Abstract
Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype lacking expression of estrogen and progesterone receptors (ER/PR) and HER2, thus limiting therapy options. We hypothesized that meta-analysis of TNBC gene expression profiles would illuminate mechanisms underlying the aggressive nature of this disease and identify therapeutic targets. Meta-analysis in the Oncomine database identified 206 genes that were recurrently deregulated in TNBC compared with non-TNBC and in tumors that metastasized or led to death within 5 years. This ‘aggressiveness gene list’ was enriched for two core functions/metagenes: chromosomal instability (CIN) and ER signaling metagenes. We calculated an ‘aggressiveness score’ as the ratio of the CIN metagene to the ER metagene, which identified aggressive tumors in breast cancer data sets regardless of subtype or other clinico-pathological indicators. A score calculated from six genes from the CIN metagene and two genes from the ER metagene recapitulated the aggressiveness score. By multivariate survival analysis, we show that our aggressiveness scores (from 206 genes or the 8 representative genes) outperformed several published prognostic signatures. Small interfering RNA screen revealed that the CIN metagene holds therapeutic targets against TNBC. Particularly, the inhibition of TTK significantly reduced the survival of TNBC cells and synergized with docetaxel in vitro. Importantly, mitosis-independent expression of TTK protein was associated with aggressive subgroups, poor survival and further stratified outcome within grade 3, lymph node-positive, HER2-positive and TNBC patients. In conclusion, we identified the core components of CIN and ER metagenes that identify aggressive breast tumors and have therapeutic potential in TNBC and aggressive breast tumors. Prognostication from these metagenes at the mRNA level was limited to ER-positive tumors. However, we provide evidence that mitosis-independent expression of TTK protein was prognostic in TNBC and other aggressive breast cancer subgroups, suggesting that protection of CIN/aneuploidy drives aggressiveness and treatment resistance.
Similar content being viewed by others
Introduction
Estrogen and progesterone receptors (ER and PR) and HER2 are standard biomarkers used in clinical practice to aid the histopathological classification of breast cancer and management decisions. Hormone receptor- and HER2-positive tumors benefit from tamoxifen and anti-HER2 therapies, respectively. On the other hand, there are currently no targeted drug therapies for management of triple-negative breast cancer (TNBC), which lacks the expression of hormone receptors and HER2. TNBCs are more sensitive to chemotherapy than hormone receptor-positive tumors, because they are generally more proliferative, and pathological complete responses after chemotherapy are more likely in TNBC than in non-TNBC.1, 2 Paradoxically, TNBC is associated with poorer survival than non-TNBC, owing to more frequent relapse in TNBC patients with residual disease.1, 2 Only 31% of TNBC patients experience pathological complete responses after chemotherapy,3 emphasizing the need for targeted therapies.
Transcriptome profiling has been used to dissect the heterogeneity of breast cancer into five intrinsic ‘PAM50’ subtypes: Luminal A, Luminal B, Basal-like, HER2 and normal-like subtypes that relate to clinical outcomes.4, 5, 6, 7, 8 Several gene signatures have been developed to predict outcome or response to treatment, including MammaPrint,9 OncotypeDx10, 11 and Theros.12, 13, 14, 15 These commercial signatures rely on models that select geneson the basis of clinical phenotypes such as tumor response or survival time. Notwithstanding their clinical utilities, these models fail to identify core biological mechanisms for the phenotypes of interest. Recently, an approach based on biological function-driven gene coexpression signatures, ‘attractor metagenes’, has been applied to the prediction of survival in multiple cancers including breast cancer.16, 17 Three attractor metagenes, chromosomal instability (CIN), mesenchymal transition and lymphocyte-specific immune recruitment, were highly predictive of breast cancer survival.17 To some extent, this approach may helpclarify some previously published signatures. For example, proliferation and cell cycle signatures have been previously reported to associate with tumor grade and prognosis.15, 18 The attractor metagene approach suggests that these signatures are essentially CIN attractors enriched with genes that function at the kinetochore–microtubule interface.16
In this study, we initially performed multiple class comparisons using the Oncomine database,19 aiming to identify genes that were commonly deregulated in subgroups exemplifying aggressive clinical behavior: TNBC compared to non-TNBC and normal breast and tumors associated with distant metastasis and/or death compared to their respective counterparts. This analysis revealed a list of 206 recurrently deregulated genes that were enriched for CIN and ER metagenes. We derived an aggressiveness score based on the ratio of the CIN metagene to the ER metagene, and found that this score identified aggressive tumors in several other data sets regardless of the molecular subtype and clinico-pathological indicators. The aggressiveness score outperformed MammaPrint,9 OncotypeDx,10, 11 proliferation per cell cycle16, 20 and CIN20 signatures in multivariate Cox proportional hazards comparisons. Next, we found that depletion of proteins involved in kinetochore binding or chromosome segregation (TTK, TPX2, NDC80 and PBK) could be therapeutic and significantly reduced the survival of TNBC cell lines in vitro, particularly TTK. TTK inhibition with small-molecule inhibitor affected the survival of TNBC cell lines. We found that both TTK mRNA and protein levels associated with aggressive tumor phenotypes. Mitosis-independent expression of TTK protein was prognostic in TNBC and other aggressive breast cancer subgroups, suggesting that protection of CIN/aneuploidy drives aggressiveness and treatment resistance. Finally, we show that the combination of TTK inhibition with chemotherapy was effective in vitro in the treatment of cells that overexpress TTK, thus providing a therapeutic option for the protected CIN phenotype.
Results
Meta-analysis of gene expression profiles in TNBC
We performed a meta-analysis of published gene expression data, irrespective of platform, using the Oncomine database19 (version 4.5). We compared the expression profiles of 492 TNBC cases vs 1382 non-TNBC cases in eight data sets and found 1600 overexpressed and 1580 underexpressed genes in the TNBC cases (cutoff median P-value across the 8 data sets <1 × 10−5 from a Student’s t-test, Supplementary Figure 1). We also compared the expression profiles of primary breast cancers from 512 patients who developed metastases vs 732 patients who did not develop metastases at 5 years (7 data sets in total) to identify 500 overexpressed and 480 underexpressed genes in the metastasis cases (cutoff median P-value across the seven data sets <0.05 from a Student’s t-test, Supplementary Figure 1). Finally, we compared the expression profiles of 232 primary breast tumors from patients who died within 5 years with 879 patients who survived in seven data sets and found 500 overexpressed and 500 underexpressed genes in the poor survivors (cutoff median P-value across the seven data sets <0.05 from a Student’s t-test, Supplementary Figure 1). The union of these analyses—genes deregulated in TNBC and in tumors that metastasized or resulted in death within 5 years—generated a gene list of 305 overexpressed and 341 underexpressed genes (Supplementary Figures 2A and B). The deregulated genes from our analyses did not consider deregulation in comparison with normal breast tissue. To identify cancer-related genes, we used the METABRIC (Molecular Taxonomy of Breast Cancer International Consortium) data set21 as a validation data set. Of the 305 overexpressed and 341 underexpressed genes identified in the meta-analysis, 117 overexpressed and 89 underexpressed genes (206 genes) were deregulated in TNBC (250 cases) vs 144 adjacent normal tissue (1.5-fold-change cutoff; Supplementary Figures 2C and D).
Clinico-pathological features of the aggressiveness gene list
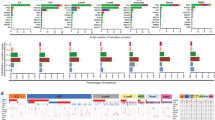
We compared the 206 genes from the above analysis, which we called the ‘aggressiveness gene list’ (Supplementary Table 1), with the recently described metagene attractors16, 17 and found that 45 of the overexpressed genes were in the CIN metagene, whereas 19 of the underexpressed genes were in the ER metagene (Supplementary Figure 3). The expression of the aggressiveness gene list was visualized in the METABRIC data set, stratified according to the histological subtypes by the GENUIS classification.22 As shown in Figure 1a, ER−/HER2− (TNBC), in comparison with adjacent normal breast tissue, showed the highest upregulation of CIN genes (red in the heat map) and downregulation of ER signaling genes (green in the heat map). Tumors of other subtypes showed a range of deregulation of these genes. To quantify these trends, we calculated the ‘aggressiveness score’ as the ratio of the CIN metagene (average of expression of CIN genes) to the ER metagene (average of expression of ER genes). The aggressiveness score was highest for ER−/HER2− (TNBC), followed by HER2+ then ER+ tumors (box plot in Figure 1). We also analyzed the aggressiveness score in the five intrinsic breast cancer subtypes predefined by the PAM50 classification8 and the ten integrative clustering subtypes defined by combined clustering of gene expression and copy number data subtypes21 (Supplementary Figure 4). The aggressiveness score was highest in the basal-like and the integrative clustering 10 subtypes, which are enriched for TNBC and have poor prognosis.
Correlation of breast cancer subtypes and the aggressiveness gene list. The METABRIC data set was visualized according to the expression of the 206 genes (Supplementary Table 1) in the aggressiveness gene list. The aggressiveness score for each tumor was calculated as the ratio of the CIN metagene (average value for CIN genes expression) to the ER metagene (average value for ER genes expression). (a) The expression of the aggressiveness gene list according to the GENIUS histological classification. The box plot shows the aggressiveness score of the histological subtypes. (b) The overall survival of patients in the METABRIC data set was analyzed according to the aggressiveness score (upper row: by quartiles; lower row: by median) in all patients, non-TNBC patients and in patients with ER+ Grade 2 tumors. The hazard ratio (HR), CI and P-value for comparisons of upper quartile vs lower quartiles (upper row) and at the dichotomy across the median (high vs low) are shown (log-rank test, GraphPad Prism). The number of patients (n) in each group is shown in brackets. The expression of the aggressiveness score according to PAM50 and intClust subtypes and survival curves for ER+ grade 3 and PAM50 subtypes according to the aggressiveness score are in Supplementary Figures 4 and 5.
Interestingly, tumors of various subtypes scored higher than the median aggressiveness score (line in box plots in Figure 1 and Supplementary Figure 4). To this end, we examined the overall survival of patients in the METABRIC data set stratified by quartiles and also dichotomized by the median of the aggressiveness score. Tumors with a high aggressiveness score had worse survival than those with a low aggressiveness score. The survival of patients with non-TNBC tumors with high aggressiveness score had poor survival that was similar to TNBC patients (Figure 1b). Among ER+ tumors, we found that a high aggressiveness score predicted poor survival in both Grade 2 (Figure 1b) and Grade 3 (Supplementary Figure 4) tumors. Tumors with a high aggressiveness score showed poor survival regardless of the PAM50 intrinsic breast cancer subtypes (Supplementary Figure 4). The PAM50 classifier was prognostic only in tumors with a low aggressiveness score (Supplementary Figure 5).
One network of direct interactions in the aggressiveness gene list associates with patient survival
We performed network analysis on the aggressiveness gene list by using the Ingenuity Pathway Analysis and found a network with direct interactions between 97 of the 206 deregulated genes (Figure 2a). To find the minimal genes that represent the aggressiveness genes and this network, the 97 genes in this network were analyzed for their correlation with the CIN or ER metagenes and overall survival in the METABRIC data-set (Supplementary Table 2). We selected genes according to the following criteria: (1) highest correlation with the metagenes (Pearson’s correlation coefficient >0.7); (2) association with overall survival (Cox proportional hazards model, P<0.001); and (3) more than two-fold deregulation with least standard deviation of expression between high and low aggressiveness score tumors. These analyses identified two genes from the ER metagene (MAPT and MYB) and six genes from the CIN metagene (MELK, MCM10, CENPA, EXO1, TTK and KIF2C). These eight genes were maintained in a directly connected network (Figure 2b). The classification of tumors (high vs low across the median) from these eight genes, again representing the ratio of CIN and ER metagenes, predicted the classification from the 206 genes with 95% sensitivity and 97% specificity by prediction of microarray (PAM) analysis (data not shown). Importantly, a high score from these eight genes identified poor survival in all patients, non-TNBC patients and ER+ Grade 2 (Figure 2c).
Network analysis of the aggressiveness gene list. (a) Ingenuity Pathway Analysis was performed using direct interactions on the 206 genes in the aggressiveness gene list (red is overexpressed and green is underexpressed). One network of high direct interactions was identified. (b) The genes in the network in A were investigated for their correlation with the aggressiveness score and overall survival (Supplementary Table 2), and eight genes (MAPT, MYB, MELK, MCM10, CENPA, EXO1, TTK and KIF2C) with the highest correlation were still connected in a direct interaction network. (c) The overall survival of patients in the METABRIC data set was analyzed according to the score from the 8 genes in C (upper row: by quartiles; lower row: by median) in all patients, non-TNBC patients and in patients with ER+ Grade 2 tumors.
Next, we explored the 8-gene score for prognosis in several molecular and histological settings in the METABRIC data set. The survival of patients with tumors with wild-type TP53 was stratified by the 8-gene score (Figure 3a). Patients with mutant TP53, which were mainly of high score, showed worse survival than those with wild-type TP53, suggesting that TP53 mutation is an independent prognostic factor. Patients with tumors with low or high expression of the proliferation marker Ki67 were stratified by the 8-gene score, suggesting that the 8-gene score is independent of proliferation (Figure 3a). We also found that the 8-gene score stratified the survival of patients from all stages of disease (Stage I—Stage III, Figure 3a). We focused on ER+ and found that, as in the case of ER+ Grade 2 tumors (Figure 2c), the 8-gene score stratified the survival of patients with ER+ Grade 3 tumors (Figure 3b). Importantly, the 8-gene score identified ER+LN− and ER+LN+ patients who had poor survival similar to ER−LN− and ER−LN+ patients, respectively (Figure 3b). High 8-gene score identified poor survival of patients with tumors of all PAM50 subtypes, and the prognostication by PAM50 classification was only evident in low 8-gene score tumors (Supplementary Figure 5).
Survival of patients stratified by the 8-gene score in the METABRIC data set. The overall survival of patients in the METABRIC data set was analyzed according to the 8-gene score in selected settings in all patients (a) or in ER-positive patients only (b). (a) TP53 mutation was compared in high vs low 8-gene score (split by the median). The expression of the proliferation marker Ki67 was divided by dichotomy across the median, and patients in each of these groups were then stratified according to their 8-gene score (split by quartiles). Disease stages (Stage I—Stage III) were stratified by the median 8-gene score. (b) ER+ Grade 3, ER+ lymph node-negative (LN−) and ER+ LN+ tumors were stratified by the quartiles.
The 8-gene aggressiveness score in multivariate survival analysis
To exclude the possibility that the aggressiveness score—calculated using the 206 genes or the 8 genes—was redundant, we performed multivariate Cox proportional hazards model analysis in the METABRIC data-set (with Illumina platform) in comparison with conventional clinical variables and current gene signatures. As detailed in Table 1, the aggressiveness scores significantly associated with patient survival when compared with conventional variables and outperformed MammaPrint,9 OncotypeDx,10, 11 proliferation per cell cycle16, 20 and CIN20 signatures. Moreover, our aggressiveness scores outperformed the CIN4 classifier,23 which was recently developed from the CIN signature.
We performed validation of the six CIN and two ER genes in univariate survival association using the online tool Kaplan–Meier plotter24 (Supplementary Results and Supplementary Tables 3 and 4). More importantly, we performed multivariate survival analysis of the 8-gene score in four data sets (with Affymetrix platform from the Gene Expression Omnibus (GEO); GSE2990, GSE3494, GSE2034 and GSE25066). Again, the score was significantly associated with survival in a multivariate Cox proportional hazards model in every data set tested (Figure 4). Altogether, we found that in multiple data sets that used different platforms the 8-gene score identified patients with poor survival independently of other clinico-pathological indicators and outperformed current signatures.
The 8-gene score associates with the survival of breast cancer patients. Four published data sets were used to validate the 8-gene score as a predictor of survival. The 8-gene score was calculated for tumors in each of the data sets, and the survival of patients was stratified according to the median 8-gene score; (a) GSE2990,15 (b) GSE3494,64 (c) GSE203465 and (d) GSE25066.51 The hazard ratio (HR), CI and P-value for comparisons of high vs low 8-gene score are shown in the Kaplan–Meier survival curves (log-rank test, GraphPad Prism). The number of patients (n) is shown in brackets. The table in each panel shows multivariate survival analysis using the Cox proportional hazard model including all available conventional indicators.
Therapeutic targets in the aggressiveness gene list
The overexpressed genes in the CIN metagene are involved in, or regulate, mitosis, spindle assembly and checkpoint, kinetochore attachment, chromosome segregation and mitotic exit. Thus, it is not surprising that several of the overexpressed genes are targets for molecular inhibitors, such as CDK125, 26 and AURKA/AURKB,27 and have been trialed preclinically and clinically.28 To this end, we performed small interfering RNA (siRNA) depletion against 25 genes of the CIN metagene in three TNBC cell lines: MDA-MB-231, SUM159PT and Hs578T. We found that knockdown of four genes (TTK, TPX2, NDC80 and PBK) consistently affected the survival of these cells (Figure 5a and Supplementary Table 2). The knockdown of TTK showed the worst survival, and as it was in the 8-gene score we selected TTK for further studies. We found that TTK protein was higher in TNBC cell lines compared with the near-normal MCF10A cell line and luminal/HER2 cell lines (Figure 5b). Next, we used the specific TTK inhibitor (TTKi) AZ3146 against a panel of breast cancer cell lines and found that TNBC cell lines were more sensitive to the TTKi (Figure 5c).
Therapeutic targets in the aggressiveness gene list. (a) The TNBC cell lines, MDA-MB-231, SUM159PT and Hs578T were treated with control siRNA (Scrambled, Sc CTRL) or siRNA targeting the specified genes, and the survival of these cells was compared on day 6. Data show the average from the three cell lines where each cell line was treated in triplicate. *P<0.05 and ***P<0.001 from one-way ANOVA analysis performed using GraphPad Prism. Data for individual cell lines are shown in Supplementary Table 2. (b) A panel of breast cancer cell lines was used to prepare lysates for immunoblotting of TTK. Tubulin was used as the loading control. (c) Dose response curves for the treatment of breast cancer cell lines in the absence or presence of escalating doses of the TTK inhibitor (TTKi) AZ3146. The survival of cells was measured using the CellTitre MTS/MTA assay carried out 6 days after treatment. Percentage survival (n=3 per dose) was calculated as the percentage of the signal from treated cells to that from control cells. (d) The concentration of TTK required to affect the survival of 50% of the cells (IC50) was measured by GraphPad Prism from the dose response curves in C for each cell line.
TTK expression in aggressive tumors and potential for combination therapy
To further study the potential of TTK as therapeutic target, we investigated TTK expression at the mRNA and protein levels in breast cancer patients. We analyzed the correlation of TTK mRNA expression, dichotomized at the median, with clinico-pathological indicators in the METABRIC data set of 2000 patients (Table 2). High TTK mRNA expression was associated with younger age of tumor diagnosis, larger tumor size, higher tumor grade, higher Ki67 expression, TP53 mutations, an ER/PR-negative tumor phenotype, HER2 positivity and TNBC. On the basis of PAM50 subtyping, high TTK mRNA was associated with luminal B, HER2-enriched and basal-like tumors.
We also analyzed TTK expression in a cohort of breast cancer patients (406 patients) by IHC. TTK and its activity are detected at all stages of the cell cycle; however, TTK is upregulated during mitosis.29 Thus, we observed TTK staining in non-mitotic cells to define high TTK levels (score of 3) in order to exclude the bias of elevated TTK level during mitosis. Similar to TTK mRNA, high TTK protein level (Table 3) was associated with high tumor grade, high Ki67 expression and TNBC status (particularly basal TNBC). Moreover, in agreement with the associations of TTK mRNA with the PAM50 intrinsic subtypes, high TTK protein was observed in HER2-positive and proliferative ER+/HER2− tumors (most related to luminal B) but low TTK protein was observed in nonproliferative ER+/HER2− tumors (most related to luminal A). In addition to these associations with aggressive phenotypes, we also found that high TTK protein significantly associated with aggressive histological features including ductal histology, pushing tumor border, lymph node involvement, nuclear pleomorphism, lymphocytic infiltration and higher mitotic scores (Table 3). Altogether, similar to the high aggressiveness score from the 206 genes or 8 genes, high levels of TTK mRNA and protein span across breast cancer subtypes marking aggressive behavior.
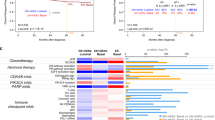
We examined the association of TTK protein level with patient survival and found that breast tumors with high TTK staining (category 3) had worse survival than other staining groups at 5 years (Figures 6a and b) and 10 and 20 years (Supplementary Figure 6). Importantly, high TTK staining (category 3) was not restricted to a particular histological subgroup or to tumors with high mitotic index (Figure 6c). Next, we focused on prognostication of aggressive subgroups (Grade 3, lymph node-positive, TNBC, HER2 or high Ki67) and found that high TTK protein level identified exceptionally aggressive tumors that lead to poor survival of less than 2 years (Figure 7a). Finally, to exploit our finding that TTK, as a part of the aggressiveness score, was associated with aggressive breast tumors and that TTK inhibition was effective in TNBC cell lines that overexpress this protein (Figure 5), we investigated the therapeutic potential of combining TTK inhibition with chemotherapy. We found that TTKi synergized with docetaxel at very low doses (sublethal doses) in the treatment of TNBC cell lines that overexpress TTK in comparison with cell lines that do not (Figure 7b), and that this combination induced apoptotic cell death (Figure 7c).
TTK protein expression associates with breast cancer survival. The overall survival of patients in a large cohort of breast cancer patients (n=409) was stratified according to TTK staining by immunohistochemistry(scores 0–3). Kaplan–Meier survival curves are shown for all patients (a) with four TTK staining (categories 0–3) and (b) two categories (0–2 vs 3). Log-rank test and P-value were used for survival curves. (c) The distribution of high TTK staining (category 3) across histological subgroups and mitotic indices. Data show the mitotic index (median+range) measured as the number of mitotic cells in 10 high-power fields (hpf). The number of tumors with high TTK staining to the total number of tumors in the cohort is shown on the right. High TTK expression distributed across subtypes and did not associate with mitotic index.
TTK associates with aggressive subtypes and is a therapeutic target. (a) Kaplan–Meier survival curves are shown for Grade 3 tumors, lymph node-positive patients (LN+) and LN+ patients with grade 3 tumors. Log-rank Test and P-value were used for these survival curves. For patients with TNBC and HER2, survival was statistically significant using the Gehan–Breslow–Wilcoxon test (P-values marked by two asterisks), which gives more weight to deaths at early time points. The poorer survival of patients with high Ki67 tumors and high TTK staining was a trend, but it did not reach significance. Survival curves and statistical analyses were performed using GraphPad Prism. (b) TNBC and non-TNBC cell lines were treated for 6 days with the specified concentrations of docetaxel (doc) alone, TTK inhibitor (TTKi) alone or the combinations. The survival of cells was measured using the MTS/MTA assay, as described in Methods. ***P<0.001 comparing the combination with single agents and with non-TNBC cell lines from two-way ANOVA in GraphPad Prism. (c) MDA-MB-231 cells were treated with docetaxel or TTKi alone or in combination and collected at 96 h to perform apoptosis assays by flow cytometry. Early apoptotic cells were defined as annexin V+/7-AAD−. **P<0.01 and ***P<0.001 comparing treatments using one-way ANOVA in GraphPad Prism.
Discussion
Our meta-analysis of gene expression in the Oncomine database identified a list of 206 genes enriched with two core biological functions/metagenes: CIN and ER signaling. We calculated the aggressiveness score, the ratio of CIN to ER metagenes, which was associated with the overall survival of breast cancer. A core of eight genes (six CIN genes and two ER signaling genes) was representative and recapitulated the correlations with outcome from the 206 genes. The score from the six CIN genes to the 2 ER signaling genes, 8-gene score, associated with survival in several breast cancer data sets. Our aggressiveness scores outperformed conventional variable and published signatures in multivariate survival analysis. Particularly in ER+ tumors, some cases have survival as poor as that of the aggressive HER2+ and TNBC subtypes. Our data suggest that the interplay of cancer-related biological functions, namely CIN and ER signaling, are better predictors of phenotypes than single genes or single functions. This notion is in line with recent studies showing that the interaction of biologically driven predictors provides better prognosis.16, 17, 30 Recently, all ER− tumors were described to have a high level of CIN metagene; however, it was not clear that ER+ tumors could be described as low CIN tumors.16 In our study, we clarify that ER+ disease contains a considerable fraction of tumors that have a high level of CIN genes and that the relationship between CIN and ER genes is a powerful predictor of survival in these patients.
The fidelity of chromosome segregation is ensured by the proper attachment of the microtubules from the mitotic spindle to the kinetochores of chromosomes in a tightly regulated process, and CIN refers to the missegregation of whole chromosomes, thus producing aneuploidy.31 Using aneuploidy as a surrogate marker for CIN, Carter et al.20 developed a gene signature and found that this ‘CIN signature’ predicts clinical outcome in multiple cancers. More recently, a minimal gene set that captures the CIN signature CIN4 (AURKA, FOXM1, TOP2A and TPX2) was described as the first clinically applicable quantitative PCR-derived measure of tumor aneuploidy from formalin-fixed, paraffin-embedded tissue. As Grade 2 tumors have heterogeneous characteristics in terms of clinical outcome, the significance of the CIN4 classifier is the stratification of Grade 2 tumors into good and poor prognosis groups.23 Our aggressiveness scores were prognostic in all tumor grades and disease stages (stages I–III and lymph node-negative and positive) and outperformed the CIN signature and the CIN4 classifier in multivariate survival analysis in the METABRIC data set. Strikingly, but in agreement with previous studies,32, 33 the prognostication using the CIN metagene and our aggressiveness scores from gene expression levels were restricted to ER+ disease but not in the TNBC or HER2 subtypes. This may be explained by the fact that ER− tumors have a high level of CIN metagene as per our results and those published previously.16 However, our results with TTK protein level clearly demonstrate that TNBC, HER2, high-grade, lymph node-positive and proliferative tumors contain subgroups with high TTK levels exclusive of mitotic cells and have poorer survival than those with low TTK expression or TTK expression in mitotic cells. We propose that there are two types of high expression of CIN genes that may not be clearly differentiated by mRNA expression studies. One form of elevated CIN genes relates to high levels of mitosis and proliferation, whereas the second form that we measured by immunohistochemistry exclusive of mitotic cells is driven by another aggressive phenotype: protection of aneuploidy and genomic instability. The recent study of the CIN4 classifier lends support to our proposition. In this study, using flow cytometry to measure aneuploidy by DNA content, the authors found that a substantial proportion of tumors with high CIN4 scores have a normal DNA ploidy and that a significant proportion of aneuploid cases had a low CIN4 score.23
Chromosome missegregation and aneuploidy enhance genetic recombination and defective DNA damage repair34 to drive a ‘mutator phenotype’ required for oncogenesis.35 Genomic instability caused by deregulated mitotic spindle assembly checkpoint and aneuploidy has been termed ‘non-oncogene addiction’.36, 37 It is tempting to suggest that CIN and aneuploidy are exploited by breast cancer stem cells, which are high in TNBCs38 owing to the link between cancer stem cells, aneuploidy and therapy resistance.39, 40 This is supported by studies that implicate several genes involved in the spindle assembly checkpoint and chromosome segregation in tumor initiation, progression and cancer stem cells, e.g., AURKA in ovarian cancer,41 MELK/FOXM1 in glioblastoma,42, 43 MELK44 and MAD245 in breast cancer and SKP2 in several cancers.46 The role of CIN genes in protecting aneuploidy could provide an insight to the paradox that TNBCs show a better response to chemotherapy owing to the higher level of proliferation, yet these tumors have poorer outcome. We propose that resistance in TNBC could be attributed to the ability of aneuploid cells to adapt and drive recurrence. At least in vivo, chemotherapy has been shown to induce the proliferation quiescent aneuploid cells as a mechanism for therapy resistance.39 We envisage that the high level of the CIN metagene in TNBC, particularly genes involved in chromosome segregation, is protective of this state. Indeed, one study found that a high level of TTK is protective of aneuploidy in breast cancer cells, and its silencing reduces the tumorigenicity of breast cancer cell lines in vivo.47 Our results from the patient cohort demonstrate that high TTK protein expression exclusive of mitosis was indeed prognostic in aggressive tumors and support the concept that protection from aneuploidy and genomic instability is an aggressive phenotype that drives poor outcome.
Our results with the TTK molecular inhibitor, in agreement with published studies using siRNA depletion,47, 48 support the idea of targeting chromosomal segregation in tumors with a high CIN phenotype as a therapeutic strategy. We also suggest that while TTK is high in TNBC, as previously described,47, 48 a considerable proportion of non-TNBC tumors that display aggressive features also show an elevated level of CIN genes, and would benefit from such targeted therapies. To our knowledge, the combination of sublethal doses of taxanes with TTK inhibition has not been investigated so far in breast cancer, but it has been investigated in other cancers.34 Our results reveal that TTK inhibition indeed sensitizes breast cancer cells with high TTK to docetaxel.
In conclusion, our study emphasizes that classification of breast cancer on the basis of biological phenotypes facilitates the understanding of the drivers of oncogenic phenotypes and therapeutic potentials. Importantly, our studies demonstrate that immunohistochemistry assessment of CIN genes, exemplified by TTK here, provide better characterization and understanding for the contribution of CIN to tumor aggressiveness and prognosis.
Materials and methods
Meta-analysis of global gene expression in TNBC
We performed a meta-analysis of global gene expression data in the Oncomine database19 (Compendia Bioscience, Ann Arbor, MI, USA) using a primary filter for breast cancer (130 data sets), sample filter to use clinical specimens and data set filters to use mRNA data sets with more than 151 patients (22 data sets). Patients of all ages, gender, disease stages or treatments were included. Three additional filters were applied to perform three independent differential analyses: (1) triple negative (TNBC cases vs non-TNBC cases, eight data sets;49, 50, 51, 52, 53, 54, 55 (2) metastatic event analysis at 5 years (metastatic events vs no metastatic events, seven data sets51, 53, 56, 57, 58, 59, 60); and (3) survival at 5 years (patients who died vs patients who survived, seven data sets52, 53, 55, 57, 60, 61, 62). Deregulated genes were selected on the basis of the median P-value of the median gene rank in overexpression or underexpression patterns across the data sets (Supplementary Figure 1). The union of these three deregulated gene lists resulted in a gene list of deregulated genes in aggressive breast cancers (Supplementary Figure 2). The METBRIC data set21 was used as the validation set for further analysis. The normalized z-score expression data of the METABRIC data set was extracted from Oncomine and imported into BRB-ArrayTools63 (V4.2, Biometric Research Branch, NCI, Bethesda, MD, USA) with built-in R Bioconductor packages. Survival curves for the METABRIC data set were constructed using GraphPad Prism v6.0 (GraphPad Software, San Diego, CA, USA), and the log-rank (Mantel–Cox) Test was used for statistical comparisons of survival curves.
Ingenuity Pathway Analysis and derivation of the eight-gene list
Pathway analysis was performed using the Ingenuity Pathway Analysis (Ingenuity Systems, Redwood City, CA, USA). For pathway analysis in IPA, we used only direct relationships. After pathway analysis, we set out to identify the minimum gene list that recapitulates the aggressiveness 206-gene list. We used the METABRIC data set to perform statistical filtering in the BRB-ArrayTools software to derive the minimum gene list as follows: (1) the correlation of each gene in the CIN metagene and the ER metagene to the metagene itself was determined by quantitative trait analysis using the Pearson’s correlation coefficient (univariate P-value threshold of 0.001); (2) the association of each gene with overall survival using univariate Cox proportional hazards model (univariate test P-value <0.001); and (3) the fold change of gene expression between high aggressiveness score tumors and low aggressiveness score tumors was calculated for each gene. We selected genes with Pearson’s correlation coefficient >0.7 to the metagenes, strongest survival association and more than two-fold deregulation between high and low agressiveness score tumors. The METABRIC data set and four publically available data sets were used to validate the 8-gene score. The four data sets (GSE25066,51 GSE3494,64 GSE299015 and GSE203465) were analyzed as described previously.66
Cell culture and drug treatments
Breast cancer cell lines were obtained from ATCC (Manassas, VA, USA) and cultured as per ATCC instructions. All cell lines were regularly tested for mycoplasma and authenticated using short tandem repeat profiling. For the siRNA screen, siRNA solutions (Shanghai Gene Pharma, Shanghai, China) were used to transfect cells (MDA-MB-231, SUM159PT and Hs578T) with 10 nM of respective siRNA using Lipofectamine RNAiMAX (Life Technologies, Carlsbad, CA, USA). For drug treatments, docetaxel and the TTK inhibitor AZ3146 were purchased from Selleck Chemicals LLC (Houston, TX, USA) and diluted in dimethylsulfoxide. Six days after siRNA knockdown or after drug treatments, the survival of cells in comparison with control was determined using the CellTiter 96 Assay, as per the manufacturer’s instructions (Promega Corporation, Fitchburg, WI, USA). For immunoblotting, standard protocols were used and membranes were probed with antibodies against TTK (anti-MPS1 mouse monoclonal antibody [N1] ab11108 (Abcam, Cambridge, UK) and γ-tubulin (Sigma-Aldrich, Sydney, NSW, Australia), and then developed using chemiluminescence reagent plus (Millipore, Billerica, MA, USA). Flow cytometry to quantify apoptosis was performed using Annexin V-Alexa488 and 7-AAD (Life Technologies), as per the manufacturer’s instruction by using the BD FACSCanto II flow cytometer (BD Biosciences, San Jose, CA, USA).
Breast cancer tissue microarrays, immunohistochemical and survival analysis
The Brisbane Breast Bank collected fresh breast tumor samples from consenting patients; the study was approved by the local ethics committees. Tissue microarrays were constructed from duplicate cores of formalin-fixed, paraffin-embedded breast tumor samples from patients undergoing resection at the Royal Brisbane and Women’s Hospital between 1987 and 1994. For biomarker analysis, whole tumor sections or tissue microarrays (depending on the marker) were stained with antibodies against ER, PR, Ki67, HER2, CK5/6, CK14, EGFR and TTK (Supplementary Table 5), and scored by trained Pathologists. The Vectastain Universal ABC kit (Vector Laboratories, Burlingame, CA, USA) was used for signal detection according to the manufacturer’s instructions. Stained sections were scanned at high resolution (ScanScope Aperio, Leica Microsystems, Wetzlar, Germany), and then images were segmented into individual cores for analysis using the Spectrum software (Aperio, Wetzlar, Germany). Survival and other clinical data were collected from the Queensland Cancer Registry and original diagnostic Pathology reports, and in addition we performed an internal histopathological review (SRL) of representative tumor sections from each case, stained with H&E. For the analysis of HER2 amplification, tissue microarrays were analyzed using HER2 CISH. Criteria for assigning prognostic subgroups in this study are summarized in Supplementary Figure 7.
Other statistical analysis
Statistical analyses were performed using GraphPad Prism v6.0. The types of tests used are stated in Figure Legends. Univariate and multivariate Cox proportional hazards regression analyses were performed using MedCalc for Windows, version 12.7 (MedCalc Software, Ostend, Belgium).
References
Liedtke C, Mazouni C, Hess KR, Andre F, Tordai A, Mejia JA et al. Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol 2008; 26: 1275–1281.
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F et al. The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 2007; 13: 2329–2334.
von Minckwitz G, Untch M, Blohmer JU, Costa SD, Eidtmann H, Fasching PA et al. Definition and impact of pathologic complete response on prognosis after neoadjuvant chemotherapy in various intrinsic breast cancer subtypes. J Clin Oncol 2012; 30: 1796–1804.
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 2001; 98: 10869–10874.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al. Molecular portraits of human breast tumours. Nature 2000; 406: 747–752.
Weigelt B, Hu Z, He X, Livasy C, Carey LA, Ewend MG et al. Molecular portraits and 70-gene prognosis signature are preserved throughout the metastatic process of breast cancer. Cancer Res 2005; 65: 9155–9158.
Hu Z, Fan C, Oh DS, Marron JS, He X, Qaqish BF et al. The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics 2006; 7: 96–107.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 2009; 27: 1160–1167.
van't Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002; 415: 530–536.
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. New Engl J Med 2004; 351: 2817–2826.
Buyse M, Loi S, van't Veer L, Viale G, Delorenzi M, Glas AM et al. Validation and clinical utility of a 70-gene prognostic signature for women with node-negative breast cancer. J Natl Cancer Inst 2006; 98: 1183–1192.
Loi S, Haibe-Kains B, Desmedt C, Lallemand F, Tutt AM, Gillet C et al. Definition of clinically distinct molecular subtypes in estrogen receptor-positive breast carcinomas through genomic grade. J Clin Oncol 2007; 25: 1239–1246.
Ma XJ, Salunga R, Dahiya S, Wang W, Carney E, Durbecq V et al. A five-gene molecular grade index and HOXB13:IL17BR are complementary prognostic factors in early stage breast cancer. Clin Cancer Res 2008; 14: 2601–2608.
Ma XJ, Wang Z, Ryan PD, Isakoff SJ, Barmettler A, Fuller A et al. A two-gene expression ratio predicts clinical outcome in breast cancer patients treated with tamoxifen. Cancer Cell 2004; 5: 607–616.
Sotiriou C, Wirapati P, Loi S, Harris A, Fox S, Smeds J et al. Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Nl Cancer Inst 2006; 98: 262–272.
Cheng WY, Ou Yang TH, Anastassiou D . Biomolecular events in cancer revealed by attractor metagenes. PLoS Comput Biol 2013; 9: e1002920.
Cheng WY, Ou Yang TH, Anastassiou D . Development of a prognostic model for breast cancer survival in an open challenge environment. Sci Transl Med 2013; 5: 181ra150.
Dai H, van't Veer L, Lamb J, He YD, Mao M, Fine BM et al. A cell proliferation signature is a marker of extremely poor outcome in a subpopulation of breast cancer patients. Cancer Res 2005; 65: 4059–4066.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia (New York, NY) 2004; 6: 1–6.
Carter SL, Eklund AC, Kohane IS, Harris LN, Szallasi Z . A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet 2006; 38: 1043–1048.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 2012; 486: 346–352.
Haibe-Kains B, Desmedt C, Rothe F, Piccart M, Sotiriou C, Bontempi G . A fuzzy gene expression-based computational approach improves breast cancer prognostication. Genome Biol 2010; 11: R18.
Szasz AM, Li Q, Eklund AC, Sztupinszki Z, Rowan A, Tokes AM et al. The CIN4 chromosomal instability qPCR classifier defines tumor aneuploidy and stratifies outcome in grade 2 breast cancer. PLoS ONE 2013; 8: e56707.
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q et al. An online survival analysis tool to rapidly assess the effect of 22 277 genes on breast cancer prognosis using microarray data of 1809 patients. Breast Cancer Res Treat 2010; 123: 725–731.
Rizzolio F, Tuccinardi T, Caligiuri I, Lucchetti C, Giordano A . CDK inhibitors: from the bench to clinical trials. Curr Drug Targets 2010; 11: 279–290.
Horiuchi D, Kusdra L, Huskey NE, Chandriani S, Lenburg ME, Gonzalez-Angulo AM et al. MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J Exp Med 2012; 209: 679–696.
Manchado E, Guillamot M, Malumbres M . Killing cells by targeting mitosis. Cell Death Differ 2012; 19: 369–377.
Janssen A, Medema RH . Mitosis as an anti-cancer target. Oncogene 2011; 30: 2799–2809.
Stucke VM, Sillje HH, Arnaud L, Nigg EA . Human Mps1 kinase is required for the spindle assembly checkpoint but not for centrosome duplication. EMBO J 2002; 21: 1723–1732.
Nagalla S, Chou JW, Willingham MC, Ruiz J, Vaughn JP, Dubey P et al. Interactions between immunity, proliferation and molecular subtype in breast cancer prognosis. Genome Biol 2013; 14: R34.
Bakhoum SF, Compton DA . Chromosomal instability and cancer: a complex relationship with therapeutic potential. J Clin Invest 2012; 122: 1138–1143.
Roylance R, Endesfelder D, Gorman P, Burrell RA, Sander J, Tomlinson I et al. Relationship of extreme chromosomal instability with long-term survival in a retrospective analysis of primary breast cancer. Cancer Epidemiol Biomarkers Prev 2011; 20: 2183–2194.
Birkbak NJ, Eklund AC, Li Q, McClelland SE, Endesfelder D, Tan P et al. Paradoxical relationship between chromosomal instability and survival outcome in cancer. Cancer Res 2011; 71: 3447–3452.
Janssen A, van der Burg M, Szuhai K, Kops GJ, Medema RH . Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 2011; 333: 1895–1898.
Kolodner RD, Cleveland DW, Putnam CD . Cancer. Aneuploidy drives a mutator phenotype in cancer. Science 2011; 333: 942–943.
Luo J, Solimini NL, Elledge SJ . Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 2009; 136: 823–837.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Al-Ejeh F, Smart CE, Morrison BJ, Chenevix-Trench G, Lopez JA, Lakhani SR et al. Breast cancer stem cells: treatment resistance and therapeutic opportunities. Carcinogenesis 2011; 32: 650–658.
Kusumbe AP, Bapat SA . Cancer stem cells and aneuploid populations within developing tumors are the major determinants of tumor dormancy. Cancer Res 2009; 69: 9245–9253.
Liang Y, Zhong Z, Huang Y, Deng W, Cao J, Tsao G et al. Stem-like cancer cells are inducible by increasing genomic instability in cancer cells. JBiol Chem 2010; 285: 4931–4940.
Do TV, Xiao F, Bickel LE, Klein-Szanto AJ, Pathak HB, Hua X et al. Aurora kinase A mediates epithelial ovarian cancer cell migration and adhesion. Oncogene 2014; 33: 539–549.
Joshi K, Banasavadi-Siddegowda Y, Mo X, Kim SH, Mao P, Kig C et al. MELK-dependent FOXM1 phosphorylation is essential for proliferation of glioma stem cells. Stem cells 2013; 31: 1051–1063.
Gu C, Banasavadi-Siddegowda YK, Joshi K, Nakamura Y, Kurt H, Gupta S et al. Tumor-specific activation of the C-JUN/MELK pathway regulates glioma stem cell growth in a p53-dependent manner. Stem cells 2013; 31: 870–881.
Hebbard LW, Maurer J, Miller A, Lesperance J, Hassell J, Oshima RG et al. Maternal embryonic leucine zipper kinase is upregulated and required in mammary tumor-initiating cells in vivo. Cancer Res 2010; 70: 8863–8873.
Schvartzman JM, Duijf PH, Sotillo R, Coker C, Benezra R . Mad2 is a critical mediator of the chromosome instability observed upon Rb and p53 pathway inhibition. Cancer Cell 2011; 19: 701–714.
Chan CH, Morrow JK, Li CF, Gao Y, Jin G, Moten A et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 2013; 154: 556–568.
Daniel J, Coulter J, Woo JH, Wilsbach K, Gabrielson E . High levels of the Mps1 checkpoint protein are protective of aneuploidy in breast cancer cells. Proc Natl Acad Sci USA 2011; 108: 5384–5389.
Maire V, Baldeyron C, Richardson M, Tesson B, Vincent-Salomon A, Gravier E et al. TTK/hMPS1 is an attractive therapeutic target for triple-negative breast cancer. PLoS ONE 2013; 8: e63712.
Bonnefoi H, Potti A, Delorenzi M, Mauriac L, Campone M, Tubiana-Hulin M et al. Validation of gene signatures that predict the response of breast cancer to neoadjuvant chemotherapy: a substudy of the EORTC 10994/BIG 00-01 clinical trial. Lancet Oncol 2007; 8: 1071–1078.
Gluck S, Ross JS, Royce M, McKenna EF Jr, Perou CM, Avisar E et al. TP53 genomics predict higher clinical and pathologic tumor response in operable early-stage breast cancer treated with docetaxel-capecitabine +/− trastuzumab. Breast Cancer Res Treatt 2012; 132: 781–791.
Hatzis C, Pusztai L, Valero V, Booser DJ, Esserman L, Lluch A et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 2011; 305: 1873–1881.
Bild AH, Yao G, Chang JT, Wang Q, Potti A, Chasse D et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 2006; 439: 353–357.
Kao KJ, Chang KM, Hsu HC, Huang AT . Correlation of microarray-based breast cancer molecular subtypes and clinical outcomes: implications for treatment optimization. BMC Cancer 2011; 11: 143.
Tabchy A, Valero V, Vidaurre T, Lluch A, Gomez H, Martin M et al. Evaluation of a 30-gene paclitaxel, fluorouracil, doxorubicin, and cyclophosphamide chemotherapy response predictor in a multicenter randomized trial in breast cancer. Clin Cancer Res 2010; 16: 5351–5361.
TCGA, Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX et al. Genes that mediate breast cancer metastasis to the brain. Nature 2009; 459: 1005–1009.
Desmedt C, Piette F, Loi S, Wang Y, Lallemand F, Haibe-Kains B et al. Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the TRANSBIG multicenter independent validation series. Clin Cancer Res 2007; 13: 3207–3214.
Schmidt M, Bohm D, von Torne C, Steiner E, Puhl A, Pilch H et al. The humoral immune system has a key prognostic impact in node-negative breast cancer. Cancer Res 2008; 68: 5405–5413.
Symmans WF, Hatzis C, Sotiriou C, Andre F, Peintinger F, Regitnig P et al. Genomic index of sensitivity to endocrine therapy for breast cancer. J Clin Oncol 2010; 28: 4111–4119.
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW et al. A gene-expression signature as a predictor of survival in breast cancer. New Engl J Med 2002; 347: 1999–2009.
Pawitan Y, Bjohle J, Amler L, Borg AL, Egyhazi S, Hall P et al. Gene expression profiling spares early breast cancer patients from adjuvant therapy: derived and validated in two population-based cohorts. Breast Cancer Res 2005; 7: R953–R964.
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 2003; 100: 8418–8423.
Zhao Y, Simon R . BRB-ArrayTools Data Archive for human cancer gene expression: a unique and efficient data sharing resource. Cancer Inform 2008; 6: 9–15.
Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A et al. An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc Natl Acad Sci USA 2005; 102: 13550–13555.
Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 2005; 365: 671–679.
Al-Ejeh F, Shi W, Miranda M, Simpson PT, Vargas AC, Song S et al. Treatment of triple-negative breast cancer using anti-EGFR-directed radioimmunotherapy combined with radiosensitizing chemotherapy and PARP inhibitor. J Nucl Med 2013; 54: 913–921.
Acknowledgements
FA is funded by the Cure Cancer Foundation Australia/Priority-driven Collaborative Cancer Research Fund (ID: 631924) and Future Fellowship from the Australian Research Council (ID: FT130101417); PTS is funded by a Fellowship from the National Breast Cancer Foundation, Australia; and SRL, GC-T and KKK are supported by NHMRC Senior Principal Research Fellowship (ID 1017028). We acknowledge The Brisbane Breast Bank (in particular Mythily Mariasegaram and Colleen Niland) for coordinating sample and clinical data collection. We thank Margaret C Cummings, Darina Vuong and Tristan Molden-Hauer for their invaluable contributions to the breast cancer tissue microarrays resource used in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogenesis website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Al-Ejeh, F., Simpson, P., Sanus, J. et al. Meta-analysis of the global gene expression profile of triple-negative breast cancer identifies genes for the prognostication and treatment of aggressive breast cancer. Oncogenesis 3, e100 (2014). https://doi.org/10.1038/oncsis.2014.14
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2014.14
Keywords
This article is cited by
-
Digital image analysis and machine learning-assisted prediction of neoadjuvant chemotherapy response in triple-negative breast cancer
Breast Cancer Research (2024)
-
Considerations for feature selection using gene pairs and applications in large-scale dataset integration, novel oncogene discovery, and interpretable cancer screening
BMC Medical Genomics (2020)
-
Cep55 overexpression promotes genomic instability and tumorigenesis in mice
Communications Biology (2020)
-
Study of Gene Expression Profiles of Breast Cancers in Indian Women
Scientific Reports (2019)
-
Estrogen receptor-α regulation of microRNA-590 targets FAM171A1—a modifier of breast cancer invasiveness
Oncogenesis (2019)