type annotation from tissue images through machine learning
Current cell annotation methods using high-plex spatial proteomics data are resource intensive and demand iterative expert input. Here, the authors present MAPS (Machine learning for Analysis of Proteomics in Spatial biology), an approach that facilitates rapid and precise cell type identification with human-level accuracy from spatial proteomics data.
- Muhammad Shaban
- , Yunhao Bai
- & Faisal Mahmood
-
Article
| Open Access Integrative genotyping of cancer and immune phenotypes by long-read sequencing
Integrative genotyping of cancer and immune phenotypes by long-read sequencing
Single-cell transcriptomics excel in cell subset classification and can be augmented by suitable genotype information. Here the authors devise a long-read sequencing workflow, termed nanoranger, for detection of molecular barcodes from single-cell cDNA and apply this to clonal tracking of acute myeloid leukemia and identification of complex immune phenotypes.
- Livius Penter
- , Mehdi Borji
- & Catherine J. Wu
-
Article
| Open Access Integrated multi-omics analyses identify anti-viral host factors and pathways controlling SARS-CoV-2 infection
Integrated multi-omics analyses identify anti-viral host factors and pathways controlling SARS-CoV-2 infection
Amnesic screening methods are useful to discover host factors that are important for SARS-CoV2 infection. Here the authors use a CRISPR screen to identify three anti-viral factors, which are associated with the coagulation system, and two pro-viral candidates and then use individual genetic deletion experiments to characterise their effect.
- Jiakai Hou
- , Yanjun Wei
- & Weiyi Peng
-
Article
| Open Access ACIDES: on-line monitoring of forward genetic screens for protein engineering
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Screening mutated proteins is a versatile strategy in protein research, producing massive datasets when combined with NGS. Here, authors present ACIDES to estimate mutated protein fitness and aid protein engineering pipelines in a range of applications, including gene therapy.
- Takahiro Nemoto
- , Tommaso Ocari
- & Ulisse Ferrari
-
Article
| Open Access Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Existing feature visualisation methods are not well-suited for regression tasks. Here, authors introduce a method to learn the manifold topology related to deep neural network output and target labels and provide insightful visualisations of the high-dimensional features while preserving the local geometry.
- Md Tauhidul Islam
- , Zixia Zhou
- & Lei Xing
-
Article
| Open Access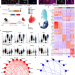 Spatiotemporal signaling underlies progressive vascular rarefaction in myocardial infarction
Spatiotemporal signaling underlies progressive vascular rarefaction in myocardial infarction
Enhancing vascularization to improve cardiac disease outcomes is a therapeutic approach with limited success. Here, the authors show that cardiac repair is governed by spatiotemporally regulated programs and underline the signaling mechanisms driving vascular deterioration.
- Lin Wei Tung
- , Elena Groppa
- & Fabio Rossi
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access Deciphering driver regulators of cell fate decisions from single-cell transcriptomics data with CEFCON
Deciphering driver regulators of cell fate decisions from single-cell transcriptomics data with CEFCON
Deciphering the roles of gene regulation in cell fate decisions is crucial. Here, authors present CEFCON, a network-based framework that reveals cell-lineage-specific gene regulatory networks and identifies driver regulators controlling cell fate decisions from single-cell transcriptomics data.
- Peizhuo Wang
- , Xiao Wen
- & Jianyang Zeng
-
Article
| Open Access Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Proteins contain modular structural and functional units called domains. Here, the authors have developed Merizo, a deep learning method for domain segmentation applicable to experimental structures as well as those generated by AlphaFold2.
- Andy M. Lau
- , Shaun M. Kandathil
- & David T. Jones
-
Article
| Open Access Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Clustering-based analysis has limited power in highly dynamic single-cell data, which is a common situation in tumour samples. Here, authors introduce GSDensity, enabling pathway-centric analysis for the direct integration of data with their domain knowledge.
- Qingnan Liang
- , Yuefan Huang
- & Ken Chen
-
Article
| Open Access High-throughput deconvolution of 3D organoid dynamics at cellular resolution for cancer pharmacology with Cellos
High-throughput deconvolution of 3D organoid dynamics at cellular resolution for cancer pharmacology with Cellos
Computational methods to analyse 3D organoids in high-throughput and with high cellular resolution remain scarce. Here, the authors propose Cellos, a high-throughput pipeline for 3D organoid segmentation using classical algorithms and a trained convolutional neural network.
- Patience Mukashyaka
- , Pooja Kumar
- & Jeffrey H. Chuang
-
Article
| Open Access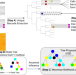 LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
Inferring lineage trees while incorporating gene expressions and lineage barcodes is a challenging task. Here, authors present LinRace, which infers improved cell lineage trees and ancestral cell states using the proposed asymmetric division model.
- Xinhai Pan
- , Hechen Li
- & Xiuwei Zhang
-
Article
| Open Access Local energetic frustration conservation in protein families and superfamilies
Local energetic frustration conservation in protein families and superfamilies
Energetic local frustration in proteins may have been positively selected by evolution when related to function such as ligand binding, allostery and other. Here the authors present a methodology to analyze local frustration patterns within protein families and superfamilies.
- Maria I. Freiberger
- , Victoria Ruiz-Serra
- & Alfonso Valencia
-
Article
| Open Access Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Co-fractionation mass spectrometry (CF-MS) is a powerful technique for mapping protein interactions under physiological conditions. Here, the authors uniformly re-process 411 CF-MS experiments and carry out meta-analyses of protein abundance, protein-protein interactions, and phosphorylation sites in the resulting resource.
- Michael A. Skinnider
- , Mopelola O. Akinlaja
- & Leonard J. Foster
-
Article
| Open Access Immune-response 3′UTR alternative polyadenylation quantitative trait loci contribute to variation in human complex traits and diseases
Immune-response 3′UTR alternative polyadenylation quantitative trait loci contribute to variation in human complex traits and diseases
Alternative polyadenylation (APA) has a key role in the post-transcriptional regulation of most human genes but is understudied in cells of the immune system. Here, the authors construct an atlas of cell type-specific APA events in various immune cell-types and stimulation conditions, providing evidence of widespread stimulation-responsiveness and association with immune-related traits.
- Lei Li
- , Xuelian Ma
- & Wei Li
-
Article
| Open Access A computational toolbox for the assembly yield of complex and heterogeneous structures
A computational toolbox for the assembly yield of complex and heterogeneous structures
Predicting the effective assembly of a set of proteins into a desired structure has traditionally been a challenging task. Here, authors demonstrate that advancements in automatic differentiation make it possible to address this problem using classical statistical mechanics.
- Agnese I. Curatolo
- , Ofer Kimchi
- & Michael P. Brenner
-
Article
| Open Access Deep learning-based phenotyping reclassifies combined hepatocellular-cholangiocarcinoma
Deep learning-based phenotyping reclassifies combined hepatocellular-cholangiocarcinoma
Combined hepatocellular-cholangiocarcinomas (cHCC-CCA) are challenging to diagnose, as they exhibit features of hepatocellular carcinoma (HCC) and intrahepatic cholangiocarcinoma (ICCA). Here, the authors use deep learning to re-classify cHCC-CCA tumours into HCC or ICCA based on histopathology images.
- Julien Calderaro
- , Narmin Ghaffari Laleh
- & Jakob Nikolas Kather
-
Article
| Open Access Directing polymorph specific calcium carbonate formation with de novo protein templates
Directing polymorph specific calcium carbonate formation with de novo protein templates
Most proteins mediating biomineralization in nature are not well structured, and the structures of the relevant protein-mineral interfaces regulating mineralization are elusive. Here, the authors computationally design proteins that modulate calcium carbonate mineralization to generate hybrid materials and elucidate the roles of designed proteins in controlling mineralization.
- Fatima A. Davila-Hernandez
- , Biao Jin
- & David Baker
-
Article
| Open Access Prediction and stratification of longitudinal risk for chronic obstructive pulmonary disease across smoking behaviors
Prediction and stratification of longitudinal risk for chronic obstructive pulmonary disease across smoking behaviors
Many people who never smoke develop COPD. Here, the authors derive and validate the Socioeconomic and Environmental Risk Score (SERS) which captures cumulative exposure risks beyond tobacco smoking to predict and stratify risk of COPD.
- Yixuan He
- , David C. Qian
- & Chirag J. Patel
-
Article
| Open Access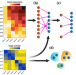 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Current methods to predict structures of proteins cannot handle large assemblies with complex symmetries. Here, the authors demonstrate that structures of proteins with cubic symmetries can be accurately predicted with a method combining AlphaFold with symmetrical assembly simulations.
- Mads Jeppesen
- & Ingemar André
-
Article
| Open Access GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
Reconstructing transcriptome-wide spatially-resolved gene expressions requires modelling nonlinear patterns and spatial structures in RNA profiling data. Here, authors introduce a graph-guided neural hierarchical tensor decomposition model that incorporates spatial and functional relations for the task.
- Tianci Song
- , Charles Broadbent
- & Rui Kuang
-
Article
| Open Access Structural variants involved in high-altitude adaptation detected using single-molecule long-read sequencing
Structural variants involved in high-altitude adaptation detected using single-molecule long-read sequencing
Here, the authors use single-molecule long-read sequencing to decipher the role of structural variations in high-altitude adaptation, finding evidence that an intergenic deletion down-regulates EPAS1 by disrupting a super-enhancer.
- Jinlong Shi
- , Zhilong Jia
- & Kunlun He
-
Article
| Open Access Genome-resolved metatranscriptomics reveals conserved root colonization determinants in a synthetic microbiota
Genome-resolved metatranscriptomics reveals conserved root colonization determinants in a synthetic microbiota
The identification of processes activated by specific microbes during microbiota colonization of plant roots is hampered by technical issues in metatranscriptomics. Here, Vannier et al. colonized germ-free plants with a defined root microbiota comprising over 100 microbial isolates, and addressed those issues in various ways to identify strain-specific processes as well as common gene sets activated by microbes during root colonization.
- Nathan Vannier
- , Fantin Mesny
- & Stéphane Hacquard
-
Comment
| Open Access Is Protein BLAST a thing of the past?
Is Protein BLAST a thing of the past?
Will protein structure search tools like AlphaFold replace protein sequence search with BLAST? We discuss the promises, using structure search for remote homology detection, and why protein BLAST, as the leading sequence search tool, should strive to incorporate structural information
- Ali Al-Fatlawi
- , Martin Menzel
- & Michael Schroeder
-
Article
| Open Access DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
Retention time (RT) alignment is a crucial step in large cohort proteomics and metabolomics studies. Here, the authors introduce DeepRTAlign, a deep learning tool for RT alignment that shows high identification sensitivity and quantitative accuracy.
- Yi Liu
- , Yun Yang
- & Cheng Chang
-
Article
| Open Access UniKP: a unified framework for the prediction of enzyme kinetic parameters
UniKP: a unified framework for the prediction of enzyme kinetic parameters
Prediction of enzyme kinetic parameters is essential for designing and optimising enzymes for various biotechnological and industrial applications. Here, authors presented a prediction framework (UniKP), which improves the accuracy of predictions for three enzyme kinetic parameters.
- Han Yu
- , Huaxiang Deng
- & Xiaozhou Luo
-
Article
| Open Access An archetype and scaling of developmental tissue dynamics across species
An archetype and scaling of developmental tissue dynamics across species
Limb tissue dynamics until basic skeletal pattern establishment exhibit a high degree of conservation between chick and frog after proper rescaling of spacetime, suggesting the presence of a species-independent archetype of morphogenetic dynamics.
- Yoshihiro Morishita
- , Sang-Woo Lee
- & Aiko Kawasumi-Kita
-
Article
| Open Access High-throughput target trial emulation for Alzheimer’s disease drug repurposing with real-world data
High-throughput target trial emulation for Alzheimer’s disease drug repurposing with real-world data
Target trial emulation (TTE) simulates randomized controlled trials using real world data (RWD). Here, authors show the effectiveness of different TTE strategies to identify drug candidates that could be potentially repurposed to Alzheimer’s disease using two large scale RWD warehouses.
- Chengxi Zang
- , Hao Zhang
- & Fei Wang
-
Article
| Open Access Deep learning of cell spatial organizations identifies clinically relevant insights in tissue images
Deep learning of cell spatial organizations identifies clinically relevant insights in tissue images
Cell spatial organization in tissue provides essential insights into diseases. Here, the authors show Ceograph, a graph convolutional network, for the analysis of pathology images to predict patient outcomes, highlighting cellular markers to guide personalized treatments and enhance biological understanding.
- Shidan Wang
- , Ruichen Rong
- & Guanghua Xiao
-
Article
| Open Access vcfdist: accurately benchmarking phased small variant calls in human genomes
vcfdist: accurately benchmarking phased small variant calls in human genomes
Accurately benchmarking small variant calling accuracy is critical for the continued improvement of human genome sequencing. Here, the authors show that current approaches are biased towards certain variant representations and develop a new approach to ensure consistent and accurate benchmarking, regardless of the original variant representations.
- Tim Dunn
- & Satish Narayanasamy
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Delineating the early dissemination mechanisms of acral melanoma by integrating single-cell and spatial transcriptomic analyses
Acral melanoma (AM) is a rare melanoma subtype with unique features, where lymph node metastasis is closely associated with clinical outcomes. Here, the authors use single-cell and spatial transcriptomics to analyse early dissemination, tumour microenvironment, and heterogeneity in AM, and infer metabolic shifts with therapeutic implications.
- Chuanyuan Wei
- , Wei Sun
- & Jianying Gu
-
Article
| Open Access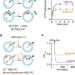 Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Mitochondria depend on phospholipids supplied by the endoplasmic reticulum. Here, using biochemical assays and molecular dynamics simulations, authors identify VDAC as a scramblase-type lipid transporter that catalyze lipid entry.
- Helene Jahn
- , Ladislav Bartoš
- & Anant K. Menon
-
Article
| Open Access The PENGUIN approach to reconstruct protein interactions at enhancer-promoter regions and its application to prostate cancer
The PENGUIN approach to reconstruct protein interactions at enhancer-promoter regions and its application to prostate cancer
The authors reconstruct high fidelity networks of protein-protein interactions between promoters and enhancers in prostate cancer and demonstrate the potential of such an analytical framework to obtain actionable insights into the disease and potential therapeutic targets.
- Alexandros Armaos
- , François Serra
- & Gian Gaetano Tartaglia
-
Article
| Open Access Design and structural validation of peptide–drug conjugate ligands of the kappa-opioid receptor
Design and structural validation of peptide–drug conjugate ligands of the kappa-opioid receptor
Despite advances in GPCR structures and peptide design, creating high-affinity ligands remains a challenge. Here the authors develop a computational method, successfully identifying peptide-based molecules for KOR: their platform shows promise for streamlined GPCR ligand discovery.
- Edin Muratspahić
- , Kristine Deibler
- & Christian W. Gruber
-
Article
| Open Access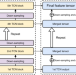 Accurate de novo peptide sequencing using fully convolutional neural networks
Accurate de novo peptide sequencing using fully convolutional neural networks
De novo peptide sequencing allows the identification of peptides without requiring target databases. Here, the authors present PepNet, a convolutional neural network model for accurate de novo peptide sequencing that is capable of analysing large-scale proteomics data.
- Kaiyuan Liu
- , Yuzhen Ye
- & Haixu Tang
-
Article
| Open Access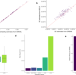 Accurate and efficient estimation of local heritability using summary statistics and the linkage disequilibrium matrix
Accurate and efficient estimation of local heritability using summary statistics and the linkage disequilibrium matrix
The authors propose “HEELS”, a new method for precise local heritability estimation. It significantly reduces the variances of summary-statistics-based heritability estimators, offering an REML-like estimator without requiring individual-level data.
- Hui Li
- , Rahul Mazumder
- & Xihong Lin
-
Article
| Open Access Programmable de novo designed coiled coil-mediated phase separation in mammalian cells
Programmable de novo designed coiled coil-mediated phase separation in mammalian cells
Membraneless liquid compartments based on phase-separating biopolymers have been observed in diverse cell types and attributed to weak multivalent interactions predominantly based on intrinsically disordered domains. Here the authors design protein liquid condensates from tunable concatenated coiled-coil dimer modules, unraveling the principles for coexisting condensates, chemical regulation, formation from either one or two polypeptide components in mammalian cells.
- Maruša Ramšak
- , Dominique A. Ramirez
- & Roman Jerala
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access Haplotype-based inference of recent effective population size in modern and ancient DNA samples
Haplotype-based inference of recent effective population size in modern and ancient DNA samples
The authors introduce a new computational method, HapNe, for inferring the recent effective size of human populations. HapNe does not require high-quality genotype data, making it suitable for the study of ancient DNA samples.
- Romain Fournier
- , Zoi Tsangalidou
- & Pier Francesco Palamara
-
Article
| Open Access Autoencoder neural networks enable low dimensional structure analyses of microbial growth dynamics
Autoencoder neural networks enable low dimensional structure analyses of microbial growth dynamics
Here, the authors apply autoencoder neural networks to show that microbial growth dynamics can be compressed into low-dimensional representations and reconstructed with high fidelity, facilitating quantitative predictions and deduction of potential mechanisms.
- Yasa Baig
- , Helena R. Ma
- & Lingchong You
-
Article
| Open Access Augmenting interpretable models with large language models during training
Augmenting interpretable models with large language models during training
Prediction and interpretation tasks may be challenging in high-stakes applications, such as medical decision-making, or systems with compute-limited hardware. The authors introduce an augmented framework for leveraging the knowledge learned by Large Language Models to build interpretable models which are both accurate and efficient.
- Chandan Singh
- , Armin Askari
- & Jianfeng Gao
-
Article
| Open Access CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
Dose-response curves are ubiquitous in pharmacology and biology, yet potency and effect size are often estimated even when there is no response. Here, authors present a statistical framework to assess curve significance and demonstrate how this aids drug mode of action analysis in large public datasets.
- Florian P. Bayer
- , Manuel Gander
- & Matthew The
-
Article
| Open Access Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
A pan-betacoronavirus vaccine will likely require the elicitation of antibodies against spike regions conserved across diverse coronaviruses. Here, authors computationally engineer and experimentally validate immunogens to elicit antibodies against two such spike regions.
- A. Brenda Kapingidza
- , Daniel J. Marston
- & Mihai L. Azoitei
-
Article
| Open Access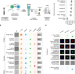 Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Therapeutic antibodies are crucial in treating severe diseases. Here, the authors introduce scifAI, an open-source explainable AI framework for analyzing imaging flow cytometry data, enabling rapid screening of therapeutic antibody candidates.
- Sayedali Shetab Boushehri
- , Katharina Essig
- & Fabian Schmich
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 ECOLE: Learning to call copy number variants on whole exome sequencing data
ECOLE: Learning to call copy number variants on whole exome sequencing data