Abstract
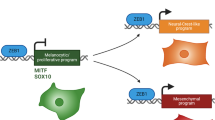
The use of adherent monolayer cultures have produced many insights into melanoma cell growth and differentiation, but often novel therapeutics demonstrated to act on these cells are not active in vivo. It is imperative that new methods of growing melanoma cells that reflect growth in vivo are investigated. To this end, a range of human melanoma cell lines passaged as adherent cultures or induced to form melanoma spheres (melanospheres) in stem cell media have been studied to compare cellular characteristics and protein expression. Melanoma spheres and tumours grown from cell lines as mouse xenografts had increased heterogeneity when compared with adherent cells and 3D-spheroids in agar (aggregates). Furthermore, cells within the melanoma spheres and mouse xenografts each displayed a high level of reciprocal BRN2 or MITF expression, which matched more closely the pattern seen in human melanoma tumours in situ, rather than the propensity for co-expression of these important melanocytic transcription factors seen in adherent cells and 3D-spheroids. Notably, when the levels of the BRN2 and MITF proteins were each independently repressed using siRNA treatment of adherent melanoma cells, members of the NOTCH pathway responded by decreasing or increasing expression, respectively. This links BRN2 as an activator, and conversely, MITF as a repressor of the NOTCH pathway in melanoma cells. Loss of the BRN2-MITF axis in antisense-ablated cell lines decreased the melanoma sphere-forming capability, cell adhesion during 3D-spheroid formation and invasion through a collagen matrix. Combined, this evidence suggests that the melanoma sphere-culture system induces subpopulations of cells that may more accurately portray the in vivo disease, than the growth as adherent melanoma cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Alonso SR, Tracey L, Ortiz P, Perez-Gomez B, Palacios J, Pollan M et al. (2007). A high-throughput study in melanoma identifies epithelial-mesenchymal transition as a major determinant of metastasis. Cancer Res 67: 3450–3460.
Carreira S, Goodall J, Denat L, Rodriguez M, Nuciforo P, Hoek KS et al. (2006). Mitf regulation of Dia1 controls melanoma proliferation and invasiveness. Genes Dev 20: 3426–3439.
Castro DS, Skowronska-Krawczyk D, Armant O, Donaldson IJ, Parras C, Hunt C et al. (2006). Proneural bHLH and Brn proteins coregulate a neurogenic program through cooperative binding to a conserved DNA motif. Dev Cell 11: 831–844.
Cheli Y, Ohanna M, Ballotti R, Bertolotto C . (2010). Fifteen-year quest for microphthalmia-associated transcription factor target genes. Pigment Cell Melanoma Res 23: 27–40.
Cook AL, Boyle GM, Leonard JH, Parsons PG, Sturm RA . (2006). BRN2 in melanocytic cell development, differentiation and transformation. In: Hearing VJ and Leong SPL (eds). From Melanocytes to Melanoma. Humana Press: Totowa, New Jersey. pp 149–167.
Cook AL, Donatien PD, Smith AG, Murphy M, Jones MK, Herlyn M et al. (2003). Human melanoblasts in culture: expression of BRN2 and synergistic regulation by fibroblast growth factor-2, stem cell factor, and endothelin-3. J Invest Dermatol 121: 1150–1159.
Cook AL, Smith AG, Smit DJ, Leonard JH, Sturm RA . (2005). Co-expression of SOX9 and SOX10 during melanocytic differentiation in vitro. Exp Cell Res 308: 222–235.
Cook AL, Sturm RA . (2008). POU domain transcription factors: BRN2 as a regulator of melanocytic growth and tumourigenesis. Pigment Cell Melanoma Res 21: 611–626.
Dodd IB, Micheelsen MA, Sneppen K, Thon G . (2007). Theoretical analysis of epigenetic cell memory by nucleosome modification. Cell 129: 813–822.
Fang D, Nguyen TK, Leishear K, Finko R, Kulp AN, Hotz S et al. (2005). A tumorigenic subpopulation with stem cell properties in melanomas. Cancer Res 65: 9328–9337.
Goodall J, Carreira S, Denat L, Kobi D, Davidson I, Nuciforo P et al. (2008). Brn-2 represses microphthalmia-associated transcription factor expression and marks a distinct subpopulation of microphthalmia-associated transcription factor-negative melanoma cells. Cancer Res 68: 7788–7794.
Hendrix MJ, Seftor EA, Seftor RE, Kasemeier-Kulesa J, Kulesa PM, Postovit LM . (2007). Reprogramming metastatic tumour cells with embryonic microenvironments. Nat Rev Cancer 7: 246–255.
Hoek KS, Eichhoff OM, Schlegel NC, Dobbeling U, Kobert N, Schaerer L et al. (2008). In vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res 68: 650–656.
Hoek KS, Goding CR . (2010). Cancer stem cells versus phenotype-switching in melanoma. Pigment Cell Melanoma Res 23: 746–759.
Houghton AN, Real FX, Davis LJ, Cordon-Cardo C, Old LJ . (1987). Phenotypic heterogeneity of melanoma. Relation to the differentiation program of melanoma cells. J Exp Med 165: 812–829.
Johansson P, Pavey S, Hayward N . (2007). Confirmation of a BRAF mutation-associated gene expression signature in melanoma. Pigment Cell Res 20: 216–221.
Kobi D, Steunou AL, Dembele D, Legras S, Larue L, Nieto L et al. (2010). Genome-wide analysis of POU3F2/BRN2 promoter occupancy in human melanoma cells reveals Kitl as a novel regulated target gene. Pigment Cell Melanoma Res 23: 404–418.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY et al. (2008). The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133: 704–715.
Miller AJ, Mihm Jr MC . (2006). Melanoma. N Engl J Med 355: 51–65.
Mouriaux F, Vincent S, Kherrouche Z, Maurage CA, Planque N, Monte D et al. (2003). Microphthalmia transcription factor analysis in posterior uveal melanomas. Exp Eye Res 76: 653–661.
Na YR, Seok SH, Kim DJ, Han JH, Kim TH, Jung H et al. (2009a). Isolation and characterization of spheroid cells from human malignant melanoma cell line WM-266-4. Tumour Biol 30: 300–309.
Na YR, Seok SH, Kim DJ, Han JH, Kim TH, Jung H et al. (2009b). Zebrafish embryo extracts promote sphere-forming abilities of human melanoma cell line. Cancer Sci 100: 1429–1433.
Newton RA, Cook AL, Roberts DW, Leonard JH, Sturm RA . (2007). Post-transcriptional regulation of melanin biosynthetic enzymes by cAMP and resveratrol in human melanocytes. J Invest Dermatol 127: 2216–2227.
Ohsie SJ, Sarantopoulos GP, Cochran AJ, Binder SW . (2008). Immunohistochemical characteristics of melanoma. J Cutan Pathol 35: 433–444.
Pavey S, Johansson P, Packer L, Taylor J, Stark M, Pollock PM et al. (2004). Microarray expression profiling in melanoma reveals a BRAF mutation signature. Oncogene 23: 4060–4067.
Perego M, Tortoreto M, Tragni G, Mariani L, Deho P, Carbone A et al. (2010). Heterogeneous phenotype of human melanoma cells with in vitro and in vivo features of tumor-initiating cells. J Invest Dermatol 130: 1877–1886.
Pinner S, Jordan P, Sharrock K, Bazley L, Collinson L, Marais R et al. (2009). Intravital imaging reveals transient changes in pigment production and Brn2 expression during metastatic melanoma dissemination. Cancer Res 69: 7969–7977.
Pinnix CC, Herlyn M . (2007). The many faces of Notch signaling in skin-derived cells. Pigment Cell Res 20: 458–465.
Pinnix CC, Lee JT, Liu ZJ, McDaid R, Balint K, Beverly LJ et al. (2009). Active Notch1 confers a transformed phenotype to primary human melanocytes. Cancer Res 69: 5312–5320.
Quintana E, Shackleton M, Sabel MS, Fullen DR, Johnson TM, Morrison SJ . (2008). Efficient tumour formation by single human melanoma cells. Nature 456: 593–598.
Richmond-Sinclair NM, Lee E, Cummings MC, Williamson R, Muller K, Green AC et al. (2008). Histologic and epidemiologic correlates of P-MAPK, Brn-2, pRb, p53, and p16 immunostaining in cutaneous melanomas. Melanoma Res 18: 336–345.
Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, Gasser M et al. (2008). Identification of cells initiating human melanomas. Nature 451: 345–349.
Smalley KS, Haass NK, Brafford PA, Lioni M, Flaherty KT, Herlyn M . (2006). Multiple signaling pathways must be targeted to overcome drug resistance in cell lines derived from melanoma metastases. Mol Cancer Ther 5: 1136–1144.
Smit DJ, Gardiner BB, Sturm RA . (2007). Osteonectin downregulates E-cadherin, induces osteopontin and focal adhesion kinase activity stimulating an invasive melanoma phenotype. Int J Cancer 121: 2653–2660.
Smith AG, Brightwell G, Smit SE, Parsons PG, Sturm RA . (1998). Redox regulation of Brn-2/N-Oct-3 POU domain DNA binding activity and proteolytic formation of N-Oct-5 during melanoma cell nuclear extraction. Melanoma Res 8: 2–10.
Thomson JAF, Murphy K, Baker E, Sutherland GR, Parsons PG, Sturm RA . (1995). The brn-2 gene regulates the melanocytic phenotype and tumorigenic potential of human melanoma cells. Oncogene 11: 691–700.
Acknowledgements
We thank Professor N Hayward (QIMR) for providing some of the cell lines that were tested for melanoma sphere formation as part of this manuscript. RAS is a Senior Research Fellow of the Australian NHMRC, and in conducting part of this work, was supported by an SSP award from the University of Queensland. This work was supported by grants from the Cancer Council Queensland to RAS and JHL and NIH grant CA-25874 to MH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Thurber, A., Douglas, G., Sturm, E. et al. Inverse expression states of the BRN2 and MITF transcription factors in melanoma spheres and tumour xenografts regulate the NOTCH pathway. Oncogene 30, 3036–3048 (2011). https://doi.org/10.1038/onc.2011.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.33
Keywords
This article is cited by
-
A unique hyperdynamic dimer interface permits small molecule perturbation of the melanoma oncoprotein MITF for melanoma therapy
Cell Research (2023)
-
De-novo and genome-wide meta-analyses identify a risk haplotype for congenital sensorineural deafness in Dalmatian dogs
Scientific Reports (2022)
-
BRN2 is a non-canonical melanoma tumor-suppressor
Nature Communications (2021)
-
Identification and quantification of notch receptors in human cutaneous melanoma using molecular biology techniques: literature review
Surgical and Experimental Pathology (2020)
-
BRN2 expression increases anoikis resistance in melanoma
Oncogenesis (2020)