Featured
-
-
Article
| Open Access Stochastic priming and spatial cues orchestrate heterogeneous clonal contribution to mouse pancreas organogenesis
Stochastic priming and spatial cues orchestrate heterogeneous clonal contribution to mouse pancreas organogenesis
The pancreas arises from a small population of cells but how individual cells contribute to organ formation is unclear. Here, the authors deconstruct pancreas organogenesis into clonal units, showing that single progenitors give rise to heterogeneous multi-lineage and endocrinogenic single-lineage clones.
- Hjalte List Larsen
- , Laura Martín-Coll
- & Anne Grapin-Botton
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access The interdependent network of gene regulation and metabolism is robust where it needs to be
The interdependent network of gene regulation and metabolism is robust where it needs to be
Although networks of interacting genes and metabolic reactions are interdependent, they have largely been treated as separate systems. Here the authors apply a statistical framework for interdependent networks to E. coli, and show that it is sensitive to gene and protein perturbations but robust against metabolic changes.
- David F. Klosik
- , Anne Grimbs
- & Marc-Thorsten Hütt
-
Article
| Open Access Accurate immune repertoire sequencing reveals malaria infection driven antibody lineage diversification in young children
Accurate immune repertoire sequencing reveals malaria infection driven antibody lineage diversification in young children
Somatic hypermutation of antibodies can occur in infants but are difficult to track. Here the authors present a new method called MIDCIRS for deep quantitative repertoire sequencing with few cells, and show infants as young as 3 months can expand antibody lineage complexity in response to malaria infection.
- Ben S. Wendel
- , Chenfeng He
- & Ning Jiang
-
Article
| Open Access Identifying topologically associating domains and subdomains by Gaussian Mixture model And Proportion test
Identifying topologically associating domains and subdomains by Gaussian Mixture model And Proportion test
Spatial organization of the genome plays a crucial role in regulating gene expression. Here the authors introduce GMAP, the Gaussian Mixture model And Proportion test, to identify topologically associating domains and subdomains in Hi-C data.
- Wenbao Yu
- , Bing He
- & Kai Tan
-
Article
| Open Access Discovery of a proteolytic flagellin family in diverse bacterial phyla that assembles enzymatically active flagella
Discovery of a proteolytic flagellin family in diverse bacterial phyla that assembles enzymatically active flagella
So far no enzymatic activity has been attributed to flagellin, the major component of bacterial flagella. Here the authors use bioinformatic analysis and identify a metallopeptidase insertion in flagellins from 74 bacterial species and show that recombinant flagellin and flagellar filaments have proteolytic activity.
- Ulrich Eckhard
- , Hina Bandukwala
- & Andrew C. Doxey
-
Article
| Open Access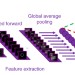 Reconstructing cell cycle and disease progression using deep learning
Reconstructing cell cycle and disease progression using deep learning
The interpretation of information-rich, high-throughput single-cell data is a challenge requiring sophisticated computational tools. Here the authors demonstrate a deep convolutional neural network that can classify cell cycle status on-the-fly.
- Philipp Eulenberg
- , Niklas Köhler
- & F. Alexander Wolf
-
Article
| Open Access Multivariate discovery and replication of five novel loci associated with Immunoglobulin G N-glycosylation
Multivariate discovery and replication of five novel loci associated with Immunoglobulin G N-glycosylation
Multivariate analysis methods can uncover the relationship between phenotypic measures characterised by modern omic techniques. Here the authors conduct a multivariate GWAS on IgG N-glycosylation phenotypes and identify 5 novel loci enriched in immune system genes.
- Xia Shen
- , Lucija Klarić
- & Yurii S. Aulchenko
-
Article
| Open Access Non-parametric genetic prediction of complex traits with latent Dirichlet process regression models
Non-parametric genetic prediction of complex traits with latent Dirichlet process regression models
Genetic prediction of complex traits with polygenic architecture has wide application from animal breeding to disease prevention. Here, Zeng and Zhou develop a non-parametric genetic prediction method based on latent Dirichlet Process regression models.
- Ping Zeng
- & Xiang Zhou
-
Article
| Open Access An in-silico approach to predict and exploit synthetic lethality in cancer metabolism
An in-silico approach to predict and exploit synthetic lethality in cancer metabolism
Exploiting synthetic lethality is a promising approach for cancer therapy. Here, the authors present an approach to identifying such interactions by finding genetic minimal cut sets (gMCSs) that block cancer proliferation, and apply it to study the lethality of RRM1 inhibition in multiple myeloma.
- Iñigo Apaolaza
- , Edurne San José-Eneriz
- & Francisco J. Planes
-
Article
| Open Access Integrative genomics of microglia implicates DLG4 (PSD95) in the white matter development of preterm infants
Integrative genomics of microglia implicates DLG4 (PSD95) in the white matter development of preterm infants
Inflammation mediated by microglia plays a key role in brain injury associated with preterm birth, but little is known about the microglial response in preterm infants. Here, the authors integrate molecular and imaging data from animal models and preterm infants, and find that microglial expression of DLG4 plays a role.
- Michelle L. Krishnan
- , Juliette Van Steenwinckel
- & Pierre Gressens
-
Article
| Open Access Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation
Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation
Metazoan genomes contain many clusters of conserved noncoding elements. Here, the authors provide evidence that these clusters coincide with distinct topologically associating domains in humans and Drosophila, revealing a conserved regulatory genomic architecture.
- Nathan Harmston
- , Elizabeth Ing-Simmons
- & Boris Lenhard
-
Article
| Open Access Stromal and epithelial transcriptional map of initiation progression and metastatic potential of human prostate cancer
Stromal and epithelial transcriptional map of initiation progression and metastatic potential of human prostate cancer
Stromal cells contribute to tumor development but the mechanisms regulating this process are still unclear. Here the authors analyze gene expression profiles in the prostate and show that stromal gene signature changes ahead of the epithelial gene signature as prostate cancer initiates and progresses.
- Svitlana Tyekucheva
- , Michaela Bowden
- & Massimo Loda
-
Article
| Open Access Using ALoFT to determine the impact of putative loss-of-function variants in protein-coding genes
Using ALoFT to determine the impact of putative loss-of-function variants in protein-coding genes
Variants causing loss of function (LoF) of human genes have clinical implications. Here, the authors present a method to predict disease-causing potential of LoF variants, ALoFT (annotation of Loss-of-Function Transcripts) and show its application to interpreting LoF variants in different contexts.
- Suganthi Balasubramanian
- , Yao Fu
- & Mark Gerstein
-
Article
| Open Access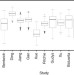 On the performance of pre-microRNA detection algorithms
On the performance of pre-microRNA detection algorithms
As the experimental discovery of microRNAs (miRNAs) is cumbersome, computational tools have been developed for the prediction of pre-miRNAs. Here the authors develop a framework to assess the performance of existing and novel pre-miRNA prediction tools and provide guidelines for selecting an appropriate approach for a given data set.
- Müşerref Duygu Saçar Demirci
- , Jan Baumbach
- & Jens Allmer
-
Article
| Open Access An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network
An integrated bioinformatics platform for investigating the human E3 ubiquitin ligase-substrate interaction network
Protein stability modulation by E3 ubiquitin ligases is an important layer of functional regulation, but screening for E3 ligase-substrate interactions is time-consuming and costly. Here, the authors take an in silico naïve Bayesian classifier approach to integrate multiple lines of evidence for E3-substrate prediction, enabling prediction of the proteome-wide human E3 ligase interaction network.
- Yang Li
- , Ping Xie
- & Fuchu He
-
Article
| Open Access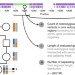 Estimating the human mutation rate from autozygous segments reveals population differences in human mutational processes
Estimating the human mutation rate from autozygous segments reveals population differences in human mutational processes
Estimates of human mutation rates differ substantially based on the approach. Here, the authors present a multi-generational estimate from the autozygous segment in a non-European population that gives insight into the contribution of post-zygotic mutations and population-specific mutational processes.
- Vagheesh M. Narasimhan
- , Raheleh Rahbari
- & Richard Durbin
-
Article
| Open Access The origin of a primordial genome through spontaneous symmetry breaking
The origin of a primordial genome through spontaneous symmetry breaking
Early molecules of life likely served both as templates and catalysts, raising the question of how functionally distinct genomes and enzymes arose. Here, the authors show that conflict between evolution at the molecular and cellular levels can drive functional differentiation of the two strands of self-replicating molecules and lead to copy number differences between the two.
- Nobuto Takeuchi
- , Paulien Hogeweg
- & Kunihiko Kaneko
-
Article
| Open Access Annotating pathogenic non-coding variants in genic regions
Annotating pathogenic non-coding variants in genic regions
While non-coding synonymous and intronic variants are often not under strong selective constraint, they can be pathogenic through affecting splicing or transcription. Here, the authors develop a score that uses sequence context alterations to predict pathogenicity of synonymous and non-coding genetic variants, and provide a web server of pre-computed scores.
- Sahar Gelfman
- , Quanli Wang
- & David B. Goldstein
-
Article
| Open Access Unraveling a tumor type-specific regulatory core underlying E2F1-mediated epithelial-mesenchymal transition to predict receptor protein signatures
Unraveling a tumor type-specific regulatory core underlying E2F1-mediated epithelial-mesenchymal transition to predict receptor protein signatures
Deregulation of E2F family transcription factors is associated with cancer progression and metastasis. Here, the authors construct a map of the regulatory network around the E2F family, and using gene expression profiles, identify tumour type-specific regulatory cores and receptor expression signatures associated with epithelial-mesenchymal transition in bladder and breast cancer.
- Faiz M. Khan
- , Stephan Marquardt
- & Brigitte M. Pützer
-
Article
| Open Access Bayesian association scan reveals loci associated with human lifespan and linked biomarkers
Bayesian association scan reveals loci associated with human lifespan and linked biomarkers
Along with various environmental factors, a complex genetic architecture influences human lifespan. Here, McDaid and colleagues reveal novel loci associated with human lifespan and linked biomarkers by Bayesian association scan.
- Aaron F. McDaid
- , Peter K. Joshi
- & Zoltán Kutalik
-
Article
| Open Access Physical limits to biomechanical sensing in disordered fibre networks
Physical limits to biomechanical sensing in disordered fibre networks
Cells in the connective tissue are surrounded by a heterogeneous network of biopolymers. Here, the authors investigate how such heterogeneity affects cellular mechanosensing by simulating the deformation response of experimental and modelled biopolymer networks to locally applied forces.
- Farzan Beroz
- , Louise M. Jawerth
- & Ned S. Wingreen
-
Article
| Open Access De novo active sites for resurrected Precambrian enzymes
De novo active sites for resurrected Precambrian enzymes
The emergence of novel catalytic functions in ancient proteins likely played a role in the evolution of modern enzymes. Here, the authors use protein sequences from Precambrian beta-lactamases and demonstrate that a single hydrophobic-to-ionizable amino acid mutation can lead to substantial Kemp eliminase activity.
- Valeria A. Risso
- , Sergio Martinez-Rodriguez
- & Jose M. Sanchez-Ruiz
-
Article
| Open Access Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets
Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets
Increased availability of large-scale molecular profiling has enabled system-level monitoring of molecular effects of candidate therapeutics. Here, the authors take advantage of such data to show that the ability of a drug to reverse cancer-associated gene expression changes is indicative of itsin vitroanti-proliferative efficacy, allowing them to identify novel compounds against liver cancer.
- Bin Chen
- , Li Ma
- & Atul J. Butte
-
Article
| Open Access Skin parasite landscape determines host infectiousness in visceral leishmaniasis
Skin parasite landscape determines host infectiousness in visceral leishmaniasis
Parasitemia has been considered the main determinant of visceral leishmaniasis transmission. By combining imaging, qPCR and experimental xenodiagnoses with mathematical models, Doehl et al. argue that the patchy landscape of parasites in the skin is necessary to explain infectiousness.
- Johannes S. P. Doehl
- , Zoe Bright
- & Paul M. Kaye
-
Article
| Open Access Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis
Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis
RNA-seq is widely used for transcriptome analysis. Here, the authors analyse a wide spectrum of RNA-seq workflows and present a comprehensive analysis protocol named RNACocktail as well as a computational pipeline leveraging the widely used tools for accurate RNA-seq analysis.
- Sayed Mohammad Ebrahim Sahraeian
- , Marghoob Mohiyuddin
- & Hugo Y. K. Lam
-
Article
| Open Access A pan-cancer genome-wide analysis reveals tumour dependencies by induction of nonsense-mediated decay
A pan-cancer genome-wide analysis reveals tumour dependencies by induction of nonsense-mediated decay
Nonsense-mediated decay (NMD) eliminates transcripts with premature stop codons and has been linked to cancer genesis. Here, the authors develop an algorithm to predict NMD and perform a pan-cancer analysis that finds that some hypermutated cancers are dependent on mutations that elicit NMD.
- Zhiyuan Hu
- , Christopher Yau
- & Ahmed Ashour Ahmed
-
Article
| Open Access Growth-coupled overproduction is feasible for almost all metabolites in five major production organisms
Growth-coupled overproduction is feasible for almost all metabolites in five major production organisms
Coupling of growth and product synthesis is an important principle in metabolic engineering, but its range of applicability is unclear. Here, the authors use a dedicated computational framework to study the feasibility of coupling the production of metabolites to growth in the genome-scale metabolic models of five production organisms, and show that coupling can be achieved for most metabolites.
- Axel von Kamp
- & Steffen Klamt
-
Article
| Open Access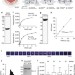 Lipid-mediated PX-BAR domain recruitment couples local membrane constriction to endocytic vesicle fission
Lipid-mediated PX-BAR domain recruitment couples local membrane constriction to endocytic vesicle fission
The spatiotemporal regulation of membrane scaffolds recruitment and coupling between membrane deformation and fission in endocytosis are unclear. Here the authors show that lipid conversion at endocytic pits recruits SNX9, which couples local membrane constriction to fission in endocytosis.
- Johannes Schöneberg
- , Martin Lehmann
- & Frank Noé
-
Article
| Open Access Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
Reconstructing cell cycle pseudo time-series via single-cell transcriptome data
In single-cell RNA sequencing data of heterogeneous cell populations, cell cycle stage of individual cells would often be informative. Here, the authors introduce a computational model to reconstruct a pseudo-time series from single cell transcriptome data, identify the cell cycle stages, identify candidate cell cycle-regulated genes and recover the methylome changes during the cell cycle.
- Zehua Liu
- , Huazhe Lou
- & Ting Chen
-
Article
| Open Access A BaSiC tool for background and shading correction of optical microscopy images
A BaSiC tool for background and shading correction of optical microscopy images
Accurate quantification of bioimaging data is often confounded by uneven illumination (shading) in space and background variation in time. Here the authors present BaSiC, a Fiji plugin solving both issues. It requires fewer input images and is more robust to artefacts than existing shading correction tools.
- Tingying Peng
- , Kurt Thorn
- & Nassir Navab
-
Article
| Open Access Cross-orientation suppression in visual area V2
Cross-orientation suppression in visual area V2
V2 neurons exhibit complex and diverse selectivity for visual features. Here the authors use a statistical analytical framework to model V2 responses to natural stimuli and find three organizing principles, chief among them is the cross-orientation suppression that increases response selectivity.
- Ryan J. Rowekamp
- & Tatyana O. Sharpee
-
Article
| Open Access A molecular portrait of microsatellite instability across multiple cancers
A molecular portrait of microsatellite instability across multiple cancers
Some cancers with DNA mismatch repair deficiency display microsatellite instability. Here the authors analyse twenty three cancer types at the exome and whole-genome level, and identify loci with recurrent microsatellite instability that could be used to identify patients who would benefit from immunotherapy.
- Isidro Cortes-Ciriano
- , Sejoon Lee
- & Peter J. Park
-
Correspondence
| Open Access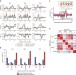 Correspondence: DNA shape is insufficient to explain binding
Correspondence: DNA shape is insufficient to explain binding
- Matthew J. Rossi
- , William K.M. Lai
- & B. Franklin Pugh
-
Article
| Open Access Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Robust quantification of the differentiation potential of single cells is a task of great importance. Here the authors integrate single-cell RNA-Seq profiles with a cellular interaction network to compute the signaling entropy, and show that it can identify normal and cancer stem-cell phenotypes.
- Andrew E. Teschendorff
- & Tariq Enver
-
Article
| Open Access Smooth 2D manifold extraction from 3D image stack
Smooth 2D manifold extraction from 3D image stack
Maximum Intensity Projection is a common tool to represent 3D biological imaging data in a 2D space, but it creates artefacts. Here the authors develop Smooth Manifold Extraction, an ImageJ/Fiji plugin, to preserve local spatial relationships when extracting the content of a 3D volume to a 2D space.
- Asm Shihavuddin
- , Sreetama Basu
- & Auguste Genovesio
-
Article
| Open Access Systematic discovery of mutation-specific synthetic lethals by mining pan-cancer human primary tumor data
Systematic discovery of mutation-specific synthetic lethals by mining pan-cancer human primary tumor data
There are no robust methods for systematically identifying mutation-specific synthetic lethal (SL) partners in cancer. Here, the authors develop a computational algorithm that uses pan-cancer data to detect mutation-andcancer-specific SL partners and they validate a novel SL interaction between mutant IDH and loss of ACACA in leukaemia.
- Subarna Sinha
- , Daniel Thomas
- & David L. Dill
-
Article
| Open Access Cancer-cell intrinsic gene expression signatures overcome intratumoural heterogeneity bias in colorectal cancer patient classification
Cancer-cell intrinsic gene expression signatures overcome intratumoural heterogeneity bias in colorectal cancer patient classification
Tumour expression profiling is currently used for prognostic and predictive purposes without taking into account the intra patient heterogeneity. Here the authors show that cancer cell specific signatures overcome the tumour heterogeneity effect and result in better classification of colorectal cancer patients.
- Philip D. Dunne
- , Matthew Alderdice
- & Mark Lawler
-
Article
| Open Access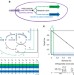 Redesigning metabolism based on orthogonality principles
Redesigning metabolism based on orthogonality principles
Growth-coupled designs for chemical production are limited by native metabolic networks’ optimality for growth. Here, the authors introduce pathway orthogonality as a measure of the independence of biomass and chemical production pathways, identify metabolic valves that allow substrate utilization to be switched between the two, and demonstrate advantages of orthogonal designs.
- Aditya Vikram Pandit
- , Shyam Srinivasan
- & Radhakrishnan Mahadevan
-
Article
| Open Access MiCASA is a new method for quantifying cellular organization
MiCASA is a new method for quantifying cellular organization
There are few methods available that can quantify relationships between cell types in tissue images. Here the authors present a quantitative method to evaluate cellular organization, validated in the mouse thymus and spinal cord, called Multitaper Circularly Averaged Spectral Analysis (MiCASA).
- Andrew Sornborger
- , Jie Li
- & Nancy R. Manley
-
Article
| Open Access A complete tool set for molecular QTL discovery and analysis
A complete tool set for molecular QTL discovery and analysis
Analysis of molecular quantitative trait loci (molQTL) can help interpret genome-wide association studies and requires efficient approaches to correct for multiple testing. This study describes a bioinformatics toolkit called QTLtool that can handle large data sets and quickly perform multiple types of molQTL analyses.
- Olivier Delaneau
- , Halit Ongen
- & Emmanouil T. Dermitzakis
-
Article
| Open Access An integrated model for detecting significant chromatin interactions from high-resolution Hi-C data
An integrated model for detecting significant chromatin interactions from high-resolution Hi-C data
Genome-wide chromosome conformation capture has helped us identify features of genome topology influencing biology but requires careful statistical analysis. Here the authors present HiC-DC, a bioinformatics method that can detect statistically significant regulatory interactions.
- Mark Carty
- , Lee Zamparo
- & Christina S. Leslie
-
Article
| Open Access A hybrid cloud read aligner based on MinHash and kmer voting that preserves privacy
A hybrid cloud read aligner based on MinHash and kmer voting that preserves privacy
Outsourcing computation for genomic data processing offers the ability to allocate massive computing power and storage on demand. Here, Popic and Batzoglou develop a hybrid cloud aligner for sequence read mapping that preserves privacy with competitive accuracy and speed.
- Victoria Popic
- & Serafim Batzoglou
-
Article
| Open Access Whole genome analysis of a schistosomiasis-transmitting freshwater snail
Whole genome analysis of a schistosomiasis-transmitting freshwater snail
Biomphalaria glabrata is a fresh water snail that acts as a host for trematode Schistosoma mansoni that causes intestinal infection in human. This work describes the genome and transcriptome analyses from 12 different tissues of B glabrata, and identify genes for snail behavior and evolution.
- Coen M. Adema
- , LaDeana W. Hillier
- & Richard K. Wilson
-
Article
| Open Access Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection
Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection
Cell lines are central to cancer research, but knowing which cell lines are the best representative of actual tumours is a major challenge. Here the authors provide a resource assessment of 65 renal cell lines to assist researchers in selecting suitable lines for studying specific renal carcinoma subtypes.
- Rileen Sinha
- , Andrew G. Winer
- & A. Ari Hakimi
-
Article
| Open Access Exchange pathways of plastoquinone and plastoquinol in the photosystem II complex
Exchange pathways of plastoquinone and plastoquinol in the photosystem II complex
Plastoquinone (PLQ) shuttles electrons between photosystem II (PSII) and cytochrome b6f. Here the authors perform molecular dynamics simulations and propose that PLQ enters the exchange cavity of PSII by a promiscuous diffusion mechanism whereby three different channels each act as entry and exit points.
- Floris J. Van Eerden
- , Manuel N. Melo
- & Siewert J. Marrink
-
Article
| Open Access Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Global patterns of gene transcription can be represented with reduced dimensionality. Here, the authors devise a method called Tradict that learns and uses 100 marker genes to predict transcriptome-wide pathway expression levels and patterns that reflect cell activity and state.
- Surojit Biswas
- , Konstantin Kerner
- & Philip A. Wigge
-
Article
| Open Access Subsampling scaling
Subsampling scaling
We can often observe only a small fraction of a system, which leads to biases in the inference of its global properties. Here, the authors develop a framework that enables overcoming subsampling effects, apply it to recordings from developing neural networks, and find that neural networks become critical as they mature.
- A. Levina
- & V. Priesemann
-
Article
| Open Access Similarity in viral and host promoters couples viral reactivation with host cell migration
Similarity in viral and host promoters couples viral reactivation with host cell migration
The coevolution of viruses and host cells can be mapped with interactomics. Here the authors identify coupling of human and viral promoters, and show that HIV-reactivation from dormancy is coincident with migration of HIV-infected cells owing to coupling of human CXCR4 and HIV LTR promoters.
- Kathrin Bohn-Wippert
- , Erin N. Tevonian
- & Roy D. Dar
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination
Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination