Article
|
Open Access
Featured
-
-
Article
| Open Access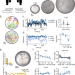 Precise visuomotor transformations underlying collective behavior in larval zebrafish
Precise visuomotor transformations underlying collective behavior in larval zebrafish
How visual social information informs movement is unclear. Here, the authors characterise the algorithm zebrafish use to transform visual inputs from neighbours into movement decisions during collective swimming behavior. The authors can also predict the neural circuits involved in transforming the visual input into movement decisions.
- Roy Harpaz
- , Minh Nguyet Nguyen
- & Florian Engert
-
Article
| Open Access An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
To characterize molecular changes during cell type transitions, the authors develop a method to simultaneously measure protein expression and thermal stability changes. They apply this approach to study differences between human pluripotent stem cells, their progenies, parental and allogeneic cells.
- Pierre Sabatier
- , Christian M. Beusch
- & Roman A. Zubarev
-
Article
| Open Access Towards inferring nanopore sequencing ionic currents from nucleotide chemical structures
Towards inferring nanopore sequencing ionic currents from nucleotide chemical structures
Nanopore sequencing allows users to identify nucleotide sequence from ionic currents. Here, the authors use deep learning to facilitate the de novo identification of modified nucleotides, particularly methylated cytosine and guanine, from the measured ionic currents without the need for controls.
- Hongxu Ding
- , Ioannis Anastopoulos
- & Benedict Paten
-
Article
| Open Access Discordant associations of educational attainment with ASD and ADHD implicate a polygenic form of pleiotropy
Discordant associations of educational attainment with ASD and ADHD implicate a polygenic form of pleiotropy
Autism Spectrum Disorder and Attention-Deficit/Hyperactivity Disorder are co-occurring neurodevelopmental conditions displaying strong, discordant polygenic associations with educational attainment. Here, the authors study genetic mechanisms underlying genome-wide correlation patterns across these traits.
- Ellen Verhoef
- , Jakob Grove
- & Beate St Pourcain
-
Article
| Open Access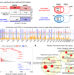 A systematic genome-wide mapping of oncogenic mutation selection during CRISPR-Cas9 genome editing
A systematic genome-wide mapping of oncogenic mutation selection during CRISPR-Cas9 genome editing
CRISPR-Cas9 gene editing can induce a p53 mediated damage response. Here the authors investigate the possibility of selection of pre-existing cancer driver mutations during CRISPR-Cas9 knockout based gene editing and identify KRAS mutants that may confer a selected advantage to edited cells.
- Sanju Sinha
- , Karina Barbosa
- & Eytan Ruppin
-
Article
| Open Access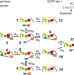 A general theoretical framework to design base editors with reduced bystander effects
A general theoretical framework to design base editors with reduced bystander effects
Base editors can edit target nucleotides, and identical ones that are within the editing window. Here the authors build an analytical model to propose general principles of editor design to reduce bystander effects.
- Qian Wang
- , Jie Yang
- & Anatoly B. Kolomeisky
-
Article
| Open Access Discovery of putative tumor suppressors from CRISPR screens reveals rewired lipid metabolism in acute myeloid leukemia cells
Discovery of putative tumor suppressors from CRISPR screens reveals rewired lipid metabolism in acute myeloid leukemia cells
CRISPR-based knockout screens in cancer cells have suggested the existence of proliferation suppressor genes (PSG). Here, the authors develop an approach to systematically identify them, and reveal a PSG module involved in fatty acid synthesis and tumour suppression in acute myeloid leukemia cell lines.
- W. Frank Lenoir
- , Micaela Morgado
- & Traver Hart
-
Article
| Open Access Co-evolution based machine-learning for predicting functional interactions between human genes
Co-evolution based machine-learning for predicting functional interactions between human genes
With the rise in number of eukaryotic species being fully sequenced, large scale phylogenetic profiling can give insights on gene function, Here, the authors describe a machine-learning approach that integrates co-evolution across eukaryotic clades to predict gene function and functional interactions among human genes.
- Doron Stupp
- , Elad Sharon
- & Yuval Tabach
-
Article
| Open Access Network analysis reveals rare disease signatures across multiple levels of biological organization
Network analysis reveals rare disease signatures across multiple levels of biological organization
Despite the clear causal relationship between genotype and phenotype in rare diseases, identifying the pathobiological mechanisms at various levels of biological organization remains a practical and conceptual challenge. Here, the authors introduce a network approach for evaluating the impact of rare gene defects across biological scales.
- Pisanu Buphamalai
- , Tomislav Kokotovic
- & Jörg Menche
-
Article
| Open Access Extended antibody-framework-to-antigen distance observed exclusively with broad HIV-1-neutralizing antibodies recognizing glycan-dense surfaces
Extended antibody-framework-to-antigen distance observed exclusively with broad HIV-1-neutralizing antibodies recognizing glycan-dense surfaces
Here, the authors analyse the distance between the body of an antibody and a protein antigen denoted as the Antibody-Framework-to-Antigen Distance (AFAD) for about 2000 non-redundant antibody-protein antigen complexes in the Protein Data Bank. They observe that antibodies with exceptionally long AFADs were all broad HIV-1-neutralizing antibodies that targeted densely glycosylated regions on the HIV-1-envelope trimer. The connection between long AFAD and dense glycan was further validated by the cryo-EM structure of antibody 2909 recognizing a glycan hole and by glycan shielding analyses based on molecular dynamics simulations.
- Myungjin Lee
- , Anita Changela
- & Peter D. Kwong
-
Article
| Open Access Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Deep learning (DL) can be used to automatically extract complex features from dynamic systems. Here, the authors combine high-content imaging, DL and mechanistic models to extract and explain drug-induced morphological changes in the growth of the fungus responsible for Asian soybean rust.
- Henry Cavanagh
- , Andreas Mosbach
- & Robert G. Endres
-
Article
| Open Access Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit
Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit
The exact protein features that control passage through the eukaryotic secretory system remain largely unknown. Here the authors report SECRiFY which they use to evaluate the secretory potential of polypeptides on a proteome-wide scale in yeast, revealing a role for flexibility and intrinsic disorder.
- Morgane Boone
- , Pathmanaban Ramasamy
- & Nico Callewaert
-
Article
| Open Access Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Normalisr removes technical bias in single-cell RNA-seq and detects gene differential and coexpression accurately and efficiently. It also infers gene regulatory and co-expression networks from conventional and CRISPR screen single-cell RNA-seq datasets.
- Lingfei Wang
-
Article
| Open Access Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen
Chromatin-accessibility estimation from single-cell ATAC-seq data with scOpen
scATAC-Seq yields data that is extremely sparse. Here, the authors present a computationally efficient imputation method called scOpen that improves the downstream analyses of scATAC-Seq data and use it to identify transcriptional regulators of kidney fibrosis.
- Zhijian Li
- , Christoph Kuppe
- & Ivan G. Costa
-
Article
| Open Access Benchmarking pipelines for subclonal deconvolution of bulk tumour sequencing data
Benchmarking pipelines for subclonal deconvolution of bulk tumour sequencing data
Subclonal deconvolution in cancer sequencing data is a complex task, and the optimal tools to use are unclear. Here, the authors systematically benchmark subclonal deconvolution pipelines with a comprehensive set of simulated tumour genomes and identify the best-performing methods.
- Georgette Tanner
- , David R. Westhead
- & Lucy F. Stead
-
Article
| Open Access Potential global impacts of alternative dosing regimen and rollout options for the ChAdOx1 nCoV-19 vaccine
Potential global impacts of alternative dosing regimen and rollout options for the ChAdOx1 nCoV-19 vaccine
The ChAdOx1 nCoV-19 vaccine requires two doses, but under limited supply single dose regimens have also been considered. Here, the authors show using static transmission modelling that under certain conditions it is optimal to more expediently administer a single dose to a larger proportion of the population.
- Ricardo Aguas
- , Anouska Bharath
- & Rima Shretta
-
Article
| Open Access Using secondary cases to characterize the severity of an emerging or re-emerging infection
Using secondary cases to characterize the severity of an emerging or re-emerging infection
Estimates of the severity of emerging infections did not consider the case ascertainment method, but secondary cases identified by contact tracing of index cases may be more reliable as they are less susceptible to ascertainment bias. Here, the authors perform a systematic review to quantify these differences and model their impacts for COVID-19.
- Tim K. Tsang
- , Can Wang
- & Benjamin J. Cowling
-
Article
| Open Access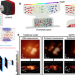 Computational optical sectioning with an incoherent multiscale scattering model for light-field microscopy
Computational optical sectioning with an incoherent multiscale scattering model for light-field microscopy
Light-field microscopy provides volumetric imaging at high speeds, but suffers from degradation in scattering tissue. Here, the authors present an incoherent multiscale scattering model which allows for quantitative 3D reconstruction in complex environments, and demonstrate dynamic imaging in vivo.
- Yi Zhang
- , Zhi Lu
- & Qionghai Dai
-
Article
| Open Access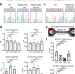 Gain-of-function cardiomyopathic mutations in RBM20 rewire splicing regulation and re-distribute ribonucleoprotein granules within processing bodies
Gain-of-function cardiomyopathic mutations in RBM20 rewire splicing regulation and re-distribute ribonucleoprotein granules within processing bodies
Mutations in the splicing factor RBM20 cause aggressive Dilated Cardiomyopathy. Here the authors generated RBM20 R636S mutants and knockout in human iPSC-derived cardiomyocytes. Mutant RBM20 showed different target RNA binding, altered splicing and localization to cytoplasmic processing bodies.
- Aidan M. Fenix
- , Yuichiro Miyaoka
- & Nathan Salomonis
-
Article
| Open Access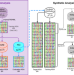 The generative capacity of probabilistic protein sequence models
The generative capacity of probabilistic protein sequence models
Generative models have become increasingly popular in protein design, yet rigorous metrics that allow the comparison of these models are lacking. Here, the authors propose a set of such metrics and use them to compare three popular models.
- Francisco McGee
- , Sandro Hauri
- & Allan Haldane
-
Article
| Open Access Accurate recognition of colorectal cancer with semi-supervised deep learning on pathological images
Accurate recognition of colorectal cancer with semi-supervised deep learning on pathological images
Machine-assisted recognition of colorectal cancer has been mainly focused on supervised deep learning that suffers from a significant bottleneck of requiring massive amounts of labeled data. Here, the authors propose a semi-supervised model based on the mean teacher architecture that provides pathological predictions at both patch- and patient-levels.
- Gang Yu
- , Kai Sun
- & Hong-Wen Deng
-
Article
| Open Access Multi-omics analysis identifies therapeutic vulnerabilities in triple-negative breast cancer subtypes
Multi-omics analysis identifies therapeutic vulnerabilities in triple-negative breast cancer subtypes
Triple negative breast cancer can be divided into additional subtypes. Here, using omics analyses, the authors show that in the mesenchymal subtype expression of MHC-1 is repressed and that this can be restored by using drugs that target subunits of the epigenetic modifier PRC2.
- Brian D. Lehmann
- , Antonio Colaprico
- & X. Steven Chen
-
Article
| Open Access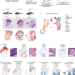 Spatially resolved transcriptomics reveals the architecture of the tumor-microenvironment interface
Spatially resolved transcriptomics reveals the architecture of the tumor-microenvironment interface
During tumor progression, cancer cells contact different neighboring cell types, but it is unclear how these interactions affect cancer cell behavior. Here, the authors use spatially resolved transcriptomics and single-cell RNA-seq to study the role of cilia at the tumormicroenvironment interface.
- Miranda V. Hunter
- , Reuben Moncada
- & Richard M. White
-
Article
| Open Access Refining models of archaic admixture in Eurasia with ArchaicSeeker 2.0
Refining models of archaic admixture in Eurasia with ArchaicSeeker 2.0
Existing methods to identify the presence of DNA from other hominin species can be limited in the ability to accurately estimate introgression waves, or can only be applied to specific populations. Here, the authors have developed a generalizable method to identify introgression in multi-wave situations.
- Kai Yuan
- , Xumin Ni
- & Shuhua Xu
-
Article
| Open Access The serine proteases dipeptidyl-peptidase 4 and urokinase are key molecules in human and mouse scar formation
The serine proteases dipeptidyl-peptidase 4 and urokinase are key molecules in human and mouse scar formation
Mechanisms triggering hypertrophic scar formation remain poorly understood. Here the authors perform scRNA-seq on mature human hypertrophic scars and developing scars in mice to identify the serine proteases dipeptidyl-peptidase 4 and urokinase as key molecules in this process.
- Vera Vorstandlechner
- , Maria Laggner
- & Michael Mildner
-
Article
| Open Access The impact of the timely birth dose vaccine on the global elimination of hepatitis B
The impact of the timely birth dose vaccine on the global elimination of hepatitis B
The timely hepatitis B birth dose vaccination is recommended for all new-borns by the WHO, but coverage is inconsistent. Here, the authors model the impact of scaling-up coverage in 110 low and middle income countries and assess how it may be affected by delays for example caused by the COVID-19 pandemic.
- Margaret J. de Villiers
- , Shevanthi Nayagam
- & Timothy B. Hallett
-
Article
| Open Access Quantifying previous SARS-CoV-2 infection through mixture modelling of antibody levels
Quantifying previous SARS-CoV-2 infection through mixture modelling of antibody levels
The proportion of a population that has previously been infected by a pathogen is typically estimated using antibody thresholds adjusted for sensitivity and specificity. Here, the authors present a model-based alternative to threshold methods which accounts for antibody waning and other sources of spectrum bias.
- C. Bottomley
- , M. Otiende
- & J. A. G. Scott
-
Article
| Open Access Precise measurements of chromatin diffusion dynamics by modeling using Gaussian processes
Precise measurements of chromatin diffusion dynamics by modeling using Gaussian processes
Although much effort has been devoted to determine the 3D structure of chromatin, there is a need for new experimental and computational methods. Here the authors present GP-FBM to extract chromatin diffusion parameters with high precision and apply it to live-imaging of embryonic stem cells, revealing that the diffusive properties of mitotic and interphase chromatin do not differ significantly.
- Guilherme M. Oliveira
- , Attila Oravecz
- & Nacho Molina
-
Article
| Open Access Designing the bioproduction of Martian rocket propellant via a biotechnology-enabled in situ resource utilization strategy
Designing the bioproduction of Martian rocket propellant via a biotechnology-enabled in situ resource utilization strategy
Returning from Mars to Earth requires propellant. The authors propose a biotechnology-enabled in situ resource utilization (bioISRU) process to produce a Mars specific rocket propellant, 2,3-butanediol, using cyanobacteria and engineered E. coli, with lower payload mass and energy usage compared to chemical ISRU strategies.
- Nicholas S. Kruyer
- , Matthew J. Realff
- & Pamela Peralta-Yahya
-
Article
| Open Access Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Deconvolution methods reveal individual cell types in complex tissues profiled by bulk methods. Here the authors present a Bayesian deconvolution method that outperforms existing methods when benchmarked on >700 datasets, especially in estimating cell-type-specific gene expression profiles.
- Bárbara Andrade Barbosa
- , Saskia D. van Asten
- & Yongsoo Kim
-
Article
| Open Access GproDIA enables data-independent acquisition glycoproteomics with comprehensive statistical control
GproDIA enables data-independent acquisition glycoproteomics with comprehensive statistical control
Data independent acquisition (DIA) proteomics provides deep coverage and high quantitative accuracy, but is not yet well established in glycoproteomics. Here, the authors develop a DIA-based glycoproteomics workflow with stringent statistical controls to enable accurate glycopeptide identification.
- Yi Yang
- , Guoquan Yan
- & Liang Qiao
-
Article
| Open Access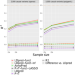 Incorporating functional priors improves polygenic prediction accuracy in UK Biobank and 23andMe data sets
Incorporating functional priors improves polygenic prediction accuracy in UK Biobank and 23andMe data sets
Incorporating functional information has shown promise for improving polygenic risk prediction of complex traits. Here, the authors describe polygenic prediction method LDpred-funct, and demonstrate its utility across 21 heritable traits in the UK Biobank.
- Carla Márquez-Luna
- , Steven Gazal
- & Alkes L. Price
-
Article
| Open Access Single cell T cell landscape and T cell receptor repertoire profiling of AML in context of PD-1 blockade therapy
Single cell T cell landscape and T cell receptor repertoire profiling of AML in context of PD-1 blockade therapy
The response rate of relapsed/refractory acute myeloid leukemia patients to PD-1 checkpoint blockade is low and unpredictable. Authors here show by single cell RNA sequencing, T cell receptor profiling and genomic analysis that the phenotypes and repertoire of CD8 + T cells and loss of chromosome 7/7q are important determinants of response.
- Hussein A. Abbas
- , Dapeng Hao
- & Andrew Futreal
-
Article
| Open Access Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
The true number of infections from SARS-Cov-2 is unknown and believed to exceed the reported numbers by several fold. National testing policies, in particular, can strongly affect the proportion of undetected cases. Here, the authors propose a method that reconstructs incidence profiles within minutes, solely from publicly available, time-stamped viral genomes.
- Maureen Rebecca Smith
- , Maria Trofimova
- & Max von Kleist
-
Article
| Open Access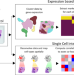 Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions
Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions
While transcriptomics have enhanced our understanding for cancer, spatial transcriptomics enable the characterisation of cellular interactions. Here, the authors integrate single cell data with spatial information for HER2 + tumours and develop tools for the prediction of interactions between tumour-infiltrating cells.
- Alma Andersson
- , Ludvig Larsson
- & Joakim Lundeberg
-
Article
| Open Access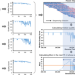 Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Current methods to identify G-quadruplex structures in DNA require specialized protocols and multiple rounds of sequencing. Here, the authors develop a method to detect G-quadruplex structures in DNA based on fluctuations in sequencing quality in a standard sequencing experiment.
- Jing Tu
- , Mengqin Duan
- & Zuhong Lu
-
Article
| Open Access Genome-wide detection of cytosine methylations in plant from Nanopore data using deep learning
Genome-wide detection of cytosine methylations in plant from Nanopore data using deep learning
Existing methods cannot profile genome-wide cytosine DNA methylations (5mCs) in all three contexts with acceptable accuracy. Here, the authors develop a deep learning tool to detect genome-wide 5mCs of all three contexts in plants with high accuracy from Nanopore reads.
- Peng Ni
- , Neng Huang
- & Jianxin Wang
-
Article
| Open Access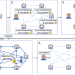 Truly privacy-preserving federated analytics for precision medicine with multiparty homomorphic encryption
Truly privacy-preserving federated analytics for precision medicine with multiparty homomorphic encryption
Existing approaches to sharing of distributed medical data either provide only limited protection of patients’ privacy or sacrifice the accuracy of results. Here, the authors propose a federated analytics system, based on multiparty homomorphic encryption (MHE), to overcome these issues.
- David Froelicher
- , Juan R. Troncoso-Pastoriza
- & Jean-Pierre Hubaux
-
Article
| Open Access Nationwide rollout reveals efficacy of epidemic control through digital contact tracing
Nationwide rollout reveals efficacy of epidemic control through digital contact tracing
The effectiveness of digital contact tracing for COVID-19 control remains uncertain. Here, the authors use data from the Smittestopp app, used in Norway in spring 2020, and estimate that 80% of nearby devices were detected by the app, and at least 11% of close contacts were not visible to manual contact tracing.
- Ahmed Elmokashfi
- , Joakim Sundnes
- & Olav Lysne
-
Article
| Open Access A frequency-amplitude coordinator and its optimal energy consumption for biological oscillators
A frequency-amplitude coordinator and its optimal energy consumption for biological oscillators
Calibrating both anomalous frequency and amplitude of biorhythm prevents physiological dysfunctions or diseases. Here, the authors propose a universal approach to design a frequency-amplitude coordinator rigorously via dynamical systems tools.
- Bo-Wei Qin
- , Lei Zhao
- & Wei Lin
-
Article
| Open Access ClusterMap for multi-scale clustering analysis of spatial gene expression
ClusterMap for multi-scale clustering analysis of spatial gene expression
In situ transcriptomics maps RNA expression patterns across intact tissues taking our understanding of gene expression to a new level. Here, the authors present a computational method that uncovers gene expression, cell niche, and tissue region patterns from 2D and 3D spatial transcriptomics.
- Yichun He
- , Xin Tang
- & Xiao Wang
-
Article
| Open Access Annotation-efficient deep learning for automatic medical image segmentation
Annotation-efficient deep learning for automatic medical image segmentation
Existing high-performance deep learning methods typically rely on large training datasets with high-quality manual annotations, which are difficult to obtain in many clinical applications. Here, the authors introduce an open-source framework to handle imperfect training datasets.
- Shanshan Wang
- , Cheng Li
- & Hairong Zheng
-
Article
| Open Access Efficient and precise single-cell reference atlas mapping with Symphony
Efficient and precise single-cell reference atlas mapping with Symphony
The number of single-cell RNA-seq datasets generated is increasing rapidly, making methods that map cell types to well-curated references increasingly important. Here, the authors propose an accurate method for mapping single cells onto a reference atlas in seconds.
- Joyce B. Kang
- , Aparna Nathan
- & Soumya Raychaudhuri
-
Article
| Open Access DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
DUBStepR is a scalable correlation-based feature selection method for accurately clustering single-cell data
Cell-type-specific genes are often strongly correlated in expression - an informative yet underexplored property of single-cell data. Here, the authors leverage gene expression correlations to develop DUBStepR, a feature selection method for accurately clustering single-cell data.
- Bobby Ranjan
- , Wenjie Sun
- & Shyam Prabhakar
-
Perspective
| Open Access A proteomics sample metadata representation for multiomics integration and big data analysis
A proteomics sample metadata representation for multiomics integration and big data analysis
The number of publicly available proteomics datasets is growing rapidly, but a standardized approach for describing the associated metadata is lacking. Here, the authors propose a format and a software pipeline to present and validate metadata, and integrate them into ProteomeXchange repositories.
- Chengxin Dai
- , Anja Füllgrabe
- & Yasset Perez-Riverol
-
Article
| Open Access Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Identifying associations of rare variants with disease is challenging due to small effect sizes, technical artefacts and population structure heterogeneity. Here, the authors present RV-EXCALIBER, a method that uses large summary-level exome data to robustly calibrate rare variant burden.
- Ricky Lali
- , Michael Chong
- & Guillaume Paré
-
Article
| Open Access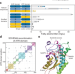 Machine learning-guided acyl-ACP reductase engineering for improved in vivo fatty alcohol production
Machine learning-guided acyl-ACP reductase engineering for improved in vivo fatty alcohol production
Fatty acyl reductases (FARs) are critical enzymes in the biosynthesis of fatty alcohols and have the ability to directly acces acyl-ACP substrates. Here, authors couple machine learning-based protein engineering framework with gene shuffling to optimize a FAR for the activity on acyl-ACP and improve fatty alcohol production.
- Jonathan C. Greenhalgh
- , Sarah A. Fahlberg
- & Philip A. Romero
-
Article
| Open Access Orchestrating and sharing large multimodal data for transparent and reproducible research
Orchestrating and sharing large multimodal data for transparent and reproducible research
It is no secret that a significant part of scientific research is difficult to reproduce. Here, the authors present a cloud-computing platform called ORCESTRA that facilitates reproducible processing of multimodal biomedical data using customizable pipelines and well-documented data objects.
- Anthony Mammoliti
- , Petr Smirnov
- & Benjamin Haibe-Kains
-
Article
| Open Access Efficient generative modeling of protein sequences using simple autoregressive models
Efficient generative modeling of protein sequences using simple autoregressive models
Deep learning is a powerful tool for the design of novel protein sequences, yet can be computationally very inefficient. Here the authors propose using simple forecasting models to efficiently generate a large number of novel protein structures.
- Jeanne Trinquier
- , Guido Uguzzoni
- & Martin Weigt
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening
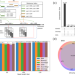
 Fast alignment and preprocessing of chromatin profiles with Chromap
Fast alignment and preprocessing of chromatin profiles with Chromap