Featured
-
-
Article
| Open Access Integrative genome-wide analyses identify novel loci associated with kidney stones and provide insights into its genetic architecture
Integrative genome-wide analyses identify novel loci associated with kidney stones and provide insights into its genetic architecture
Kidney stone disease is a complex disorder with high heritability and prevalence. Here, the authors perform a large genome-wide association study meta-analysis, identifying 28 new loci and genes potentially involved in disease etiology.
- Xingjie Hao
- , Zhonghe Shao
- & Chaolong Wang
-
Article
| Open Access Direct-acting antiviral resistance of Hepatitis C virus is promoted by epistasis
Direct-acting antiviral resistance of Hepatitis C virus is promoted by epistasis
This study reveals that mutations of the hepatitis C virus act collectively to confer resistance against direct-acting antiviral drugs. This can aid the development of drugs that are less prone to resistance.
- Hang Zhang
- , Ahmed Abdul Quadeer
- & Matthew R. McKay
-
Article
| Open Access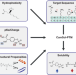 Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids
Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids
Posttranslationally modified amino acids are crucial in physiology and drug development as they alter physicochemical properties such as the solubility of proteins. Here the authors describe CamSolPTM, a software that accurately predicts the solubility of proteins containing these residues.
- Marc Oeller
- , Ryan J. D. Kang
- & Michele Vendruscolo
-
Article
| Open Access ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
Inverse Protein Folding is a critical component of protein design. Here, authors introduce ProRefiner, a deep-learning model for IPF that exhibits both high performance and memory efficiency, thereby contributing to advancements in protein design.
- Xinyi Zhou
- , Guangyong Chen
- & Pheng Ann Heng
-
Article
| Open Access Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
The authors compare the heat shock response between different cell lines and stimuli and reveal the genome-wide binding of its master transcription factor HSF1 as a platform for context-specific transcription activation.
- Sayantani Ghosh Dastidar
- , Bony De Kumar
- & Sergei Nechaev
-
Article
| Open Access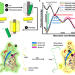 Simulation-guided engineering of split GFPs with efficient β-strand photodissociation
Simulation-guided engineering of split GFPs with efficient β-strand photodissociation
Green fluorescent proteins (GFPs) are ubiquitous for protein tagging and live cell imaging. Here, authors have used computational methods to engineer a fast-dissociating split GFP, which could be used to study macromolecular interactions.
- Yasmin Shamsudin
- , Alice R. Walker
- & Steven G. Boxer
-
Article
| Open Access Gut microbial structural variation associates with immune checkpoint inhibitor response
Gut microbial structural variation associates with immune checkpoint inhibitor response
Here, using datasets from the gut microbiome of 996 patients from seven clinical trials, the authors characterize gut microbial genomic structural variants, located in species such as Akkermansia muciniphila, Dorea formicigenerans, and Bacteroides caccae, that associate with hosts’ response and survival after immune checkpoint inhibitors treatment.
- Rong Liu
- , You Zou
- & Dao-Ming Wang
-
Article
| Open Access Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
The authors develop deep learning models to identify genome-wide polyA sites at nucleotide resolution and calculate site strength. They further examine genomic parameters regulating site usage and reveal genetic variants altering polyA activity.
- Emily Kunce Stroup
- & Zhe Ji
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access Functional annotation of enzyme-encoding genes using deep learning with transformer layers
Functional annotation of enzyme-encoding genes using deep learning with transformer layers
Functional annotation of open reading frames in microbial genomes remains substantially incomplete. Here, Kim et al. present a deep learning model that utilizes transformer layers as a neural network architecture to predict specific catalytic functions for enzyme-encoding genes of unknown function.
- Gi Bae Kim
- , Ji Yeon Kim
- & Sang Yup Lee
-
Article
| Open Access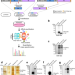 Interactome profiling of Crimean-Congo hemorrhagic fever virus glycoproteins
Interactome profiling of Crimean-Congo hemorrhagic fever virus glycoproteins
Here, Ning et al report the cellular interactomes of Crimean-Congo haemorrhagic fever virus glycoproteins and uncover a host restriction factor HAX1 that hijacks the viral glycoproteins to mitochondria, disabling progeny virion packaging.
- Shiyu Dai
- , Yuan-Qin Min
- & Yun-Jia Ning
-
Article
| Open Access Unistrand piRNA clusters are an evolutionarily conserved mechanism to suppress endogenous retroviruses across the Drosophila genus
Unistrand piRNA clusters are an evolutionarily conserved mechanism to suppress endogenous retroviruses across the Drosophila genus
To control transposable elements, fruit flies rely on distinct genomic regions called piRNA clusters. Here, new piRNA clusters were identified across diverse Drosophila species, displaying a conserved and specialised role in the control of endogenous retroviruses in ovarian somatic cells.
- Jasper van Lopik
- , Azad Alizada
- & Benjamin Czech Nicholson
-
Article
| Open Access CLOOME: contrastive learning unlocks bioimaging databases for queries with chemical structures
CLOOME: contrastive learning unlocks bioimaging databases for queries with chemical structures
Artificial intelligence can assist in obtaining knowledge from bioimaging data, but need human annotation. Here the authors use multimodal contrastive learning to link chemical structures and cell phenotypes, which can lead to foundation models for microscopy images.
- Ana Sanchez-Fernandez
- , Elisabeth Rumetshofer
- & Günter Klambauer
-
Article
| Open Access A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Pseudotime analysis is prevalent in single-cell RNA-seq, but it remains challenging to perform it across multiple samples and experimental conditions. Here, the authors develop Lamian, a computational framework for multi-sample pseudotime analysis that adjusts for biological and technical variation to detect gene program changes along cell trajectories and across conditions.
- Wenpin Hou
- , Zhicheng Ji
- & Hongkai Ji
-
Article
| Open Access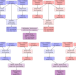 Leveraging information between multiple population groups and traits improves fine-mapping resolution
Leveraging information between multiple population groups and traits improves fine-mapping resolution
Statistical fine-mapping helps to pinpoint likely causal variants underlying genetic association signals, and can be enhanced by using multi-ancestry datasets. Here, the authors introduce MGflashfm, a fine-mapping method for pinpointing likely causal variants amongst multiple traits and population groups.
- Feng Zhou
- , Opeyemi Soremekun
- & Jennifer L. Asimit
-
Article
| Open Access Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial omics technologies reveal the organisation of cells in various biological systems. Here, authors propose SLAT, a graph-based algorithm for aligning heterogenous data across technologies, modalities and timepoints, enabling spatiotemporal reconstruction of complex developmental processes.
- Chen-Rui Xia
- , Zhi-Jie Cao
- & Ge Gao
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang
-
Article
| Open Access trRosettaRNA: automated prediction of RNA 3D structure with transformer network
trRosettaRNA: automated prediction of RNA 3D structure with transformer network
Here, authors develop trRosettaRNA, a deep learning-based approach for predicting RNA 3D structures. Blind tests demonstrate that the automated predictions compete effectively with top human predictions on natural RNAs.
- Wenkai Wang
- , Chenjie Feng
- & Jianyi Yang
-
Article
| Open Access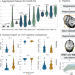 Automated temporalis muscle quantification and growth charts for children through adulthood
Automated temporalis muscle quantification and growth charts for children through adulthood
Temporalis muscle thickness is a promising marker of lean muscle mass but has had limited utility due to its unknown normal growth trajectory and lack of standardized measurement. Here, the authors develop an automated deep learning pipeline to accurately measure temporalis muscle thickness from routine brain magnetic resonance imaging.
- Anna Zapaishchykova
- , Kevin X. Liu
- & Benjamin H. Kann
-
Article
| Open Access EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
The study reveals limitations in widely used RNA-seq aligners, which create 'phantom' introns in reference databases. The authors introduce EASTR, a computational tool that not only enhances alignment accuracy but also uncovers existing annotation errors. This improvement bolsters the dependability of subsequent RNA-seq analyses.
- Ida Shinder
- , Richard Hu
- & Mihaela Pertea
-
Article
| Open Access Asymmetric conformations and lipid interactions shape the ATP-coupled cycle of a heterodimeric ABC transporter
Asymmetric conformations and lipid interactions shape the ATP-coupled cycle of a heterodimeric ABC transporter
Multidrug resistance through active extrusion of molecules by transporters is a pressing clinical problem. Here, authors dissect the mechanism by which an ABC transporter from B. Subtilis binds and removes drugs by consuming the energy of ATP hydrolysis.
- Qingyu Tang
- , Matt Sinclair
- & Hassane S. Mchaourab
-
Article
| Open Access A single cell genomics atlas of the Drosophila larval eye reveals distinct photoreceptor developmental timelines
A single cell genomics atlas of the Drosophila larval eye reveals distinct photoreceptor developmental timelines
The Drosophila eye is a powerful model system to study the dynamics of cell differentiation, cell state transitions, cell maturation, and pattern formation. Here, the authors report transcriptomic and chromatin accessibility data for all known cell types in the developing larval eye.
- Komal Kumar Bollepogu Raja
- , Kelvin Yeung
- & Graeme Mardon
-
Article
| Open Access Inflammation in the tumor-adjacent lung as a predictor of clinical outcome in lung adenocarcinoma
Inflammation in the tumor-adjacent lung as a predictor of clinical outcome in lung adenocarcinoma
Lung adenocarcinoma is often curable when diagnosed at an early stage, but a subsection of patients will progress. Here, the authors use multi-omics profiling to show that gene expression data can predict clinical outcome.
- Igor Dolgalev
- , Hua Zhou
- & Aristotelis Tsirigos
-
Article
| Open Access Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Speos: an ensemble graph representation learning framework to predict core gene candidates for complex diseases
Understanding phenotype-genotype relationships is a grand challenge of current biological research. Here, the authors use graph representation learning to identify human genes which display key characteristics of core genes for five complex diseases.
- Florin Ratajczak
- , Mitchell Joblin
- & Matthias Heinig
-
Article
| Open Access replicAnt: a pipeline for generating annotated images of animals in complex environments using Unreal Engine
replicAnt: a pipeline for generating annotated images of animals in complex environments using Unreal Engine
Deep learning-based computer vision tools are transforming animal behavioural research; however, many challenges remain. Here, Plum et al. present replicAnt, a novel tool for generating synthetic data to train computer vision models for animal behaviour studies, reducing the need for manual annotation.
- Fabian Plum
- , René Bulla
- & David Labonte
-
Article
| Open Access NIPMAP: niche-phenotype mapping of multiplex histology data by community ecology
NIPMAP: niche-phenotype mapping of multiplex histology data by community ecology
Multiplex histology faces the challenge of integrating tissue architecture with the identification of relevant spatial cellular phenotypes. Using community ecology principles, the authors propose NIPMAP, a tool for niche-phenotype mapping of multiplex histology data.
- Anissa El Marrahi
- , Fabio Lipreri
- & Jean Hausser
-
Article
| Open Access Unappreciated subcontinental admixture in Europeans and European Americans and implications for genetic epidemiology studies
Unappreciated subcontinental admixture in Europeans and European Americans and implications for genetic epidemiology studies
European ancestry individuals are not typically treated as admixed in genetic studies. Here, the authors detect higher than expected admixture in European populations, which could potentially affect the results of genetic studies if it is not accounted for.
- Mateus H. Gouveia
- , Amy R. Bentley
- & Daniel Shriner
-
Article
| Open Access Evolutionary design of explainable algorithms for biomedical image segmentation
Evolutionary design of explainable algorithms for biomedical image segmentation
Deep learning frameworks require large human-annotated datasets for training and the resulting ‘black box’ models are difficult to interpret. Here, the authors present Kartezio; a modular Cartesian Genetic Programming-based computational strategy that generates fully transparent and easily interpretable image processing pipelines.
- Kévin Cortacero
- , Brienne McKenzie
- & Sylvain Cussat-Blanc
-
Article
| Open Access Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Successful induction of broadly neutralizing antibodies is a main challenge in HIV vaccine development. The authors provide a framework to determine probabilities of antibody sequence development and show that uninfected and chronically infected individuals have the same chances to develop HIV-1 neutralizing antibodies.
- Christoph Kreer
- , Cosimo Lupo
- & Florian Klein
-
Article
| Open Access A generative adversarial network model alternative to animal studies for clinical pathology assessment
A generative adversarial network model alternative to animal studies for clinical pathology assessment
Generative AI has the potential to transform the way chemical and drug safety research is conducted. Here the authors show AnimalGAN, a model developed using Generative Adversarial Networks, which simulates virtual animal experiments to generate multidimensional rat clinical pathology measurements.
- Xi Chen
- , Ruth Roberts
- & Weida Tong
-
Article
| Open Access Tracing cancer evolution and heterogeneity using Hi-C
Tracing cancer evolution and heterogeneity using Hi-C
It is challenging to analyse chromosomal rearrangements in heterogeneous solid cancers. Here the authors present HiDENSEC, a method to jointly infer absolute copy number, ploidy, tumor purity and large-scale rearrangements from Hi-C data. The increased statistical power afforded by joint inference enables novel insights into cancer genome evolution.
- Dan Daniel Erdmann-Pham
- , Sanjit Singh Batra
- & Dirk Hockemeyer
-
Article
| Open Access LensAge index as a deep learning-based biological age for self-monitoring the risks of age-related diseases and mortality
LensAge index as a deep learning-based biological age for self-monitoring the risks of age-related diseases and mortality
Age is closely related to health, but chronologically defined age often disagrees with biological age. Here, the authors develop an indicator of biological age - LensAge index - to reveal individuals’ aging level, and it can be implemented with smartphones, showing potential for self-monitoring of aging.
- Ruiyang Li
- , Wenben Chen
- & Haotian Lin
-
Article
| Open Access Poor sleep and shift work associate with increased blood pressure and inflammation in UK Biobank participants
Poor sleep and shift work associate with increased blood pressure and inflammation in UK Biobank participants
Circadian disruption is linked to increased blood pressure and heart disease risk. Here, the authors show a positive association between circadian disruption and blood pressure (SBP/DBP) regulation in males and females irrespective of age, weight and inflammatory status.
- Monica Kanki
- , Artika P. Nath
- & Morag J. Young
-
Article
| Open Access Fetal biometry and amniotic fluid volume assessment end-to-end automation using Deep Learning
Fetal biometry and amniotic fluid volume assessment end-to-end automation using Deep Learning
Fetal biometry and amniotic fluid volume are essential but strenuous measurements in fetal ultrasound screening. Here, the authors show that deep learning models can automate these measurements with high accuracy, using a large and diverse dataset of Moroccan fetal ultrasound images.
- Saad Slimani
- , Salaheddine Hounka
- & El Houssine Bouyakhf
-
Article
| Open Access Dissecting the human leptomeninges at single-cell resolution
Dissecting the human leptomeninges at single-cell resolution
The meninges protect the central nervous system at the brain border, and its dysfunction can lead to neural inflammation and cell damage. Here, the authors uncover the gene signatures of diverse cell types in the aged human leptomeninges and highlight their changes in Alzheimer’s Disease.
- Nicola A. Kearns
- , Artemis Iatrou
- & Yanling Wang
-
Article
| Open Access Genome-wide association analysis of plasma lipidome identifies 495 genetic associations
Genome-wide association analysis of plasma lipidome identifies 495 genetic associations
The human plasma lipidome captures risk for cardiometabolic diseases. Here, the authors perform univariate and multivariate genome-wide analyses of 179 lipid species in 7174 Finnish individuals, revealing genetic links between diseases and lipid species beyond the standard lipids HDL-Cholesterol, LDL-Cholesterol, Triglycerides, and total Cholesterol.
- Linda Ottensmann
- , Rubina Tabassum
- & Matti Pirinen
-
Article
| Open Access Proteomics reveal biomarkers for diagnosis, disease activity and long-term disability outcomes in multiple sclerosis
Proteomics reveal biomarkers for diagnosis, disease activity and long-term disability outcomes in multiple sclerosis
Precise biomarkers for multiple sclerosis prognosis are vital for treatment decisions. Here, the authors identify specific proteins in cerebrospinal fluid that can predict short-term disease activity and long-term disability outcomes in persons with multiple sclerosis.
- Julia Åkesson
- , Sara Hojjati
- & Mika Gustafsson
-
Article
| Open Access Isoform-resolved transcriptome of the human preimplantation embryo
Isoform-resolved transcriptome of the human preimplantation embryo
Human embryo development involves extensive transcriptional remodeling. In this study, the authors apply long- and short-read RNA-Seq to profile the transcriptomes of 73 human preimplantation embryos spanning zygotic to blastocyst stages, identifying tens of thousands of additional isoforms transcribed from both known and unannotated gene loci.
- Denis Torre
- , Nancy J. Francoeur
- & Robert Sebra
-
Article
| Open Access XMAP: Cross-population fine-mapping by leveraging genetic diversity and accounting for confounding bias
XMAP: Cross-population fine-mapping by leveraging genetic diversity and accounting for confounding bias
Fine-mapping prioritizes risk variants identified by genome-wide association studies to uncover biological mechanisms underlying complex traits. Here, the authors develop a reliable fine-mapping method (XMAP) by leveraging genetic diversity and accounting for confounding bias.
- Mingxuan Cai
- , Zhiwei Wang
- & Can Yang
-
Article
| Open Access Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Chameleolyser enables the accurate identification of genetic variants hidden within complex regions of the genome. Its application uncovers the disease-explanatory variant in 25 previously undiagnosed patients.
- Wouter Steyaert
- , Lonneke Haer-Wigman
- & Christian Gilissen
-
Article
| Open Access Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Predicting dynamic RNA-RBP interactions in diverse cell lines is an important challenge in unravelling RNA function and post-transcriptional regulatory mechanisms. Here, authors develop HDRNet, an end-to-end deep-learning-based framework for accurately predicting dynamic RBP binding events across various cellular conditions.
- Haoran Zhu
- , Yuning Yang
- & Xiangtao Li
-
Article
| Open Access Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Unnatural base pairing xenonucleic acids (XNAs) can be used to expand life’s alphabet beyond ATGC. Here, authors show strategies for enzymatic synthesis and next-generation nanopore sequencing of XNA base pairs for reading and writing 12-letter DNA (ATGCBSPZXKJV).
- Hinako Kawabe
- , Christopher A. Thomas
- & Jorge A. Marchand
-
Article
| Open Access Systematic review and integrated data analysis reveal diverse pangolin-associated microbes with infection potential
Systematic review and integrated data analysis reveal diverse pangolin-associated microbes with infection potential
The diversity and spillover potential of pangolin-associated microbes are not fully understood. Here, the authors describe the distribution and spectrum of reported pangolin microbes by integrating data from multiple sources and assess their potential to emerge as human pathogens.
- Run-Ze Ye
- , Xiao-Yang Wang
- & Wu-Chun Cao
-
Article
| Open Access De novo design of knotted tandem repeat proteins
De novo design of knotted tandem repeat proteins
This study reports the successful de novo design of a trefoil knotted protein fold for which the crystal structure agrees closely with the intended trefoil knot topology.
- Lindsey A. Doyle
- , Brittany Takushi
- & Philip Bradley
-
Article
| Open Access A deep population reference panel of tandem repeat variation
A deep population reference panel of tandem repeat variation
Tandem repeats (TRs) comprise some of the most polymorphic regions of the human genome but are difficult to study. Here, the authors develop an ensemble-based genotyping method and characterize 1.7 million TRs across 3,550 humans from diverse populations.
- Helyaneh Ziaei Jam
- , Yang Li
- & Melissa Gymrek
-
Article
| Open Access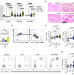 Immune checkpoint inhibitor-induced colitis is mediated by polyfunctional lymphocytes and is dependent on an IL23/IFNγ axis
Immune checkpoint inhibitor-induced colitis is mediated by polyfunctional lymphocytes and is dependent on an IL23/IFNγ axis
Immune checkpoint inhibitors (CPI) could effectively target cancers that are resistant to traditional therapy but may initiate immune related adverse effects, such as colitis. Here, authors characterise the gut immune microenvironment during CPI-colitis by bulk RNA sequencing, single-cell RNA sequencing and flow cytometry, and find that interleukin 23 plays an important role in promoting inflammation via cytotoxic polyfunctional IFNγ-producing lymphocytes.
- Jonathan W. Lo
- , Domenico Cozzetto
- & Nick Powell
-
Article
| Open Access A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
A genomic appraisal of invasive Salmonella Typhimurium and associated antibiotic resistance in sub-Saharan Africa
Invasive Salmonella Typhimurium bloodstream infection causes a significant public health burden in sub-Saharan Africa. Here, the authors analyse whole genome sequences of 1,302 S. Typhimurium isolates from Africa and describe its evolution, geographic spread, and antimicrobial resistance characteristics.
- Sandra Van Puyvelde
- , Tessa de Block
- & Octavie Lunguya
-
Article
| Open Access De novo genome assembly depicts the immune genomic characteristics of cattle
De novo genome assembly depicts the immune genomic characteristics of cattle
The genomic organisation of the cattle genome has been assembled to a limited level of resolution. Here using long range nanopore sequencing the authors present a cattle genome assembly concentrating on characterising the immunogenomic loci, particularly T cell receptor (TR), immunoglobulin (IG) and MHC genes, from one animal.
- Ting-Ting Li
- , Tian Xia
- & Tao Li
-
Article
| Open Access Ligand activation mechanisms of human KCNQ2 channel
Ligand activation mechanisms of human KCNQ2 channel
The potassium channel KCNQ2 can be activated by analgesics and antiepileptic drugs via an unclear mechanism. Here authors report structures of KCNQ2-CaM in complex with cannabidiol, PIP2, and HN37 and elucidate the mechanisms of activation.
- Demin Ma
- , Yueming Zheng
- & Jiangtao Guo
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Genome-wide CRISPR off-target prediction and optimization using RNA-DNA interaction fingerprints
Genome-wide CRISPR off-target prediction and optimization using RNA-DNA interaction fingerprints