Featured
-
-
Article
| Open Access Integrating gene expression and clinical data to identify drug repurposing candidates for hyperlipidemia and hypertension
Integrating gene expression and clinical data to identify drug repurposing candidates for hyperlipidemia and hypertension
Prioritizing drug repurposing candidates for downstream studies remains challenging. Here, the authors present a high-throughput approach to identify and validate drug repurposing candidates, integrating human gene expression, drug perturbation, and clinical data from publicly available resources.
- Patrick Wu
- , QiPing Feng
- & Wei-Qi Wei
-
Article
| Open Access Helical structure motifs made searchable for functional peptide design
Helical structure motifs made searchable for functional peptide design
Here, we present TP-DB; a pattern-based search engine based on 1.67 million helices from the Protein Database (PDB). We demonstrate the utility of TP-DB in identifying microbe-specific antigens, as well as the design of antimicrobial peptides and Protein-protein interaction blockers.
- Cheng-Yu Tsai
- , Emmanuel Oluwatobi Salawu
- & Lee-Wei Yang
-
Article
| Open Access Prediction of biomarkers and therapeutic combinations for anti-PD-1 immunotherapy using the global gene network association
Prediction of biomarkers and therapeutic combinations for anti-PD-1 immunotherapy using the global gene network association
A lot of cancer patients are not responsive to anti-PD1 therapy. Here, the authors develop a network approach to identify genes, pathways and potential therapeutic combinations and develop an MHC-I gene immunoscore associated with tumour response to anti-PD1.
- Chia-Chin Wu
- , Y. Alan Wang
- & P. Andrew Futreal
-
Article
| Open Access Mini-batch optimization enables training of ODE models on large-scale datasets
Mini-batch optimization enables training of ODE models on large-scale datasets
Ordinary differential equation (ODE) models are widely used to understand multiple processes. Here the authors show how the concept of mini-batch optimization can be transferred from the field of Deep Learning to ODE modelling.
- Paul Stapor
- , Leonard Schmiester
- & Jan Hasenauer
-
Article
| Open Access Harnessing protein folding neural networks for peptide–protein docking
Harnessing protein folding neural networks for peptide–protein docking
AlphaFold2 has originally been developed to provide highly accurate predictions of protein monomer structures. Here, the authors present a simple adaptation of AlphaFold2 that enables structural modeling of peptide–protein complexes, and explore the underlying mechanisms and limitations of this approach.
- Tomer Tsaban
- , Julia K. Varga
- & Ora Schueler-Furman
-
Article
| Open Access Polyply; a python suite for facilitating simulations of macromolecules and nanomaterials
Polyply; a python suite for facilitating simulations of macromolecules and nanomaterials
To facilitate the rational design of (nano)-materials and biomacromolecules by MD simulations, the authors present the polyply suite, featuring a graph matching algorithm and a random walk protocol for generating multi-scale polymeric topologies and initial coordinates.
- Fabian Grünewald
- , Riccardo Alessandri
- & Siewert J. Marrink
-
Article
| Open Access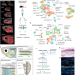 Lifelong single-cell profiling of cranial neural crest diversification in zebrafish
Lifelong single-cell profiling of cranial neural crest diversification in zebrafish
Cranial neural crest generates a wide diversity of cell types. Here the authors perform single-cell profiling of neural crest to identify key enhancers and transcription factors for cell fate competency, thus revealing progressive acquisition of fate potential.
- Peter Fabian
- , Kuo-Chang Tseng
- & J. Gage Crump
-
Article
| Open Access Association of mutation signature effectuating processes with mutation hotspots in driver genes and non-coding regions
Association of mutation signature effectuating processes with mutation hotspots in driver genes and non-coding regions
In cancer, associations between mutational signatures and driver mutations have been proposed but not fully explored. Here, the authors develop sigDriver to find associations between mutational signatures and mutation hotspots in order to predict coding and non-coding driver mutations in pan-cancer genomics data.
- John K. L. Wong
- , Christian Aichmüller
- & Marc Zapatka
-
Article
| Open Access Phylogenetically and functionally diverse microorganisms reside under the Ross Ice Shelf
Phylogenetically and functionally diverse microorganisms reside under the Ross Ice Shelf
The Ross Ice Shelf is the most extensive ice shelf of Antarctica and isolates the underlying ocean from sunlight. Here the authors use multi-omics to unravel the phylogenetic and functional diversity of microbial life in this ecosystem.
- Clara Martínez-Pérez
- , Chris Greening
- & Federico Baltar
-
Article
| Open Access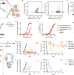 Maximizing response to intratumoral immunotherapy in mice by tuning local retention
Maximizing response to intratumoral immunotherapy in mice by tuning local retention
Understanding the pharmacokinetics of locally-injected drugs could aid in the design of immunotherapies to maximize their therapeutic effect. Here, by evaluating different IL-2 fusion proteins, the authors show that molecular weight and matrix binding affect anti-tumor immune response and report a pharmacokinetic framework to predict response to intratumoral IL-2 therapy.
- Noor Momin
- , Joseph R. Palmeri
- & K. Dane Wittrup
-
Article
| Open Access Topographic mapping of the glioblastoma proteome reveals a triple-axis model of intra-tumoral heterogeneity
Topographic mapping of the glioblastoma proteome reveals a triple-axis model of intra-tumoral heterogeneity
Gioblastoma tumours consist of different niches defined by histology. Here, the authors use proteomics and machine learning to assign protein expression programs to these niches, and reveal that KRAS and hypoxia are associated with drug resistance.
- K. H. Brian Lam
- , Alberto J. Leon
- & Phedias Diamandis
-
Article
| Open Access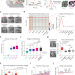 Face detection in untrained deep neural networks
Face detection in untrained deep neural networks
Face-selective neurons are observed in the primate visual pathway and are considered as the basis of face detection in the brain. Here, using a hierarchical deep neural network model of the ventral visual stream, the authors suggest that face selectivity arises in the complete absence of training.
- Seungdae Baek
- , Min Song
- & Se-Bum Paik
-
Article
| Open Access Simultaneous estimation of bi-directional causal effects and heritable confounding from GWAS summary statistics
Simultaneous estimation of bi-directional causal effects and heritable confounding from GWAS summary statistics
Mendelian Randomization approaches are being increasingly refined, but certain statistical limitations hinder their application to GWAS. Here, the authors propose a new Mendelian Randomization method to estimate bi- directional causal effects and explicitly account for heritable confounding.
- Liza Darrous
- , Ninon Mounier
- & Zoltán Kutalik
-
Article
| Open Access A machine and human reader study on AI diagnosis model safety under attacks of adversarial images
A machine and human reader study on AI diagnosis model safety under attacks of adversarial images
While active efforts are advancing medical AI model development and clinical translation, safety issues of medical AI models have emerged. Here, the authors investigate the effects on an AI model and on human experts of potential fake/adversarial images for breast cancer diagnosis.
- Qianwei Zhou
- , Margarita Zuley
- & Shandong Wu
-
Article
| Open Access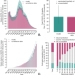 The effect of COVID-19 vaccination in Italy and perspectives for living with the virus
The effect of COVID-19 vaccination in Italy and perspectives for living with the virus
Vaccination campaigns against COVID-19 are allowing the progressive release of physical distancing restrictions in many countries. Here, the authors assess the impact of the vaccination program in Italy and evaluate possible prospects for reopening the society while at the same time keeping COVID-19 under control.
- Valentina Marziano
- , Giorgio Guzzetta
- & Stefano Merler
-
Article
| Open Access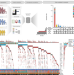 Clonal architecture predicts clinical outcomes and drug sensitivity in acute myeloid leukemia
Clonal architecture predicts clinical outcomes and drug sensitivity in acute myeloid leukemia
Individual studies have been underpowered to draw clear associations between clonal heterogeneity and response to therapy in acute myeloid leukemia (AML). Here, the authors aggregate multiple AML cohorts and are able to correlate the clonal abundance of somatic mutations with clinical outcomes and drug sensitivity.
- Brooks A. Benard
- , Logan B. Leak
- & Ravindra Majeti
-
Article
| Open Access Emulator-based Bayesian optimization for efficient multi-objective calibration of an individual-based model of malaria
Emulator-based Bayesian optimization for efficient multi-objective calibration of an individual-based model of malaria
Individual-based models have become important tools in the global battle against infectious diseases, yet model complexity can make calibration challenging. Here, the authors propose a Bayesian optimization framework to calibrate a complex malaria transmission simulator.
- Theresa Reiker
- , Monica Golumbeanu
- & Melissa A. Penny
-
Article
| Open Access RNA modifications detection by comparative Nanopore direct RNA sequencing
RNA modifications detection by comparative Nanopore direct RNA sequencing
Nanopore direct RNA Sequencing data contain information about the presence of RNA modifications, but their detection poses substantial challenges. Here the authors introduce Nanocompore, a new methodology for modification detection from Nanopore data.
- Adrien Leger
- , Paulo P. Amaral
- & Tony Kouzarides
-
Article
| Open Access Improved analyses of GWAS summary statistics by reducing data heterogeneity and errors
Improved analyses of GWAS summary statistics by reducing data heterogeneity and errors
Analyses of summary statistics from GWAS are subject to biases due to errors in the discovery GWAS or linkage disequilibrium reference data set or heterogeneity between data sets. Here, the authors propose a quality control method to be added to analysis of GWAS summary data that can reduce such biases.
- Wenhan Chen
- , Yang Wu
- & Jian Yang
-
Article
| Open Access Sliding of HIV-1 reverse transcriptase over DNA creates a transient P pocket – targeting P-pocket by fragment screening
Sliding of HIV-1 reverse transcriptase over DNA creates a transient P pocket – targeting P-pocket by fragment screening
Here the authors observe a transient P-pocket created when HIV reverse transcriptase slides over DNA substrate, identify fragments targeting this pocket, and develop a cryo-EM platform for lead optimization.
- Abhimanyu K. Singh
- , Sergio E. Martinez
- & Kalyan Das
-
Article
| Open Access Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Idiopathic pulmonary arterial hypertension is a rare and fatal disease with a heterogeneous treatment response. Here the authors show that unsupervised machine learning of whole blood transcriptomes from 359 patients with idiopathic pulmonary arterial hypertension identifies 3 subgroups (endophenotypes) that improve risk stratification and provide new molecular insights.
- Sokratis Kariotis
- , Emmanuel Jammeh
- & Richard C. Trembath
-
Article
| Open Access DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
The authors present DeepRank, a deep learning framework for the data mining of large sets of 3D protein-protein interfaces (PPI). They use DeepRank to address two challenges in structural biology: distinguishing biological versus crystallographic PPIs in crystal structures, and secondly the ranking of docking models.
- Nicolas Renaud
- , Cunliang Geng
- & Li C. Xue
-
Article
| Open Access Higher order genetic interactions switch cancer genes from two-hit to one-hit drivers
Higher order genetic interactions switch cancer genes from two-hit to one-hit drivers
In the classic two-hit model, both alleles of a tumour suppressor gene need to be inactivated in order to promote cancer. Here, the authors challenge this model, finding that many cancer genes can be either one-hit or two-hit drivers depending on the context and other mutations in a tumor.
- Solip Park
- , Fran Supek
- & Ben Lehner
-
Article
| Open Access Probabilistic inference of the genetic architecture underlying functional enrichment of complex traits
Probabilistic inference of the genetic architecture underlying functional enrichment of complex traits
Improving inference in large-scale genetic data linked to electronic medical record data requires the development of novel computationally efficient regression methods. Here, the authors develop a Bayesian approach for association analyses to improve SNP-heritability estimation, discovery, fine-mapping and genomic prediction.
- Marion Patxot
- , Daniel Trejo Banos
- & Matthew R. Robinson
-
Comment
| Open AccessComputational tools for genomic data de-identification: facilitating data protection law compliance
In this opinion piece, we discuss why computational tools to limit the identifiability of genomic data are a promising avenue for privacy-preservation and legal compliance. Even where these technologies do not eliminate all residual risk of individual identification, the law may still consider such data anonymised.
- Alexander Bernier
- , Hanshi Liu
- & Bartha Maria Knoppers
-
Article
| Open Access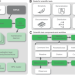 Ensuring scientific reproducibility in bio-macromolecular modeling via extensive, automated benchmarks
Ensuring scientific reproducibility in bio-macromolecular modeling via extensive, automated benchmarks
Computational methods are becoming an increasingly important part of biological research. Using the Rosetta framework as an example, the authors demonstrate how community-driven development of computational methods can be done in a reproducible and reliable fashion.
- Julia Koehler Leman
- , Sergey Lyskov
- & Richard Bonneau
-
Article
| Open Access High-throughput mediation analysis of human proteome and metabolome identifies mediators of post-bariatric surgical diabetes control
High-throughput mediation analysis of human proteome and metabolome identifies mediators of post-bariatric surgical diabetes control
Factors underlying the effects of gastric bypass surgery on glucose homeostasis are incompletely understood. Here the authors developed and applied high-throughput mediation analysis to identify proteome/metabolome mediators of improved glucose homeostasis after to gastric bypass surgery, and report that improved glycemia was mediated by the growth hormone receptor.
- Jonathan M. Dreyfuss
- , Yixing Yuchi
- & Mary Elizabeth Patti
-
Article
| Open Access An atlas of protein turnover rates in mouse tissues
An atlas of protein turnover rates in mouse tissues
Protein turnover underpins biology but is challenging to measure in vivo across the entire proteome. Here, the authors provide a comprehensive resource of protein turnover in mouse tissues and develop a visualization platform to analyze these data.
- Zach Rolfs
- , Brian L. Frey
- & Nathan V. Welham
-
Article
| Open Access Whole-genome analysis of Nigerian patients with breast cancer reveals ethnic-driven somatic evolution and distinct genomic subtypes
Whole-genome analysis of Nigerian patients with breast cancer reveals ethnic-driven somatic evolution and distinct genomic subtypes
Breast cancer heterogeneity and tumour evolutionary trajectories remain largely unknown among women of African ancestry. Here, the authors perform whole genome and transcriptome sequencing of Nigerian breast cancer patients and identify unique evolutionary phenomena.
- Naser Ansari-Pour
- , Yonglan Zheng
- & Olufunmilayo I. Olopade
-
Article
| Open Access Cryo-EM snapshots of a native lysate provide structural insights into a metabolon-embedded transacetylase reaction
Cryo-EM snapshots of a native lysate provide structural insights into a metabolon-embedded transacetylase reaction
How is acetyl-CoA produced in the context of the endogenous, eukaryotic pyruvate dehydrogenase complex metabolon? Here the authors dissect the embedded transacetylase reaction through biochemical, cryo-EM, HADDOCKing and molecular dynamics methods.
- Christian Tüting
- , Fotis L. Kyrilis
- & Panagiotis L. Kastritis
-
Article
| Open Access Network medicine for disease module identification and drug repurposing with the NeDRex platform
Network medicine for disease module identification and drug repurposing with the NeDRex platform
There is an unmet need for adaptable tools allowing biomedical researchers to employ network-based drug repurposing approaches for their individual use cases. Here, the authors close this gap with NeDRex, an integrative and interactive platform.
- Sepideh Sadegh
- , James Skelton
- & Tim Kacprowski
-
Article
| Open Access SARS-CoV-2 transmission across age groups in France and implications for control
SARS-CoV-2 transmission across age groups in France and implications for control
In this study, Tran Kiem et al. examine the contribution of different age groups to COVID-19 transmission. Using data from the French epidemic in summer 2020, they report that while individuals aged 80 years and older are more at risk, pandemic control in the absence of vaccines required measures targeted at all age groups.
- Cécile Tran Kiem
- , Paolo Bosetti
- & Simon Cauchemez
-
Article
| Open Access A benchmark study of simulation methods for single-cell RNA sequencing data
A benchmark study of simulation methods for single-cell RNA sequencing data
Simulation is useful for developing and evaluating computational methods. Here, the authors develop a comprehensive evaluation framework, SimBench, to benchmark Single-cell RNA-seq simulation methods through a diverse collection of experimental datasets.
- Yue Cao
- , Pengyi Yang
- & Jean Yee Hwa Yang
-
Article
| Open Access scCODA is a Bayesian model for compositional single-cell data analysis
scCODA is a Bayesian model for compositional single-cell data analysis
Imbalance and loss of cell types is a hallmark in many diseases. Still, quantifying compositional changes in scRNAseq data remains challenging. Here the authors present scCODA, a Bayesian model to assess cell type compositions in scRNA-seq data.
- M. Büttner
- , J. Ostner
- & B. Schubert
-
Article
| Open Access Ecological memory preserves phage resistance mechanisms in bacteria
Ecological memory preserves phage resistance mechanisms in bacteria
One might think that complete extinction of a virulent pathogen is the most effective way of saving a population. For a bacteria-phage system, Skanata and Kussell show that sustaining a minimum pathogen level is actually favorable to prevent a complete loss of immunity in the long run.
- Antun Skanata
- & Edo Kussell
-
Article
| Open Access Bioinformatic and cell-based tools for pooled CRISPR knockout screening in mosquitos
Bioinformatic and cell-based tools for pooled CRISPR knockout screening in mosquitos
Forward genetic approaches such as CRISPR screens are powerful ways to identify essential genes and those that influence host-pathogen interactions. Here the authors design a bioinformatics portal for sgRNA design and a recombination-mediated cassette system for delivery into mosquito cell lines.
- Raghuvir Viswanatha
- , Enzo Mameli
- & Norbert Perrimon
-
Article
| Open Access A unified drug–target interaction prediction framework based on knowledge graph and recommendation system
A unified drug–target interaction prediction framework based on knowledge graph and recommendation system
Prediction of drug-target interactions (DTI) plays a vital role in drug development through applications in various areas, such as virtual screening for lead discovery, drug repurposing and identification of potential drug side effects. Here, the authors develop a unified framework for DTI prediction by combining a knowledge graph and a recommendation system.
- Qing Ye
- , Chang-Yu Hsieh
- & Tingjun Hou
-
Article
| Open Access Machine learning of genomic features in organotropic metastases stratifies progression risk of primary tumors
Machine learning of genomic features in organotropic metastases stratifies progression risk of primary tumors
The location and timing of metastasis are still fundamentally unpredictable. Here the authors present the Metastatic Network model, a machine learning framework that integrates clinical data and DNA alterations to predict the risk of metastasis to specific organs as well as clinical outcomes
- Biaobin Jiang
- , Quanhua Mu
- & Jiguang Wang
-
Article
| Open Access DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
The coverage and throughput of data-independent acquisition (DIA)-based phosphoproteomics is limited by its dependence on experimental spectral libraries. Here the authors develop a DIA workflow based on in silico spectral libraries generated by a novel deep neural network to expand phosphoproteome coverage.
- Ronghui Lou
- , Weizhen Liu
- & Wenqing Shui
-
Article
| Open Access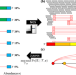 Jumper enables discontinuous transcript assembly in coronaviruses
Jumper enables discontinuous transcript assembly in coronaviruses
@melkebir @psashittal et al. develop a graph-based method for the assembly of discontinuous transcripts produced in Coronaviruses and other Nidovirales, enabling the discovery of transcriptional changes missed by existing methods.
- Palash Sashittal
- , Chuanyi Zhang
- & Mohammed El-Kebir
-
Article
| Open Access Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Obtaining accurate variant calls from multiple displacement amplified single cell DNA sequencing data needs dedicated models that account for amplification bias and copy errors. Here, the authors describe ProSolo, a model for calling single nucleotide variants with control over the false discovery rate.
- David Lähnemann
- , Johannes Köster
- & Alexander Schönhuth
-
Article
| Open Access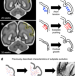 A model of tension-induced fiber growth predicts white matter organization during brain folding
A model of tension-induced fiber growth predicts white matter organization during brain folding
Associations have been established between brain folding and white matter connectivity. Here the authors show that axon elongation, in response to mechanical stresses during cortical expansion and folding, may be sufficient to induce tissue remodeling consistent with white matter organization.
- Kara E. Garcia
- , Xiaojie Wang
- & Christopher D. Kroenke
-
Article
| Open Access Age-seroprevalence curves for the multi-strain structure of influenza A virus
Age-seroprevalence curves for the multi-strain structure of influenza A virus
Multi-strain pathogens, such as influenza, present challenges for interpretation of seroprevalence data as estimates may vary by strain. Here, the authors develop a method for estimating age-specific seroprevalence based on principal components analysis and apply it to influenza data from Vietnam.
- Dao Nguyen Vinh
- , Nguyen Thi Duy Nhat
- & Maciej F. Boni
-
Article
| Open Access Model-based assessment of Chikungunya and O’nyong-nyong virus circulation in Mali in a serological cross-reactivity context
Model-based assessment of Chikungunya and O’nyong-nyong virus circulation in Mali in a serological cross-reactivity context
O’nyong nyong and Chikungunya virus are arboviruses present in Africa but their prevalence is unknown, partly due to high antibody cross-reactivity with one another. Here, the authors develop a statistical model that accounts for cross-reactivity to characterise circulation of both viruses from seroprevalence surveys.
- Nathanaël Hozé
- , Issa Diarra
- & Simon Cauchemez
-
Article
| Open Access Mechanical forces drive a reorientation cascade leading to biofilm self-patterning
Mechanical forces drive a reorientation cascade leading to biofilm self-patterning
Bacterial biofilms exhibit complex spatiotemporal pattern formation. Here the authors report a collective cell reorientation cascade in growing Vibrio cholerae biofilms that leads to a differentially ordered, spatiotemporally coupled core-rim structure.
- Japinder Nijjer
- , Changhao Li
- & Jing Yan
-
Article
| Open Access scPower accelerates and optimizes the design of multi-sample single cell transcriptomic studies
scPower accelerates and optimizes the design of multi-sample single cell transcriptomic studies
scRNASeq data is revolutionizing our understanding of biological systems, but is still expensive to generate. Here, the authors present a statistical framework that facilitates informed multi-sample experimental design to reduce unnecessary costs and maximize the utility of the generated data.
- Katharina T. Schmid
- , Barbara Höllbacher
- & Matthias Heinig
-
Article
| Open Access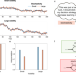 A model for learning based on the joint estimation of stochasticity and volatility
A model for learning based on the joint estimation of stochasticity and volatility
Human learning depends on opposing effects of two noise factors: volatility and stochasticity. Here the authors present a model of learning that shows how and why joint estimation of these factors is important for understanding healthy and pathological learning.
- Payam Piray
- & Nathaniel D. Daw
-
Article
| Open Access Mesmerize is a dynamically adaptable user-friendly analysis platform for 2D and 3D calcium imaging data
Mesmerize is a dynamically adaptable user-friendly analysis platform for 2D and 3D calcium imaging data
Calcium imaging is valuable for understanding neuro and cell biology, but is challenging to analyze, organize, and access. Here, the authors present an efficient, expandable and user-friendly platform, which encapsulates the entire analysis process all to way to interactive visualizations.
- Kushal Kolar
- , Daniel Dondorp
- & Marios Chatzigeorgiou
-
Article
| Open Access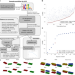 Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana
Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana
Methods to predict transcription factor binding sites typically focus on sequence motifs without considering DNA shape. Here the authors use a random forest machine learning approach that incorporates DNA shape and improves binding site prediction for Arabidopsis thaliana transcription factors.
- Janik Sielemann
- , Donat Wulf
- & Andrea Bräutigam
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Rapid antigen testing as a reactive response to surges in nosocomial SARS-CoV-2 outbreak risk
Rapid antigen testing as a reactive response to surges in nosocomial SARS-CoV-2 outbreak risk